Substitution Numbers and Scoring Matrices Substitution Numbers q

Substitution Numbers and Scoring Matrices

Substitution Numbers q The number of observed substitutions K is an important quantity in molecular evolutionary analysis q A simple count may be misleading, so statistical models are developed to estimate the number of substitutions Ø Ø Ø Jukes-Cantor model Kimura model (both are for nucleotides, but the ideas can extend to amino acids)
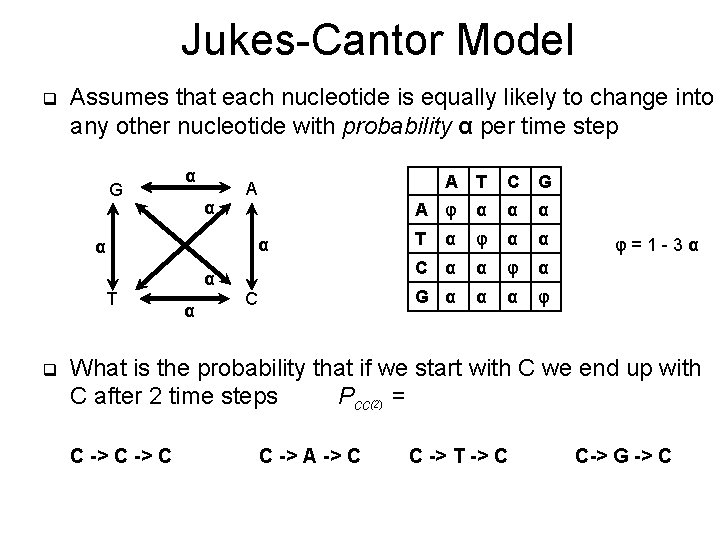
Jukes-Cantor Model q Assumes that each nucleotide is equally likely to change into any other nucleotide with probability α per time step G α α A T A α α α T q α C C G A φ α α α T α φ α α C α α φ α G α α α φ φ=1 -3α What is the probability that if we start with C we end up with C after 2 time steps Pcc(2) = C -> C C -> A -> C C -> T -> C C-> G -> C
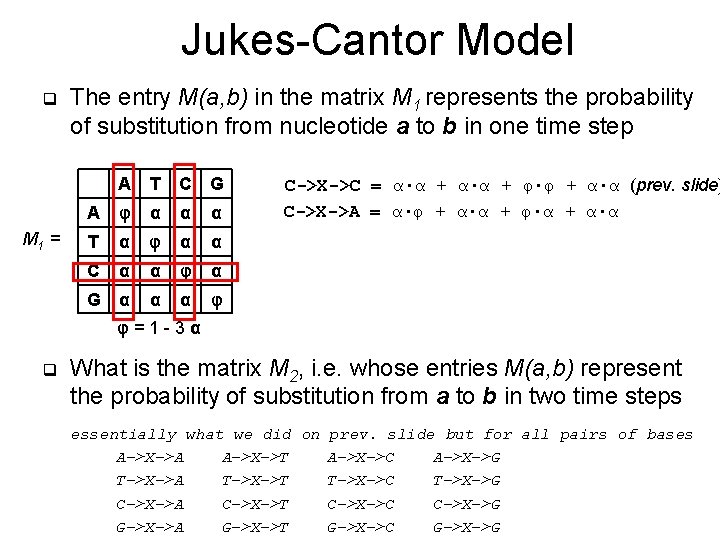
Jukes-Cantor Model q M 1 = The entry M(a, b) in the matrix M 1 represents the probability of substitution from nucleotide a to b in one time step A T C G A φ α α α T α φ α α C α α φ α G α α α φ C->X->C = α∙α + φ∙φ + α∙α (prev. slide) C->X->A = α∙φ + α∙α + φ∙α + α∙α φ=1 -3α q What is the matrix M 2, i. e. whose entries M(a, b) represent the probability of substitution from a to b in two time steps essentially what we did on prev. slide but for all pairs of bases A->X->A A->X->T A->X->C A->X->G T->X->A T->X->T T->X->C T->X->G C->X->A C->X->T C->X->C C->X->G G->X->A G->X->T G->X->C G->X->G

Jukes-Cantor Model q Turns out that Mn = (M 1)n i. e. whose entries M(a, b) represent the probability of substitution from a to b in n time steps q In general under the J. C. model the probability that a site will contain a C after t time steps is given by: Pc(t) = ¼ + (¾)e-4αt q This model can be used to derive an estimate of the number of substitutions that have occurred between the sequences K = -¾ ln[ 1 – (4/3) p ] p – the fraction of nucleotides that are considered mismatch
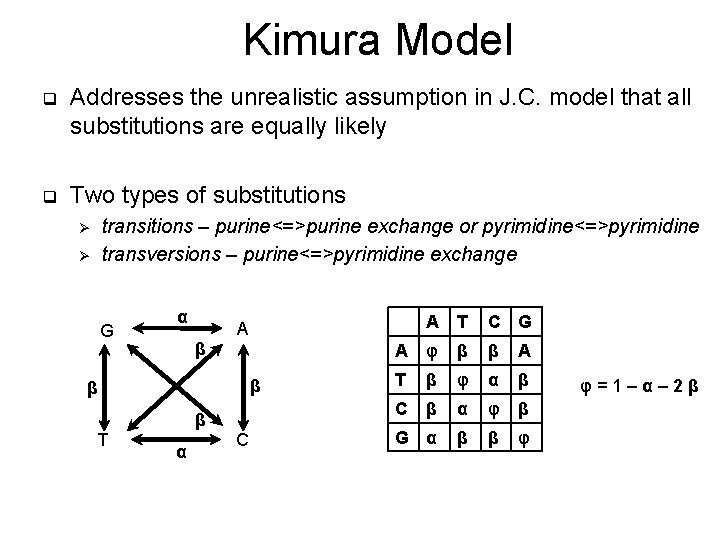
Kimura Model q Addresses the unrealistic assumption in J. C. model that all substitutions are equally likely q Two types of substitutions transitions – purine<=>purine exchange or pyrimidine<=>pyrimidine transversions – purine<=>pyrimidine exchange Ø Ø G α β β T A α C C G A φ β β Α T β φ α β C β α φ β G α β β φ φ=1–α– 2β

Kimura Model q What is the probability that if we start with C we end up with C after 2 time steps Pcc(2) = C -> C q C -> A -> C C -> T -> C C-> G -> C In general under the Kimura model the probability that a site will contain a C after t time steps is given by: Pc(t) = ¼ + (¼)e-4βt + (½)e-2(α+β)t q Estimated number of substitutions (TR – transitions, TV – transverions K = ½ ln[ 1 / (1 – 2*TR – TV)] + ¼ ln[ 1 / (1 – 2*TV)]

Scoring Matrices

Alignment Score q Alignment score attempts to measure likelihood of a common evolutionary ancestor q Two possible ways to explain a given pairwise alignment Ø Ø q Under random model Ø Ø q random model – the alignment could be produced purely by chance evolutionary model – there is high correlation between aligned pairs each position is independent of the others probability of amino acid a occurring at each position is pa Under non-random model Ø probability of amino acid a depends on matched residue b – qab

Substitution Matrices q Given a (non-gapped) pairwise alignment of sequences A = a 1 a 2 a 3 a 4…an B = b 1 b 2 b 3 b 4…bn Ø under non-random model probability of the alignment Pnon-random = qa 1 b 1 qa 2 b 2 qa 3 b 3 qa 4 b 4…qanbn Ø under random model probability of the alignment Prandom = pa 1 pa 2 pa 3 pa 4…pan pb 1 pb 2 pb 3 pb 4…pbn = pa 1 pb 1 pa 2 pb 2 pa 3 pb 3 qa 4 pb 4…panpbn q Use ratio of probabilities (odds ratio) to compare the models Pnon-random r = –––– r > 1, non-random more likely Prandom

Substitution Matrices q Ratio of probabilities (odds ratio) Pnon-random qa 1 b 1 qa 2 b 2 qa 3 b 3 qa 4 b 4 …qanbn r = ––––––––––––––––––– Prandom pa 1 pb 1 pa 2 pb 2 pa 3 pb 3 qa 4 pb 4…panpbn qa 1 b 1 qa 2 b 2 qa 3 b 3 qa 4 b 4 … qanbn = ––––––––––––––– pa 1 pb 1 pa 2 pb 2 pa 3 pb 3 qa 4 pb 4 …panpbn q Typically the log-odds ratio is used qa 1 b 1 qa 2 b 2 qa 3 b 3 qa 4 b 4 … qanbn log(r) = log( ––––––––––––––– ) Entry (a 1, b 1) in the substitution matrix pa 1 pb 1 pa 2 pb 2 pa 3 pb 3 qa 4 pb 4 …panpbn qa 1 b 1 qanbn qa 2 b 2 qa 3 b 3 = log(––––––)+ log( ––––––) +. . . +log( ––––––) pa 1 pb 1 pa 2 pb 2 panpbn pa 3 pb 3
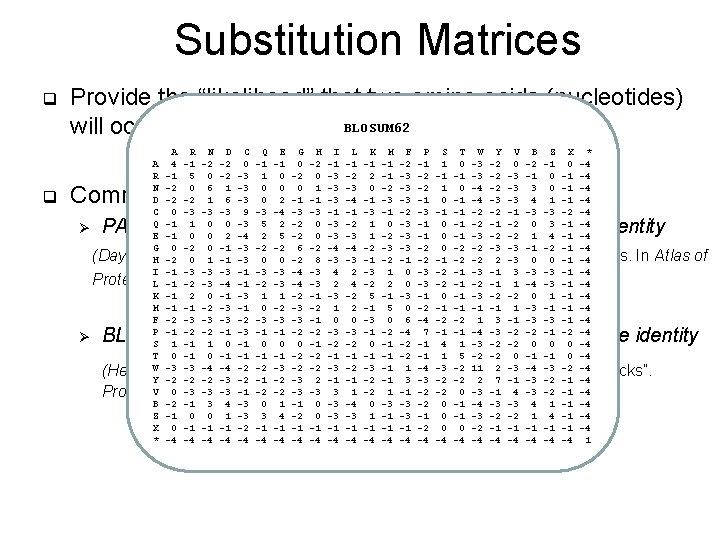
Substitution Matrices q q Provide the “likelihood” that two amino acids (nucleotides) will occur as aligned pair BLOSUM 62 A R N D C Q E G H I L K M F P S T W Y V B Z X * A 4 -1 -2 -2 0 -1 -1 0 -2 -1 -1 -2 -1 1 0 -3 -2 0 -2 -1 0 -4 R -1 5 0 -2 -3 1 0 -2 0 -3 -2 2 -1 -3 -2 -1 -1 -3 -2 -3 -1 0 -1 -4 N -2 0 6 1 -3 0 0 0 1 -3 -3 0 -2 -3 -2 1 0 -4 -2 -3 3 0 -1 -4 D -2 -2 1 6 -3 0 2 -1 -1 -3 -4 -1 -3 -3 -1 0 -1 -4 -3 -3 4 1 -1 -4 C 0 -3 -3 -3 9 -3 -4 -3 -3 -1 -1 -3 -1 -2 -3 -1 -1 -2 -2 -1 -3 -3 -2 -4 Q -1 1 0 0 -3 5 2 -2 0 -3 -2 1 0 -3 -1 0 -1 -2 0 3 -1 -4 E -1 0 0 2 -4 2 5 -2 0 -3 -3 1 -2 -3 -1 0 -1 -3 -2 -2 1 4 -1 -4 G 0 -2 0 -1 -3 -2 -2 6 -2 -4 -4 -2 -3 -3 -2 0 -2 -2 -3 -3 -1 -2 -1 -4 H -2 0 1 -1 -3 0 0 -2 8 -3 -3 -1 -2 -2 2 -3 0 0 -1 -4 I -1 -3 -3 -3 -1 -3 -3 -4 -3 4 2 -3 1 0 -3 -2 -1 -3 -1 3 -3 -3 -1 -4 L -1 -2 -3 -4 -3 2 4 -2 2 0 -3 -2 -1 1 -4 -3 -1 -4 K -1 2 0 -1 -3 1 1 -2 -1 -3 -2 5 -1 -3 -1 0 -1 -3 -2 -2 0 1 -1 -4 M -1 -1 -2 -3 -1 0 -2 -3 -2 1 2 -1 5 0 -2 -1 -1 1 -3 -1 -1 -4 F -2 -3 -3 -3 -1 0 0 -3 0 6 -4 -2 -2 1 3 -1 -3 -3 -1 -4 P -1 -2 -2 -1 -3 -1 -1 -2 -2 -3 -3 -1 -2 -4 7 -1 -1 -4 -3 -2 -2 -1 -2 -4 S 1 -1 1 0 -1 0 0 0 -1 -2 -2 0 -1 -2 -1 4 1 -3 -2 -2 0 0 0 -4 T 0 -1 -1 -2 -2 -1 -1 -2 -1 1 5 -2 -2 0 -1 -1 0 -4 W -3 -3 -4 -4 -2 -2 -3 -1 1 -4 -3 -2 11 2 -3 -4 -3 -2 -4 Y -2 -2 -2 -3 -2 -1 -2 -3 2 -1 -1 -2 -1 3 -3 -2 -2 2 7 -1 -3 -2 -1 -4 V 0 -3 -3 -3 -1 -2 -2 -3 -3 3 1 -2 1 -1 -2 -2 0 -3 -1 4 -3 -2 -1 -4 B -2 -1 3 4 -3 0 1 -1 0 -3 -4 0 -3 -3 -2 0 -1 -4 -3 -3 4 1 -1 -4 Z -1 0 0 1 -3 3 4 -2 0 -3 -3 1 -1 -3 -1 0 -1 -3 -2 -2 1 4 -1 -4 X 0 -1 -1 -1 -2 -1 -1 -1 -2 0 0 -2 -1 -1 -1 -4 * -4 -4 -4 -4 -4 -4 1 Common substitution matrices for protein alignment Ø PAM family – derived from alignments of high sequence identity (Dayhoff, Schwartz, and Orcutt. “A model of evolutionary change in proteins. In Atlas of Protein Sequence and Structure Volume 5. 1978: 345 -352) Ø BLOSUM family – derived from alignments of low sequence identity (Henikoff and Henikoff. “Amino acid substitution matrices from protein blocks”. Proc. Natl. Acad. Sci. 1992. 89(22): 10915– 10919. )

BLOSUM Matrices q Based on ungapped multiple local alignments of conserved regions of proteins with low sequence identity q These alignments are used to derive qab pa pb which give the substitution score for amino acids a and b q ab score(a, b) = log(––––––) pa pb q Procedure Ø Ø obtain known ungapped multiple local alignments split into clusters, so that every pair in a cluster has ≥ C% identity for each pair of amino acids a and b calculate qab = frequency of a, b pair / total # pairs (sequences within a cluster are given weight 1 / size_of_cluster)
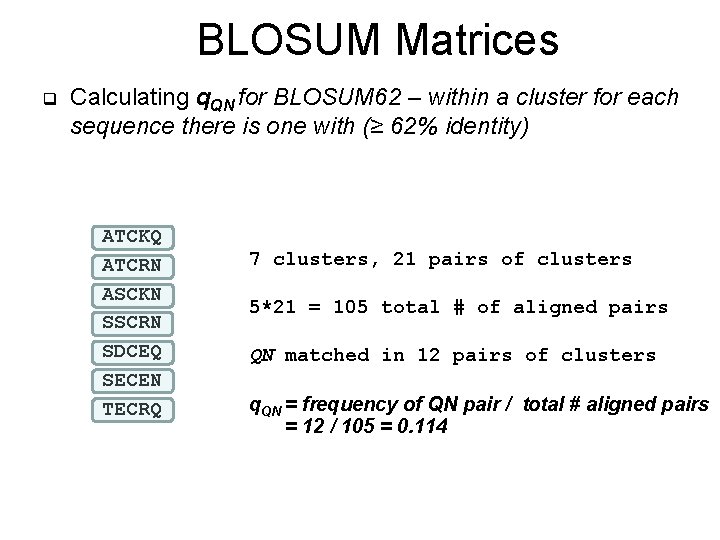
BLOSUM Matrices q Calculating q. QN for BLOSUM 62 – within a cluster for each sequence there is one with (≥ 62% identity) ATCKQ ATCRN ASCKN SSCRN SDCEQ SECEN TECRQ 7 clusters, 21 pairs of clusters 5*21 = 105 total # of aligned pairs QN matched in 12 pairs of clusters q. QN = frequency of QN pair / total # aligned pairs = 12 / 105 = 0. 114

BLOSUM Matrices q Calculating q. QN for BLOSUM 50 – within a cluster for each sequence there is one with (≥ 50% identity) ATCKQ ATCRN ASCKN SSCRN SDCEQ SECEN TECRQ 3 clusters, 3 pairs of clusters 5 bases * 3 clusters = 15 total # of aligned pairs QN match frequency (between clusters): top, mid: top, bot: mid, bot: total: q. QN = frequency of QN pair / total # aligned pairs = 14/8 / 15 = 0. 1166

BLOSUM Matrices q Calculating q. QN for BLOSUM 50 – within a cluster for each sequence there is one with (≥ 50% identity) ATCKQ ATCRN ASCKN SSCRN SDCEQ SECEN TECRQ 3 clusters, 3 pairs of clusters 5 bases * 3 clusters = 15 total # of aligned pairs QN match frequency (between clusters): top, mid: ¼*½ + ¾*½ top, bot: ¾*1 mid, bot: ½*1 total: 1/8+3/4+1/2 = 14/8 q. QN = frequency of QN pair / total # aligned pairs = 14/8 / 15 = 0. 1166

BLOSUM Matrices q So far calculated qab. N (i. e. probability that a and b will be paired up under non-random model) q To compute the substitution score need to know pa and pb (i. e. probability that a and b occur by chance) pa = qaa + ½ Σa≠bqab ≈ fraction of all amino acids that are type a q The entry computed in the substitution matrix is: qab score(a, b) = log(––––––) pa pb

PAM Matrices q Based on ungapped multiple local alignments of conserved regions of proteins with high sequence identity (> 85%) q Uses phylogenetic trees to compute the entries in the substitution matrix q Procedure Ø Ø Ø build a phylogenetic tree for sequence of high identity compute relative mutability, ma, of each amino acid (frequency of a substitutions in the phylogenetic tree) compute Fab (number of substitutions of a with b) compute Mab (mutation probability that a will be replaced by b) Mab = mb Fab / Σc. Fcb compute entry in scoring matrix score(a, b) = log(Mab / frequency of a)
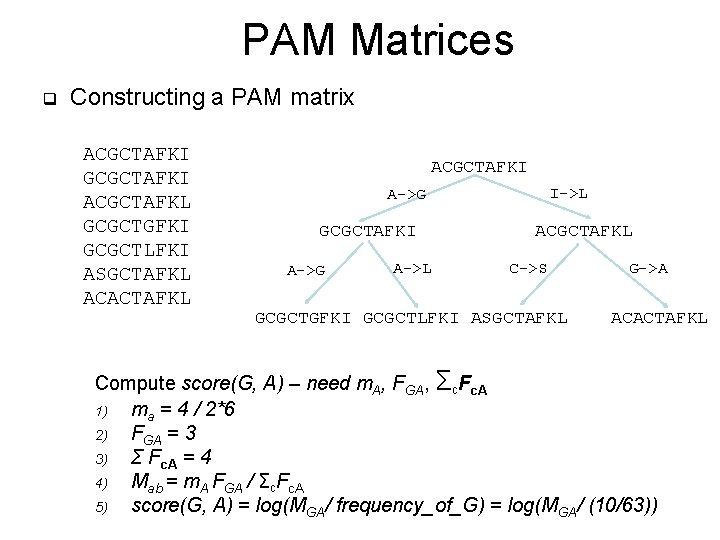
PAM Matrices q Constructing a PAM matrix ACGCTAFKI GCGCTAFKI ACGCTAFKL GCGCTGFKI GCGCTLFKI ASGCTAFKL ACACTAFKL ACGCTAFKI I->L A->G GCGCTAFKI A->G A->L ACGCTAFKL C->S GCGCTGFKI GCGCTLFKI ASGCTAFKL G->A ACACTAFKL Compute score(G, A) – need m. A, FGA, Σc. Fc. A 1) ma = 4 / 2*6 2) FGA = 3 3) Σ Fc. A = 4 4) Mab = m. A FGA / Σc. Fc. A 5) score(G, A) = log(MGA/ frequency_of_G) = log(MGA/ (10/63))
- Slides: 19