Practical Session 2 Table of contents Scoring matrices

Practical Session 2 +

Table of contents • Scoring matrices – PAM – BLOSUM • Intro to Python
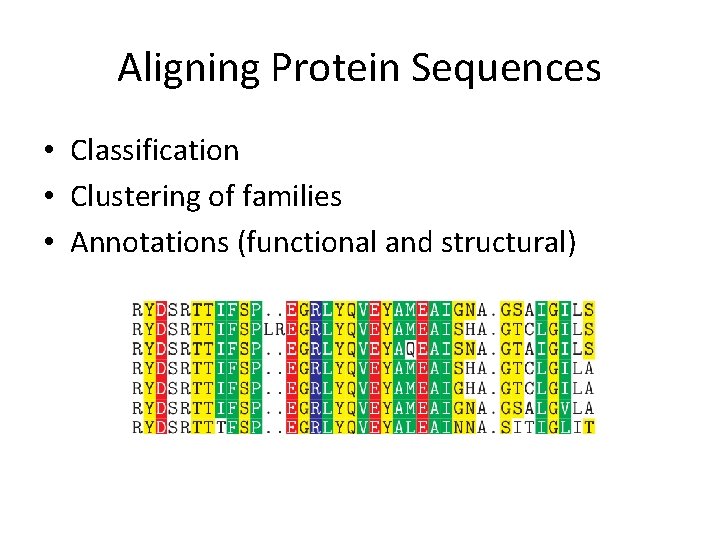
Aligning Protein Sequences • Classification • Clustering of families • Annotations (functional and structural)

Aligning Protein Sequences • Proteins consist of 20 amino acids. Task given: align two protein sequences. • Can the previous alignment algorithms be used? • How do amino acids differ from one another?

Aligning Protein Sequences When evaluating the probability of one amino acid mutating to another, we need to consider: • Mutational Distance • Chemical properties - similarity/difference • Evolutionary time

Mutational Distance Assume we start with Methionine, which is encoded by a single codon: ATG Thr (Threonine) is encoded by AC[ACGT] In order to mutate Met to Thr, one SNP (single nucleotide point) mutation is enough ACG ATG

Mutational Distance Assume we start with Methionine, which is encoded by a single codon: ATG Thr (Threonine) is encoded by AC[ACGT] In order to mutate Met to Thr, one SNP (single nucleotide point) mutation is enough • 3 point mutations are required to mutate Met to His - encoded by CA[TC] • Therefore, His is more distant to Met.
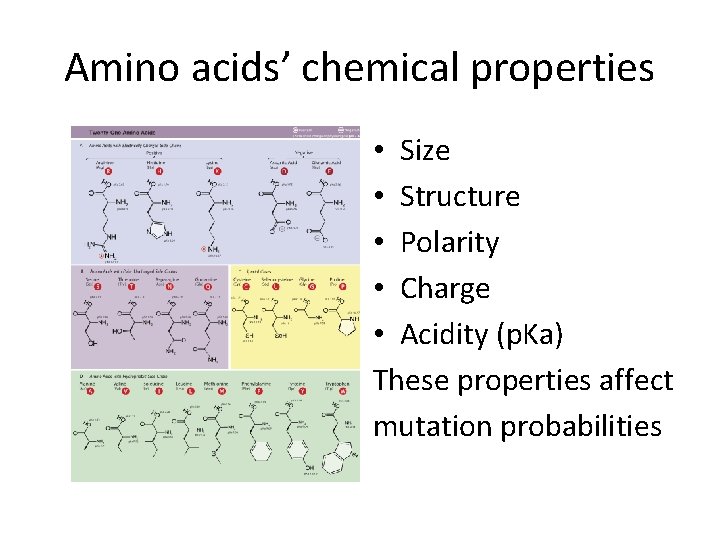
Amino acids’ chemical properties • Size • Structure • Polarity • Charge • Acidity (p. Ka) These properties affect mutation probabilities
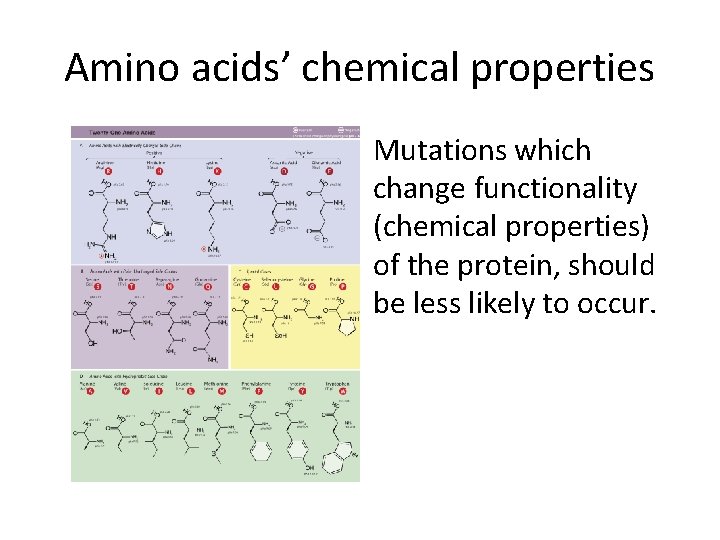
Amino acids’ chemical properties Mutations which change functionality (chemical properties) of the protein, should be less likely to occur.

Evolutionary time • Time is another aspect which needs attention. • Does longer time permits less or more mutation? • How can that be included in the scoring system ?

Evolutionary Substitution Matrix •

Evolutionary Substitution Matrix •

PAM Matrices PAM – Percent/Point Accepted Mutations. • The first widely used scoring scheme used for amino acid alignment. • Devised by Margaret Oakley Dayhoff and Co. in 1978.

PAM – point accepted mutation • Substitution of an amino acid in a protein with another amino acid, which is accepted by the process of natural selection. • Silent or lethal mutations are not point accepted mutations

PAM Matrices • PAM matrices are noted as PAMn matrices • PAM 1 represents the time period over which we expect 1% of the amino acids to undergo point accepted mutations

Constructing PAM Matrices • Examined 1572 substitutions in 71 families of proteins (71 phylogenetic trees) • The proteins sequences were at least 85% identical

Constructing PAM Matrices •

Constructing PAM Matrices
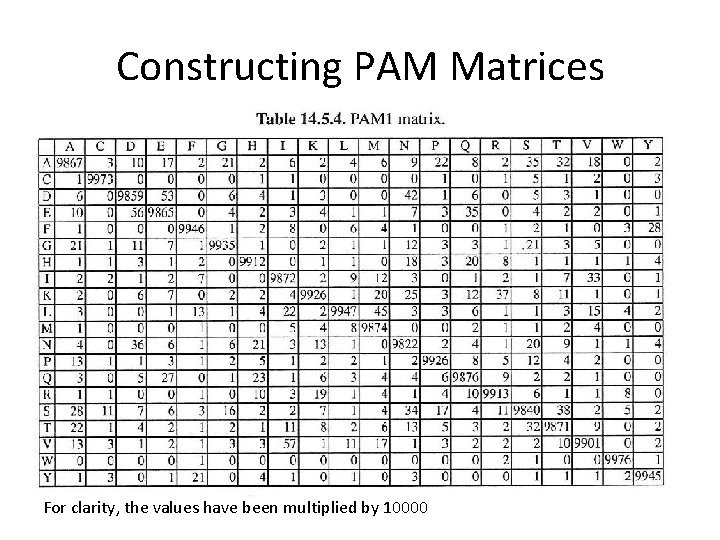
Constructing PAM Matrices For clarity, the values have been multiplied by 10000
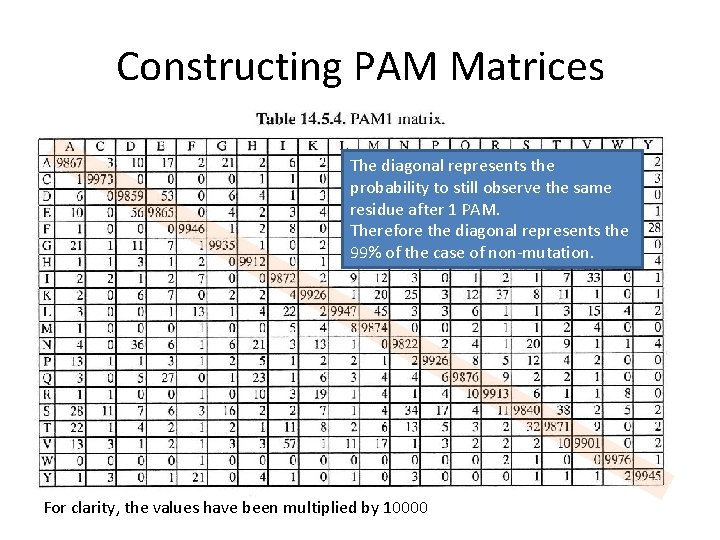
Constructing PAM Matrices The diagonal represents the probability to still observe the same residue after 1 PAM. Therefore the diagonal represents the 99% of the case of non-mutation. For clarity, the values have been multiplied by 10000

Deriving PAMn matrices •

Deriving PAMn matrices •


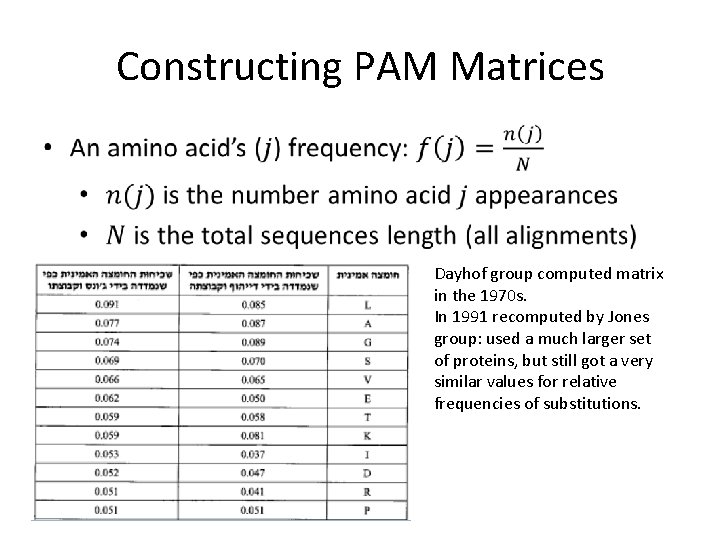
Constructing PAM Matrices Dayhof group computed matrix in the 1970 s. In 1991 recomputed by Jones group: used a much larger set of proteins, but still got a very similar values for relative frequencies of substitutions.

From probabilities to scores • Observed frequency Expected frequency by chance

Constructing PAM Matrices Observed frequency Expected frequency by chance
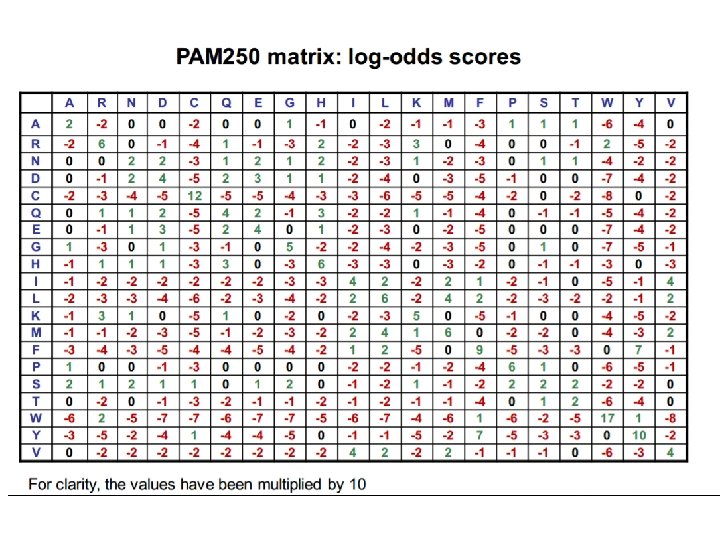

Choosing the right PAM matrix •

Choosing the right PAM matrix •

The model’s assumptions • Only mutations are allow – no indels. • Sites evolve independently – mutation in one site, has no effect on another. • Evolution model: Next mutation is dependent on current state and is independent on previous mutations.

Problem PAM matrices work quite well for closely related sequences, especially during short evolutionary time. However, they seems to lack the ability to represent more distant/divergent sequences, on a larger evolutionary time scale.

BLOSUM (BLOcks SUbstitutions Matrix) Devised by Henikoff & Henikoff in 1992.

BLOSUM (BLOcks SUbstitutions Matrix) •

BLOSUM (BLOcks SUbstitutions Matrix) • BLOSUM 62 is the default matrix for the standard protein BLAST program • BLOSUM 62 is derived from Blocks containing >62% identity in ungapped sequence alignment

Constructing BLOSUM Henikoff and Henikoff developed a database of >2, 000 blocks “blocks” based on sequences from >500 groups of related proteins with shared subsequences AABCDA. . . BBCDA DABCDA. A. BBCBB BBBCDABA. BCCAA AAACDAC. DCBCDB CCBADAB. DBBDCC AAACAA. . . BBCCC

Why blocks? • Don’t want insertions and deletions to complicate estimation of substitution probabilities • Interested in detecting conserved regions of protein sequences, so restrict attention to these regions when computing the scoring matrix

Constructing BLOSUM •
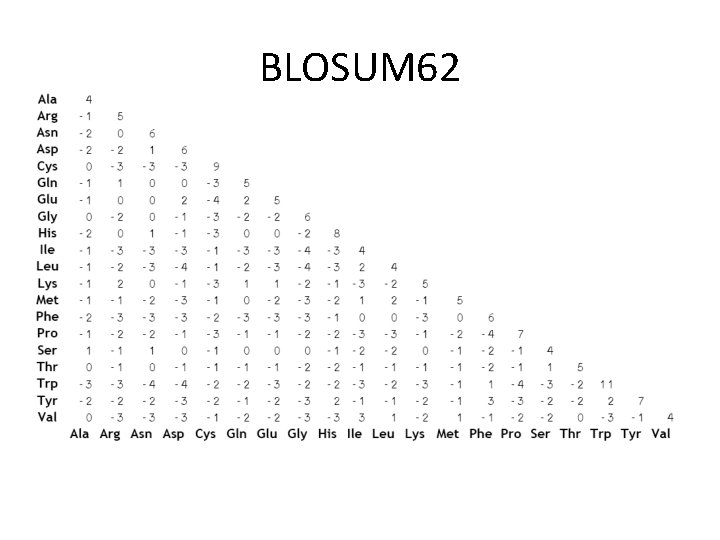
BLOSUM 62

Differences between PAM and BLOSUM

Intro to Python
Why Python? *By Code. Eval - a platform used by developers to showcase their skills.

Why Python? • • • Quick development Easy to learn Huge community Fast enough for most applications Capable of interacting with most of the other languages and platforms

![Strings http: //www. codeskulptor. org/ s = 'hi‘ print s[1] # i print len(s) Strings http: //www. codeskulptor. org/ s = 'hi‘ print s[1] # i print len(s)](http://slidetodoc.com/presentation_image/229598215dd40dba6effab175ed2d04d/image-45.jpg)
Strings http: //www. codeskulptor. org/ s = 'hi‘ print s[1] # i print len(s) # 2 print s + ' there' # hi there pi = 3. 14 text = 'The value of pi is ' + pi # does not work text = 'The value of pi is ' + str(pi) # yes s = 3
![String Slices • s[1: 4] – 'ell' -- chars starting at index 1 and String Slices • s[1: 4] – 'ell' -- chars starting at index 1 and](http://slidetodoc.com/presentation_image/229598215dd40dba6effab175ed2d04d/image-46.jpg)
String Slices • s[1: 4] – 'ell' -- chars starting at index 1 and extending up to but not including index 4 • s[1: ] – 'ello' -- omitting either index defaults to the start or end of the string • s[: ] – 'Hello' -- omitting both always gives us a copy of the whole thing (this is the pythonic way to copy a sequence like a string or list) • s[1: 100] – 'ello' -- an index that is too big is truncated down to the string length • s[-1] – 'o' -- last char (1 st from the end) • s[-3: ] – 'llo' -- starting with the 3 rd char from the end and extending to the end of the string.

If statement if speed >= 80: print 'License and registration please' Indentation is very if mood == 'terrible' or speed >= 100: important! print 'You have the right to remain silent. ' elif mood == 'bad' or speed >= 90: print "I'm going to have to write you a ticket. " write_ticket() else: print "Let's try to keep it under 80 ok? " • Note there are no {} or ;
![Lists • my_list = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10] Lists • my_list = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]](http://slidetodoc.com/presentation_image/229598215dd40dba6effab175ed2d04d/image-48.jpg)
Lists • my_list = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10] • my_list[1: 5] # [2, 3, 4, 5] • my_list[: : 2] – [1, 3, 5, 7, 9] • my_list[: : -1] – reverse [10, 9, 8, 7, 6, 5, 4, 3, 2, 1] • Lists can contain different types of variables: • pi = ['pi', 3. 14159, True]
![Lists are dynamic • students = ['Itay', 9255587, 'Alon', 744554] • students. append('Michal') # Lists are dynamic • students = ['Itay', 9255587, 'Alon', 744554] • students. append('Michal') #](http://slidetodoc.com/presentation_image/229598215dd40dba6effab175ed2d04d/image-49.jpg)
Lists are dynamic • students = ['Itay', 9255587, 'Alon', 744554] • students. append('Michal') # ['Itay', 9255587, 'Alon', 744554, 'Michal'] • students[0: 2] = [‘Noa‘] # [‘Noa’, 'Alon', 744554, 'Michal']

Range range(10) # returns an ordered list [0, 1, 2, 3, 4, 5, 6, 7, 8, 9] range(0, 10, 2) #[0, 2, 4, 6, 8] ## print the numbers from 0 through 99 for i in range(100): print i
![List iteration • squares = [1, 4, 9, 16] sum = 0 for num List iteration • squares = [1, 4, 9, 16] sum = 0 for num](http://slidetodoc.com/presentation_image/229598215dd40dba6effab175ed2d04d/image-51.jpg)
List iteration • squares = [1, 4, 9, 16] sum = 0 for num in squares: sum += num print sum ## 30

Dict ## Can build up a dict by starting with the empty dict {} ## and storing key/value pairs into the dict like this: ## dict[key] = value-for-that-key dict = {} dict['a'] = 'alpha' dict['g'] = 'gamma' dict['o'] = 'omega' print dict ## {'a': 'alpha', 'o': 'omega', 'g': 'gamma'} print dict['a'] ## Simple lookup, returns 'alpha‘ dict['a'] = 6 ## Put new key/value into dict 'a' in dict ## True print dict['z'] ## Throws Key. Error if 'z' in dict: print dict['z'] ## Avoid Key. Error print dict. get('z') ## None (instead of Key. Error)

Dict dict = {'a': 'alpha', 'o': 'omega', 'g': 'gamma'} ## By default, iterating over a dict iterates over its keys. ## Note that the keys are in a random order. for key in dict: print key ## prints a g o ## Exactly the same as above for key in dict. keys(): print key ## Get the. keys() list: print dict. keys() ## ['a', 'o', 'g'] ## Likewise, there's a. values() list of values print dict. values() ## ['alpha', 'omega', 'gamma']

Dict dict = {'a': 'alpha', 'o': 'omega', 'g': 'gamma'} ## Common case -- loop over the keys in sorted order, ## accessing each key/value for key in sorted(dict. keys()): print key, dict[key] ##. items() is the dict expressed as (key, value) tuples print dict. items() ## [('a', 'alpha'), ('o', 'omega'), ('g', 'gamma')] ## This loop syntax accesses the whole dict by looping ## over the. items() tuple list, accessing one (key, value) ## pair on each iteration. for k, v in dict. items(): print k, '>', v ## a > alpha o > omega g > gamma

Reading and Writing to a file is simple # Print the contents of a file f = open('foo. txt', 'r') for line in f: ## iterates over the lines of the file print line, ## trailing , so print does not add an end-of-line char ## since 'line' already includes the end-of line. f. close() f = open(“testfile. txt”, ”w”) f. write(“Hello World”) f. write(“This is our new text file”) f. write(“and this is another line. ”) f. write(“Why? Because we can. ”) f. close()

Functions # Function definition is here def print_info( name, age = 35 ): print "Name: ", name print "Age ", age # Now you can call printinfo function print_info( age=50, name="miki" ) Name: miki Age 50 print_info( name="miki" ) Name: miki Age 35

Classes class My. Class(object): common = 10 def __init__(self): self. my_variable = 3 def my_function(self, arg 1, arg 2): return self. my_variable # This is the class instantiation class_instance = My. Class() class_instance. my_function(1, 2) #3 class_instance 2 = My. Class() # This variable is shared by all instances class_instance. common #10 class_instance 2. common #10 My. Class. common = 30


Some tutorials • • https: //developers. google. com/edu/python/ http: //www. pythonforbeginners. com/ https: //www. codecademy. com/learn/python http: //www. learnpython. org/

Important Python Packages for the data scientist Biopython - collection of non-commercial Python tools for computational biology and bioinformatics Num. Py – mathematical package Sci. Py – scientific package Matplotlib – 2 D plotting Pandas – data structures and analysis

Development • Py. Charm - Free community version

Python 2 or Python 3? Python 2. x is legacy, Python 3. x is the present and future of the language

Main differences • Python 2 print 'Hello, World!' print('Hello, World!') print ‘text’, print 'print more text on the same line‘ • Python 3 print('Hello, World!') print("some text, ", end="") print(' print more text on the same line') Read more

Python 2 vs. Python 3 • Many prefer to use Python 2 because of larger library support • Coding style is very similar – not hard to transition from Python 2 to Python 3 • In the assignment you will use Python 2

Next practical session • Blast • Fasta
- Slides: 65