Simon Funk Netflix provided a database of 100

Simon Funk: Netflix provided a database of 100 M ratings (1 to 5) of 17 K movies by 500 K users. as a triplet of numbers: (User, Movie, Rating). The challenge: For (User, Movie, ? ) not in the database, predict how the given User would rate the given Movie. Think of the data as a big sparsely filled matrix, with user. IDs across the top and movie. IDs down the side (or vice versa then transpose everything), and each cell contains an observed rating (1 -5) for that movie (row) by that user (column), or is blank meaning you don't know. This matrix would have 8. 5 B entries, but you are only given values for 1/85 th of those 8. 5 B cells (or 100 M of them). The rest are all blank. Netflix posed a "quiz" of a bunch of question marks plopped into previously blank slots, and your job is to fill in best-guess ratings in their place. Squared error (se) measures accuracy (Your guess = 1. 5, actual = 2, you get docked for (2 -1. 5)^2 or. 25. They use root mean squared error ( rmse), but rmse and mse monotonically related. ) There is a date for ratings and question marks (so a cell can potentially have >=1 rating in it. Any movie can be described in terms of some aspects or attributes such as overall quality, action(y/n? ), comedy(y/n? ), stars, producer, etc. Every user's preferences can be roughly described in terms of whether they tend to rate quality/action/comedy/star/producer/etc. high or low. If true, then ratings ought to be explainable by a lot less than 8. 5 billion numbers (e. g. , a single number specifying how much action a particular movie has may help explain why a few million action-buffs like that movie. ) SVD assumes rating(u, m) is sum of preferences about the various aspects. E. g. , take 40 aspects - a movie, m, is described by 40 values, m(f), saying how much that movie exemplifies that aspect, and a user is described by 40 values, u(f), saying how much they prefer each aspect. A rating(u, m) = u(f) dot m(f) (40*(17 K+500 K) values =~20 M << 8. 5 B. ) or: ratings. Matrix[user][movie] = sum (user. Feature[f][user] * movie. Feature[f][movie]) for f from 1 to 40 (or 1 to F in general) R = UT o I ru, i = u o i = f=1. . F ru, f * rf, i The original matrix has been decomposed to 2 oblong matrices: 17 Kx 40 movie aspect matrix, 500 Kx 40 user preference matrix. SVD is a trick for finding the 2 smaller matrices which minimize the R i 1 i i. Test. Size. I UT f 1 f f. F I i i i 1 Test. Size. I resulting approx error--specifically the mean squared error. f 1 u 1 So, if we take the rank=40 SVD of the 8. 5 B matrix, we have the best o f rf, i (least error) approx we can within the limits of our user-movie. : f. F rating model. I. e. , the SVD has found "best" generalizations. u ru, i = u ru, f Take the derivative of the approx error and follow it. This has the bonus . : that we can ignore the unknown error on the 8. 4 B empty slots. Take the derivative of the equations for the error--just the given values, not the empties--with respect to the parameters: u. Test. Size. U user. Value[user] += lrate*err*movie. Value[movie]; movie. Value[movie] += lrate*err*user. Value[user]; With Horizontal data, the code is evaluated for each rating. So, to train for one sample: real *user. Value= user. Feature[feature. Being. Trained]; real *movie. Value= movie. Feature[feature. Being. Trained]; real lrate = 0. 001; More correctly: ru, f = uv = user. Value[user] += err * movie. Value[movie]; rf, i = movie. Value[movie] += err * uv; finds the most prominent feature remaining (most reduces error). When it's good, shift it onto done features, start a new one (cache residuals of the 100 M. "What does that mean for us? ? ? ). This Gradient descent has no local minima, which means it doesn't really matter how it's initialized. u+ = lrate ( u, i * i. T - * u ) where u, i = ru, i - r^u, i and r^u, i = actual rating

Refinements: Prior to starting SVD, Note: Avg. Rating(movie), Avg. Offset(User. Rating, Movie. Avg. Rating), for every user. I. e. : static inline real predict. Rating_Baseline(int movie, int user) {return average. Rating[movie] + average. Offset[user]; } So, that's the return value of predict. Rating before the first SVD feature even starts training. You'd think avg rating for a movie would just be. . . its average rating! Alas, Occam's razor was a little rusty that day. If m only appears once with r(m, u)=1 say, Avg. Rating(m)=1? Probably not! View r(m, u)=1 as a draw from a true prob dist who's avg you want. . . View that true average itself as a draw from a prob dist of averages--the histogram of average movie ratings. Assume both distributions Gaussian, then the best-guess mean should be lin combo of observed mean and apriori mean, with a blending ratio equal to the ratio of variances. If Ra and Va are the mean and variance (squared standard deviation) of all of the movies' average ratings (which defines your prior expectation for a new movie's average rating before you've observed any actual ratings) and Vb is the average variance of individual movie ratings (which tells you how indicative each new observation is of the true mean-- e. g, . if the average variance is low, then ratings tend to be near the movie's true mean, whereas if the avg variance is high, ratings tend to be more random and less indicative) then: Bogus. Mean = sum(Observed. Ratings)/count(Observed. Ratings) K = Vb/Va Better. Mean = [Global. Average*K + sum(Observed. Ratings)] / [K + count(Observed. Ratings)] The point here is simply that any time you're averaging a small number of examples, the true average is most likely nearer the apriori average than the sparsely observed average. Note if the number of observed ratings for a particular movie is zero, the Better. Mean (best guess) above defaults to the global average movie rating as one would expect. Moving on: 20 M free params is a lot for a 100 M Train. Set. Seems neat to just ignore all blanks, but we have expectations about them. As-is, this modified SVD algorithm tends to make a mess of sparsely observed movies or users. If you have a user who has only rated 1 movie, say American Beauty=2 while the avg is 4. 5, and further that their offset is only -1, we'd, prior to SVD, expect them to rate it 3. 5. So the error given to the SVD is -1. 5 (the true rating is 1. 5 less than we expect). m(Action) is training up to measure the amount of Action, say, . 01 for American Beauty ( ust slightly more than avg). SVD optimize predictions, which it can do by eventually setting our user's preference for Action to a huge -150. I. e. , the alg naively looks at the only example it has of this user's preferences and in the context of only the one feature it knows about so far (Action), determines that our user so hates action movies that even the tiniest bit of action in American Beauty makes it suck a lot more than it otherwise might. This is not a problem for users we have lots of observations for because those random apparent correlations average out and the true trends dominate. We need to account for priors. As with the average movie ratings, blend our sparse observations in with some sort of prior, but it's a little less clear how to do that with this incremental algorithm. But if you look at where the incremental algorithm theoretically converges, you get: user. Value[user] = [sum residual[user, movie]*movie. Value[movie]] / [sum (movie. Value[movie]^2)] The numerator there will fall in a roughly zero-mean Gaussian distribution when charted over all users, which through various gyrations: user. Value[user] = [sum residual[user, movie]*movie. Value[movie]] / [sum (movie. Value[movie]^2 + K)] And finally back to: user. Value[user] += lrate * (err * movie. Value[movie] - K * user. Value[user]); movie. Value[movie] += lrate * (err * user. Value[user] - K * movie. Value[movie]); This is equivalent to penalizing the magnitude of the features. To cut over fitting, allowing use of more features.

Moving on: Linear models are limiting. We've bastardized the whole matrix analogy so much that we aren't really restricted to linear models: We can add non-linear outputs such that instead of predicting with: sum (user. Feature[f][user] * movie. Feature[f][movie]) for f from 1 to 40. We can use: sum G(user. Feature[f][user] * movie. Feature[f][movie]) for f from 1 to 40. Two choices for G proved useful. 1. clip the prediction to 1 -5 after each component is added. E. g. , each feature is limited to only swaying rating within the valid range, and any excess beyond that is lost rather than carried over. So, if the first feature suggests +10 on a scale of 1 -5, and the second feature suggests -1, then instead of getting a 5 for the final clipped score, it gets a 4 because the score was clipped after each stage. The intuitive rationale here is that we tend to reserve the top of our scale for the perfect movie, and the bottom for one with no redeeming qualities whatsoever, and so there's a sort of measuring back from the edges that we do with each aspect independently. More pragmatically, since the target range has a known limit, clipping is guaranteed to improve our perf, and having trained a stage with clipping on, use it with clipping on. I did not really play with this extensively enough to determine there wasn't a better strategy. A second choice for G is to introduce some functional non-linearity such as a sigmoid. I. e. , G(x) = sigmoid(x). Even if G is fixed, this requires modifying the learning rule slightly to include the slope of G, but that's straightforward. The next question is how to adapt G to the data. I tried a couple of options, including an adaptive sigmoid, but the most general and the one that worked the best was to simply fit a piecewise linear approximation to the true output/output curve. That is, if you plot the true output of a given stage vs the average target output, the linear model assumes this is a nice 45 degree line. But in truth, for the first feature for instance, you end up with a kink around the origin such that the impact of negative values is greater than the impact of positive ones. That is, for two groups of users with opposite preferences, each side tends to penalize more strongly than the other side rewards for the same quality. Or put another way, below-average quality (subjective) hurts more than above-average quality helps. There is also a bit of a sigmoid to the natural data beyond just what is accounted for by the clipping. The linear model can't account for these, so it just finds a middle compromise; but even at this compromise, the inherent non-linearity shows through in an actual-output vs. average-target-output plot, and if G is then simply set to fit this, the model can further adapt with this new performance edge, which leads to potentially more beneficial nonlinearity and so on. . . This introduces new free parameters and encourages over fitting especially for the later features which tend to represent small groups. We found it beneficial to use this non-linearity only for the first twenty or so features and to disable it after that. Moving on: Despite the regularization term in the final incremental law above, over fitting remains a problem. Plotting the progress over time, the probe rmse eventually turns upward and starts getting worse (even though the training error is still inching down). We found that simply choosing a fixed number of training epochs appropriate to the learning rate and regularization constant resulted in the best overall performance. I think for the numbers mentioned above it was about 120 epochs per feature, at which point the feature was considered done and we moved on to the next before it started over fitting. Note that now it does matter how you initialize the vectors: Since we're stopping the path before it gets to the (common) end, where we started will affect where we are at that point. I wonder if a better regularization couldn't eliminate overfitting altogether, something like Dirichlet priors in an EM approach--but I tried that and a few others and none worked as well as the above. Here is the probe and training rmse for the first few features with and w/o regularization term "decay" enabled. Same thing, just the probe set rmse, further along where you can see the regularized version pulling ahead: This time showing probe rmse (vertical) against train rmse (horizontal). Note how the regularized version has better probe performance relative to the training performance: Anyway, that's about it. I've tried a few other ideas over the last couple of weeks, including a couple of ways of using the date information, and while many of them have worked well up front, none held their advantage long enough to actually improve the final result. If you notice any obvious errors or have reasonably quick suggestions for better notation or whatnot to make this explanation more clear, let me know. And of course, I'd love to hear what y'all are doing and how well it's working, whether it's improvements to the above or something completely different. Whatever you're willing to share,

//============================ // SVD Sample Code (C) 2007 Timely Development (www. timelydevelopment. com) // STANDARD DISCLAIMER: // - THIS CODE AND INFORMATION IS PROVIDED "AS IS" WITHOUT WARRANTY // - OF ANY KIND, EITHER EXPRESSED OR IMPLIED, INCLUDING BUT NOT // - LIMITED TO THE IMPLIED WARRANTIES OF MERCHANTABILITY AND/OR // - FITNESS FOR A PARTICULAR PURPOSE. //========================== #define WIN 32_LEAN_AND_MEAN #include <windows. h> #include <stdio. h> #include <math. h> #include <tchar. h> #include <map> using namespace std; //======================== // Constants and Type Declarations //======================== #define TRAINING_PATH L"C: etflix raining_set*. txt" #define TRAINING_FILE L"C: etflix raining_set%s" #define FEATURE_FILE L"C: etflixfeatures. txt" #define TEST_PATH L"C: etflix%s" #define PREDICTION_FILE L"C: etflixprediction. txt" #define MAX_RATINGS 00480508 //Ratings in entire training set (+1) #define MAX_CUSTOMERS 480190 //Custs in entire training set (+1) #define MAX_MOVIES 17771 //Movies in entire training set (+1) #define MAX_FEATURES 64 //Number of features to use #define MIN_EPOCHS 120 //Min number of epochs per feature #define MAX_EPOCHS 200 // Max epochs per feature #define MIN_IMPROVEMENT 0. 0001 //Min improve req cont current feature #define INIT 0. 1 // Initialization value for features #define LRATE 0. 001 // Learning rate parameter #define K 0. 015 // Reg param to min over-fitting typedef unsigned char BYTE; typedef map<int, int> Id. Map; typedef Id. Map: : iterator Id. Itr; struct Movie { int Rating. Count; int Rating. Sum; double Rating. Avg; double Pseudo. Avg; //Wtd avg to deal with small movie counts }; struct Customer { int Customer. Id; int Rating. Count; int Rating. Sum; }; struct Data { int Cust. Id; short Movie. Id; BYTE Rating; float Cache; }; class Engine { private: int m_n. Rating. Count; // Current number of loaded ratings Data m_a. Ratings[MAX_RATINGS]; //Array of ratings data Movie m_a. Movies[MAX_MOVIES]; //Array of movie metrics Customer m_a. Customers[MAX_CUSTOMERS]; //Array of customer metrics float m_a. Movie. Features[MAX_FEATURES][MAX_MOVIES]; //Array of feat by mov float m_a. Cust. Features[MAX_FEATURES][MAX_CUSTOMERS]; //Array feas by cust Id. Map m_m. Cust. Ids; // Map for one time translation of ids to compact array index inline double Predict. Rating(short movie. Id, int cust. Id, int feature, float cache, bool b. Trailing=true); inline double Predict. Rating(short movie. Id, int cust. Id); bool Read. Number(wchar_t* pwz. Buffer. In, int n. Length, int &n. Position, wchar_t* pwz. Buffer. Out); bool Parse. Int(wchar_t* pwz. Buffer, int n. Length, int &n. Position, int& n. Value); bool Parse. Float(wchar_t*pwz. Buffer, int n. Length, int &n. Position, float& f. Value); public: Engine(void); ~Engine(void) { }; void Calc. Metrics(); void Calc. Features(); void Load. History(); void Process. Test(wchar_t* pwz. File); void Process. File(wchar_t* pwz. File); }; //======= // Program Main int _tmain(int argc, _TCHAR* argv[]) { Engine* engine = new Engine(); engine->Load. History(); engine->Calc. Metrics(); engine->Calc. Features(); engine->Process. Test(L"qualifying. txt"); wprintf(L" Done "); getchar(); return 0; }

//=================== // Engine Class // Initialization Engine: : Engine(void) { m_n. Rating. Count = 0; for (int f=0; f<MAX_FEATURES; f++) { for (int i=0; i<MAX_MOVIES; i++) m_a. Movie. Features[f][i] = (float)INIT; for (int i=0; i<MAX_CUSTOMERS; i++) m_a. Cust. Features[f][i] = (float)INIT; } } //---------------------// Calculations - This Paragraph contains all of the relevant code // Calc. Metrics // - Loop through the history and pre-calculate metrics used in the training // - Also re-number the customer id's to fit in a fixed array void Engine: : Calc. Metrics() { int i, cid; Id. Itr itr; wprintf(L" Calculating intermediate metrics "); // Process each row in the training set for (i=0; i<m_n. Rating. Count; i++) { Data* rating = m_a. Ratings + i; // Increment movie stats m_a. Movies[rating->Movie. Id]. Rating. Count++; m_a. Movies[rating->Movie. Id]. Rating. Sum += rating->Rating; // Add customers (using a map to re-number id's to array indexes) itr = m_m. Cust. Ids. find(rating->Cust. Id); if (itr == m_m. Cust. Ids. end()) { cid = 1 + (int)m_m. Cust. Ids. size(); // Reserve new id and add lookup m_m. Cust. Ids[rating->Cust. Id] = cid; // Store off old sparse id for later m_a. Customers[cid]. Customer. Id = rating->Cust. Id; // Init vars to zero m_a. Customers[cid]. Rating. Count = 0; m_a. Customers[cid]. Rating. Sum = 0; } else { cid = itr->second; // Swap sparse id for compact one rating->Cust. Id = cid; m_a. Customers[cid]. Rating. Count++; m_a. Customers[cid]. Rating. Sum += rating->Rating; } // Do a follow-up loop to calc movie averages for (i=0; i<MAX_MOVIES; i++) { Movie* movie = m_a. Movies+i; movie->Rating. Avg = movie->Rating. Sum / (1. 0 * movie->Rating. Count); movie->Pseudo. Avg = (3. 23 * 25 + movie->Rating. Sum) / (25. 0 + movie->Rating. Count); } } // Calc. Features - Iteratively train each feature on entire data set // - Once sufficient progress has been made, move on void Engine: : Calc. Features() { int f, e, i, cust. Id, cnt = 0; Data* rating; double err, p, sq, rmse_last, rmse = 2. 0; short movie. Id; float cf, mf; for (f=0; f<MAX_FEATURES; f++) { wprintf(L" --- Calculating feature: %d --- ", f); // Keep looping until you have passed a minimum number // of epochs or have stopped making significant progress for (e=0; (e < MIN_EPOCHS) || (rmse <= rmse_last - MIN_IMPROVEMENT); e++) { cnt++; sq = 0; rmse_last = rmse; for (i=0; i<m_n. Rating. Count; i++) { rating = m_a. Ratings + i; movie. Id = rating->Movie. Id; cust. Id = rating->Cust. Id; // Predict rating and calc error p = Predict. Rating(movie. Id, cust. Id, f, rating->Cache, true); err = (1. 0 * rating->Rating - p); sq += err*err; // Cache off old feature values cf = m_a. Cust. Features[f][cust. Id]; mf = m_a. Movie. Features[f][movie. Id]; // Cross-train the features m_a. Cust. Features[f][cust. Id] += (float)(LRATE * (err * mf - K * cf)); m_a. Movie. Features[f][movie. Id] += (float)(LRATE * (err * cf - K * mf)); } rmse = sqrt(sq/m_n. Rating. Count); wprintf(L" <set x='%d' y='%f' /> ", cnt, rmse); }

// Cache off old predictions for (i=0; i<m_n. Rating. Count; i++) { rating = m_a. Ratings + i; rating->Cache = (float)Predict. Rating(rating->Movie. Id, rating->Cust. Id, f, rating->Cache, false); } } // Predict. Rating - During training there is no need to loop through all of the features // - Use a cache for the leading features and do a quick calculation for the trailing // - The trailing can be optionally removed when calculating a new cache value double Engine: : Predict. Rating(short movie. Id, int cust. Id, int feature, float cache, bool b. Trailing) { // Get cached value for old features or default to an average double sum = (cache > 0) ? cache : 1; //m_a. Movies[movie. Id]. Pseudo. Avg; //Add contribution of current feature sum += m_a. Movie. Features[feature][movie. Id] * m_a. Cust. Features[feature][cust. Id]; if (sum > 5) sum = 5; if (sum < 1) sum = 1; // Add up trailing defaults values if (b. Trailing) { sum += (MAX_FEATURES-feature-1) * (INIT * INIT); if (sum > 5) sum = 5; if (sum < 1) sum = 1; } return sum; } // Predict. Rating - This version is used for calculating the final results // - It loops through the entire list of finished features double Engine: : Predict. Rating(short movie. Id, int cust. Id) { double sum = 1; //m_a. Movies[movie. Id]. Pseudo. Avg; for (int f=0; f<MAX_FEATURES; f++) { sum += m_a. Movie. Features[f][movie. Id] * m_a. Cust. Features[f][cust. Id]; if (sum > 5) sum = 5; if (sum < 1) sum = 1; } return sum; } // Data Loading / Saving // Load. History // - Loop through all of the files in the training directory void Engine: : Load. History() { WIN 32_FIND_DATA Find. File. Data; HANDLE h. Find; bool b. Continue = true; // Loop through all of the files in the training directory h. Find = Find. First. File(TRAINING_PATH, &Find. File. Data); if (h. Find == INVALID_HANDLE_VALUE) return; while (b. Continue) { this->Process. File(Find. File. Data. c. File. Name); b. Continue = (Find. Next. File(h. Find, &Find. File. Data) != 0); //if (++count > 999) break; // TEST: Uncomment to only test with the first X movies } Find. Close(h. Find); } // Process. File Load a history: <Movie. Id>: <Customer. Id>, <Rating>. . . void Engine: : Process. File(wchar_t* pwz. File) { FILE *stream; wchar_t pwz. Buffer[1000]; wsprintf(pwz. Buffer, TRAINING_FILE, pwz. File); int cust. Id, movie. Id, rating, pos = 0; wprintf(L"Processing file: %s ", pwz. Buffer); if (_wfopen_s(&stream, pwz. Buffer, L"r") != 0) return; // First line is the movie id fgetws(pwz. Buffer, 1000, stream); Parse. Int(pwz. Buffer, (int)wcslen(pwz. Buffer), pos, movie. Id); m_a. Movies[movie. Id]. Rating. Count = 0; m_a. Movies[movie. Id]. Rating. Sum = 0; // Get all remaining rows fgetws(pwz. Buffer, 1000, stream); while ( !feof( stream ) ) { pos = 0; Parse. Int(pwz. Buffer, (int)wcslen(pwz. Buffer), pos, cust. Id); Parse. Int(pwz. Buffer, (int)wcslen(pwz. Buffer), pos, rating); m_a. Ratings[m_n. Rating. Count]. Movie. Id = (short)movie. Id; m_a. Ratings[m_n. Rating. Count]. Cust. Id = cust. Id; m_a. Ratings[m_n. Rating. Count]. Rating = (BYTE)rating; m_a. Ratings[m_n. Rating. Count]. Cache = 0; m_n. Rating. Count++; fgetws(pwz. Buffer, 1000, stream); } // Cleanup fclose( stream ); } // Process. Test - Load a sample set in the following format // <Movie 1 Id>: <Customer. Id>. . . <Movie 2 Id>: <Customer. Id> // - And write results: <Movie 1 Id>: <Rating> <Raing> . . . void Engine: : Process. Test(wchar_t* pwz. File) { FILE *stream. In, *stream. Out; wchar_t pwz. Buffer[1000]; int cust. Id, movie. Id, pos = 0; double rating; bool b. Movie. Row; wsprintf(pwz. Buffer, TEST_PATH, pwz. File);

Processing test: %s ", pwz. Buffer); // Find start of number if (_wfopen_s(&stream. In, pwz. Buffer, L"r") != 0) return; while (start < n. Length) if (_wfopen_s(&stream. Out, PREDICTION_FILE, L"w") != 0) return; { wc = pwz. Buffer. In[start]; fgetws(pwz. Buffer, 1000, stream. In); if ((wc >= 48 && wc <= 57) || (wc == 45)) break; while ( !feof( stream. In ) ) start++; { } b. Movie. Row = false; for (int i=0; i<(int)wcslen(pwz. Buffer); i++) // Copy each character into the output buffer { n. Position = start; b. Movie. Row |= (pwz. Buffer[i] == 58); while(n. Position<n. Length&&((wc>=48&&wc<=57)||wc==69 || wc==101 || wc==45 || wc==46)) } { pwz. Buffer. Out[count++] = wc; pos = 0; wc = pwz. Buffer. In[++n. Position]; if (b. Movie. Row) } { Parse. Int(pwz. Buffer, (int)wcslen(pwz. Buffer), pos, movie. Id); // Null terminate and return // Write same row to results pwz. Buffer. Out[count] = 0; fputws(pwz. Buffer, stream. Out); return (count > 0); } } else { bool Engine: : Parse. Float(wchar_t* pwz. Buffer, int n. Length, int &n. Position, float& f. Value) Parse. Int(pwz. Buffer, (int)wcslen(pwz. Buffer), pos, cust. Id); { cust. Id = m_m. Cust. Ids[cust. Id]; wchar_t pwz. Number[20]; rating = Predict. Rating(movie. Id, cust. Id); bool b. Result = Read. Number(pwz. Buffer, n. Length, n. Position, pwz. Number); f. Value = (b. Result) ? (float)_wtof(pwz. Number) : 0; // Write predicted value return false; swprintf(pwz. Buffer, 1000, L"%5. 3 f ", rating); } fputws(pwz. Buffer, stream. Out); } bool Engine: : Parse. Int(wchar_t* pwz. Buffer, int n. Length, int &n. Position, int& n. Value) { //wprintf(L"Got Line: %d %d %d ", movie. Id, cust. Id, rating); wchar_t pwz. Number[20]; fgetws(pwz. Buffer, 1000, stream. In); bool b. Result = Read. Number(pwz. Buffer, n. Length, n. Position, pwz. Number); } n. Value = (b. Result) ? _wtoi(pwz. Number) : 0; return b. Result; // Cleanup } fclose( stream. In ); fclose( stream. Out ); } //--------------------// Helper Functions //--------------------bool Engine: : Read. Number(wchar_t* pwz. Buffer. In, int n. Length, int &n. Position, wchar_t* pwz. Buffer. Out) { int count = 0; int start = n. Position; wchar_t wc = 0;

Maximizing the. Variance Given any table, X(X 1, . . . , Xn), and any unit vector, d, in n-space, let Xod=Fd(X)=DPP d(X) x 1 x 2 : x. N d 1 dn x 1 od x 2 od =x Nod How do we use this theory? For Dot Product gap based Clustering, we can hill-climb akk below to a d that gives us the global maximum variance. Heuristically, higher variance means more prominent gaps. M 1 For Dot Product Gap based Classification, we can start M 2 V(d)≡Variance. Xod=(Xod)2 - (Xod)2 with X = the table of the C Training Set Class Means, : 2 1 where Mk≡Mean. Vector. Of. Classk. MC ( Xj dj )2 = ( j=1. . n xi, jdj) - j=1. . n N i=1. . N Then Xi = Mean(X)i and These computations are O(C) (C=number of 1 ( x d ) = - ( X d ) ( X d ) ( x d ) i, j j k k i, k k k N i j and Xi. Xj = Mean Mi 1 Mj 1 classes) and are instantaneous. Once we have j k 2 x x d d 2 2 1 x 2 d 2 . the matrix A, we can hill-climb to obtain a d + = N i j i, j j N j<k i, j i, k j k - j. Xj dj +2 j<k Xj. Xkdjdk FAUST Classifier MVDI : that maximizes the variance of the dot product (Maximized Variance Mi. C Mj. C +2 j<k. Xj. Xkdjdk - " = Xj 2 dj 2 projections of the class means. Definite Indefinite: j = +(2 j=1. . n<k=1. . n (Xj. Xk - (Xj 2 - Xj 2)dj 2 + Xj. Xk)djdk ) Build a Decision tree. 1. Find d that maximizes variance of dot product subject to i=1. . ndi 2=1 projections of class means each round. 2. Apply DI each round d. T o A o d = V(d) FAUST technology relies on: 1. a distance dominating functional, F. d 1 V d 1 . . . dn We can separate out the diagonal : 2. Use of gaps in range(F) to separate. or not: dn For Unsupervised (Clustering) Hierarchical Divisive? Piecewise Linear? i Xi. Xj-Xi. X, j V(d)= jajjdj 2 + j kajkdjdk other? Perf Anal (which approach is best for which type of table? ) : V(d) = ijaijdidj For Supervised (Classification), Decision Tree? Nearest Nbr? Piecewise Linear? Perf Anal (which is best for training set? ) d 1≡ (V(d 0)); d 0, one can hill-climb it to locally maximize the variance, V, as follows: d 2≡ (V(d 1)): . . . where White papers: Terabyte Head Wall. The Only Good Data is Data in Motion V(d)≡Gradient(V)=2 Aod Multilevel p. Trees: k=0, 1 suffices! A PTree. Set is defined by specifying a table, ora d V(d)= 2 a 11 d 1 + j 1 1 j j an array of stride_lengths (usually equi-length so just that one length is specified) 2 a 22 d 2 + j 2 a 2 jdj 2 a 11 2 a 12. . . 2 a 1 n d 1 and a stride_predicate (TF condition on a stride (stride=bag [or array? ] of bits): 2 a 21 2 a 22. . . 2 a 2 n : : So the metadata of PTree. Set(T, sl, sp) specifies T, sl and sp. : d i A “raw” PTree. Set has sl=1 and the identity predicate (sl and sp not used). 2 anndn + j nanjdj ' : A “cooked” PTree. Set (AKA Level-1 PTree. Set) for a table with sl 1 Ubhaya Theorem 1: 2 an 1 . . . 2 ann dn (main purpose: provide compact summary information on the table. ) Let PTS(T) be a raw PTree. Set, then it, plus PTS(T, 64, p), . . . , PTS(T, 64^k, p) form a k {1, . . . , n} s. t. d=ek will hill-climb V to its globally max. tree of vertical summarizations of T. Theorem 2 (working on it): Note that P(T, 64*64, p) is different from P(P(T, 64, p), but both make sense Let d=ek s. t. akk is a maximal diagonal element of A, since P(t, 64, p) is a table and P(P(T, 64, p) is just a cooked p. Tree on it. d=ek will hill-climb V to its globally maximum. j=1. . n
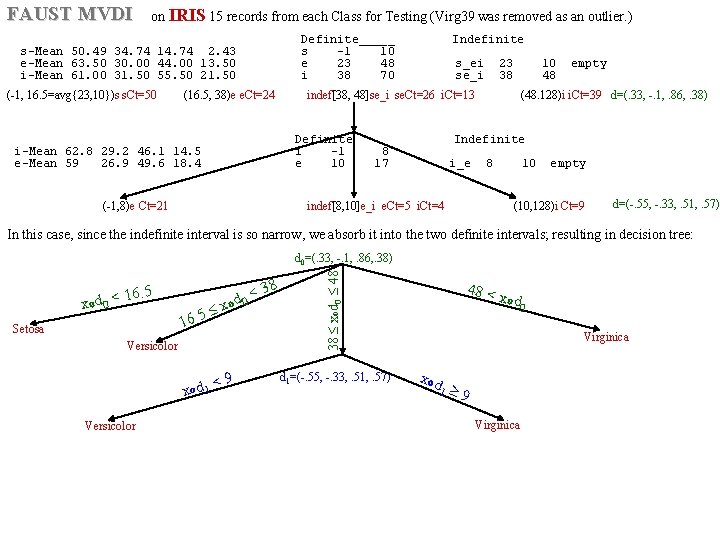
FAUST MVDI on IRIS 15 records from each Class for Testing (Virg 39 was removed as an outlier. ) Definite_____ Indefinite s-Mean 50. 49 34. 74 14. 74 2. 43 s -1 10 e-Mean 63. 50 30. 00 44. 00 13. 50 e 23 48 s_ei 23 10 empty i-Mean 61. 00 31. 50 55. 50 21. 50 i 38 70 se_i 38 48 (-1, 16. 5=avg{23, 10})s s. Ct=50 (16. 5, 38)e e. Ct=24 (48. 128)i i. Ct=39 d=(. 33, -. 1, . 86, . 38) indef[38, 48]se_i se. Ct=26 i. Ct=13 Definite Indefinite i-Mean 62. 8 29. 2 46. 1 14. 5 i -1 8 e-Mean 59 26. 9 49. 6 18. 4 e 10 17 i_e 8 10 empty (-1, 8)e Ct=21 (10, 128)i Ct=9 indef[8, 10]e_i e. Ct=5 i. Ct=4 d=(-. 55, -. 33, . 51, . 57) In this case, since the indefinite interval is so narrow, we absorb it into the two definite intervals; resulting in decision tree: 16. 5 xod 0 < Setosa 38 < od 0 x 5. 16 Versicolor < 9 xod 1 Versicolor 38 xod 0 48 d 0=(. 33, -. 1, . 86, . 38) d 1=(-. 55, -. 33, . 51, . 57) 48 < x od 0 Virginica xod 1 9 Virginica
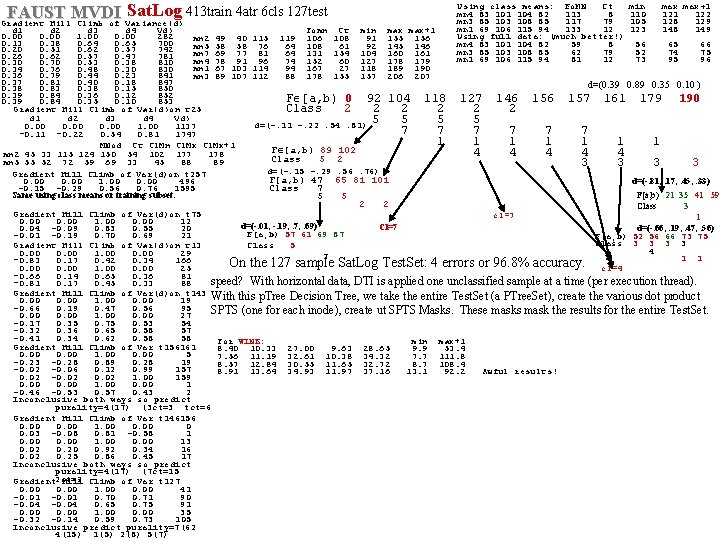
FAUST MVDI Sat. Log 413 train 4 atr 6 cls 127 test Gradient Hill Climb of Variance(d) Using class means: Fo. MN Ct mn 4 83 101 104 82 113 8 mn 3 85 103 108 85 117 79 mn 1 69 106 115 94 133 12 Using full data: (much better!) mn 4 83 101 104 82 59 8 mn 3 85 103 108 85 62 79 mn 1 69 106 115 94 81 12 min 110 105 123 max+1 122 128 129 148 149 d 1 d 2 d 3 d 4 Vd) Fomn Ct min max+1 0. 00 1. 00 0. 00 282 mn 2 49 40 115 119 106 108 91 155 156 0. 13 0. 38 0. 64 0. 65 700 56 65 66 mn 5 58 58 76 64 108 61 92 145 146 0. 20 0. 51 0. 62 0. 57 742 52 74 75 mn 7 69 77 81 64 131 154 104 160 161 0. 26 0. 62 0. 57 0. 47 781 73 95 96 mn 4 78 91 96 74 152 60 127 178 179 0. 30 0. 70 0. 53 0. 38 810 mn 1 67 103 114 94 167 27 118 189 190 0. 34 0. 76 0. 48 0. 30 830 0. 36 0. 79 0. 44 0. 23 841 mn 3 89 107 112 88 178 155 157 206 207 0. 37 0. 81 0. 40 0. 18 847 d=(0. 39 0. 89 0. 35 0. 10 ) 0. 38 0. 83 0. 38 0. 15 850 0. 39 0. 84 0. 36 0. 12 852 F [a, b) 0 92 104 118 127 146 157 161 179 190 0. 39 0. 84 0. 35 0. 10 853 Class 2 2 2 Gradient Hill Climb of Var(d)on t 25 d 1 d 2 d 3 d 4 Vd) 5 5 d=(-. 11 -. 22. 54. 81) 0. 00 1137 7 7 7 -0. 11 -0. 22 0. 54 0. 81 1747 1 1 MNod Ct Cl. Mn Cl. Mx+1 F [a, b) 89 102 4 4 4 mn 2 45 33 115 124 150 54 102 177 178 Class 5 2 mn 5 55 52 72 59 69 33 45 88 89 3 3 3 3 d=(-. 15 -. 29. 56. 76) Gradient Hill Climb of Var(d)on t 257 F[a, b) 47 65 81 101 d=(-. 81, . 17, . 45, . 33) 0. 00 1. 00 0. 00 496 -0. 15 -0. 29 0. 56 0. 76 1595 Class 7 F[a, b) 21 35 41 59 Same using class means or training subset. 5 5 2 2 Class 3 Gradient Hill Climb of Var(d)on t 75 cl=7 1 0. 00 1. 00 0. 00 12 d=(-. 01, -. 19, . 7, . 69) Cl=7 d=(-. 66, . 19, . 47, . 56) 0. 04 -0. 09 0. 83 0. 55 20 F[a, b) 57 61 69 87 -0. 01 -0. 19 0. 70 0. 69 21 F[a, b) 52 56 66 73 75 Class 3 3 Class 5 Gradient Hill Climb of Var(d)on t 13 4 0. 00 1. 00 0. 00 29 7 1 1 -0. 83 0. 17 0. 42 0. 34 166 On the 127 sample Sat. Log Test. Set: 4 errors or 96. 8% accuracy. 0. 00 1. 00 0. 00 25 cl=4 -0. 66 0. 14 0. 65 0. 36 81 speed? With horizontal data, DTI is applied one unclassified sample at a time (per execution thread). -0. 81 0. 17 0. 45 0. 33 88 Gradient Hill Climb of Var(d)on t 143 With this p. Tree Decision Tree, we take the entire Test. Set (a PTree. Set), create the various dot product 0. 00 1. 00 0. 00 19 -0. 66 0. 19 0. 47 0. 56 95 SPTS (one for each inode), create ut SPTS Masks. These masks mask the results for the entire Test. Set. 0. 00 1. 00 0. 00 27 -0. 17 0. 35 0. 75 0. 53 54 -0. 32 0. 36 0. 65 0. 58 57 -0. 41 0. 34 0. 62 0. 58 58 For WINE: min max+1 Gradient Hill Climb of Var t 156161 8. 40 10. 33 27. 00 9. 63 28. 65 9. 9 53. 4 0. 00 1. 00 0. 00 5 7. 56 11. 19 32. 61 10. 38 34. 32 7. 7 111. 8 -0. 23 -0. 28 0. 89 0. 28 19 8. 57 12. 84 30. 55 11. 65 32. 72 8. 7 108. 4 -0. 02 -0. 06 0. 12 0. 99 157 8. 91 13. 64 34. 93 11. 97 37. 16 13. 1 92. 2 Awful results! 0. 02 -0. 02 1. 00 159 0. 00 1. 00 0. 00 1 -0. 46 -0. 53 0. 57 0. 43 2 Inconclusive both ways so predict purality=4(17) (3 ct=3 tct=6 Gradient Hill Climb of Var t 146156 0. 00 1. 00 0 0. 03 -0. 08 0. 81 -0. 58 1 0. 00 1. 00 0. 00 13 0. 02 0. 20 0. 92 0. 34 16 0. 02 0. 25 0. 86 0. 45 17 Inconclusive both ways so predict purality=4(17) (7 ct=15 Gradient 2 ct=2 Hill Climb of Var t 127 0. 00 1. 00 0. 00 41 -0. 01 0. 70 0. 71 90 -0. 04 0. 65 0. 75 91 0. 00 1. 00 0. 00 35 -0. 32 -0. 14 0. 59 0. 73 105 Inconclusive predict purality=7(62 4(15) 1(5) 2(8) 5(7)
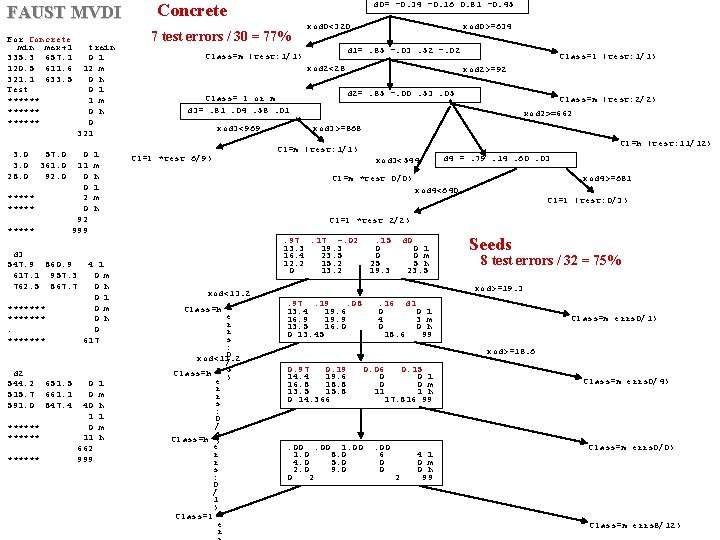
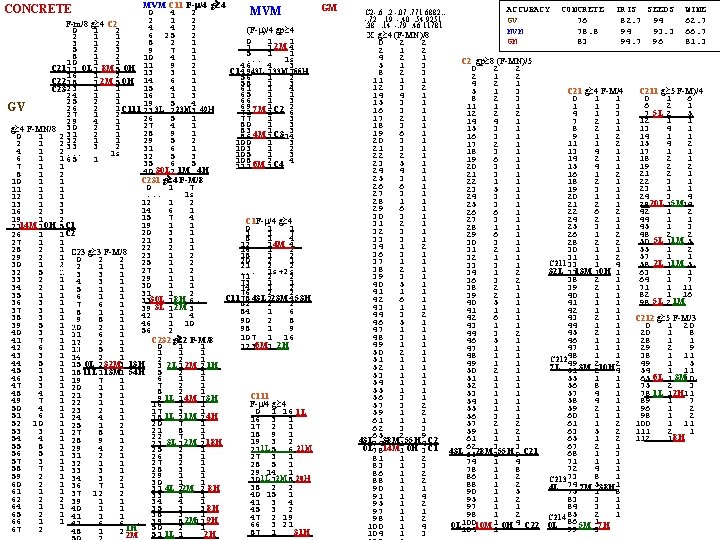
FAUST MVDI For Concrete min max+1 335. 3 657. 1 120. 5 611. 6 321. 1 633. 5 Test ****** 3. 0 28. 0 57. 0 361. 0 92. 0 train 0 l 12 m 0 h 0 l 1 m 0 h 0 321 ***** 0 11 0 0 2 0 92 999 ***** d 3 547. 9 860. 9 617. 1 957. 3 762. 5 867. 7 ******* d 2 544. 2 515. 7 591. 0 ****** 651. 5 661. 1 847. 4 l m h d 0= -0. 34 -0. 16 0. 81 -0. 45 7 test errors / 30 = 77% xod 0<320 xod 0>=634 d 1=. 85 -. 03. 52 -. 02 Class=m (test: 1/1) xod 2<28 Class= l or m d 3=. 81. 04. 58. 01 xod 3<969 Cl=l *test 6/9) Class=l (test: 1/1) xod 2>=92 d 2=. 85 -. 00. 53. 05 Class=m (test: 2/2) xod 2>=662 xod 3>=868 Cl=h (test: 11/12) Cl=m (test: 1/1) xod 3<544 d 4 =. 79. 14. 60. 03 Cl=m *test 0/0) xod 4>=681 xod 4<640 Cl=l (test: 0/3) Cl=l *test 2/2) 4 l 0 m 0 h 0 617 0 0 40 1 0 11 662 999 Concrete l m h . 97. 17 -. 02 13. 3 19. 3 16. 4 23. 5 12. 2 15. 2 0 13. 2 xod<13. 2 Class=h e r r s : 0 xod<13. 2 / Class=h 5 e ) r r s : 0 / 5 Class=h ) e r r s : 0 / 1 ) Class=l e r . 15 0 0 25 19. 3 d 0 0 l 0 m 5 h 23. 5 Seeds 8 test errors / 32 = 75% xod>=19. 3. 97. 19. 08 13. 4 19. 6 16. 9 19. 9 13. 5 16. 0 0 13. 45 . 16 d 1 0 0 l 4 3 m 0 0 h 18. 6 99 Class=m errs 0/1) xod>=18. 6 0. 97 0. 19 14. 4 19. 6 16. 8 18. 8 13. 5 15. 8 0 14. 366 . 00 1. 0 8. 0 4. 0 5. 0 2. 0 9. 0 0 2 0. 06 0. 15 0 0 l 0 0 m 11 1 h 17. 816 99 . 00 6 0 0 2 4 l 0 m 0 h 99 Class=m errs 0/4) Class=m errs 0/0) Class=m errs 8/12)

FAUST Classifier 0. Cut in middle of the means: means D≡m. R m. V d=D/|D| PR=Pxod<a PV=Pxod a a= (m. R+(m. V-m. R)/2)od = (m. R+m. V)/2 od 1. Cut in the middle of: Vector. Of. Medians (VOM), not the means. Use stdev ratio not middle for even better cut placement? 2. Cut in the middle of {Max{Rod}, Min{Vod}. (assuming m. Rod m. Vod) If no gap, move cut to minimize Rerrors + Verrors. 3. Hill-climb d to maximize gap or to minimize training set errors or (simplest) to minimize dis(max{rod}, min{vod}). 4. Replace mr, mv with the avg of the margin points? 5. PR=Px d<Cut. R o PV=Px d>Cut. V Min{Vod} Max{Rod} Cut. R=Cut. V=avg{min. Vod, min. Rod}, else Cut. R≡Min{Vod}, Cut≡Max{Rod} o y PR or y PV , Definite classifications; else re-do on Indefinite region, PCut. R xod Cut. V until actual gap (AND with certain stop cond? E. g. , "On nth round, use definite only (cut at midpt(m. R, m. V). " V Another way to view FAUST DI is that it is a Decision Tree Method. With each non-empty indefinite set, descend down the tree to a new level For each definite set, terminate the descent and make the classification. R dim 2 vom. R d 2 -l ine Each round, it may be advisable to go through an outlier removal process on each class before setting Min{Vod} and Max{Rod} (E. g. , Iteratively check if F 1(Min{Vod}) consists of V-outliers). vom. V Max. Rod Mn. Vod r v v r m. R r v v v v r r v m. V v r v d-line dim 1 d a

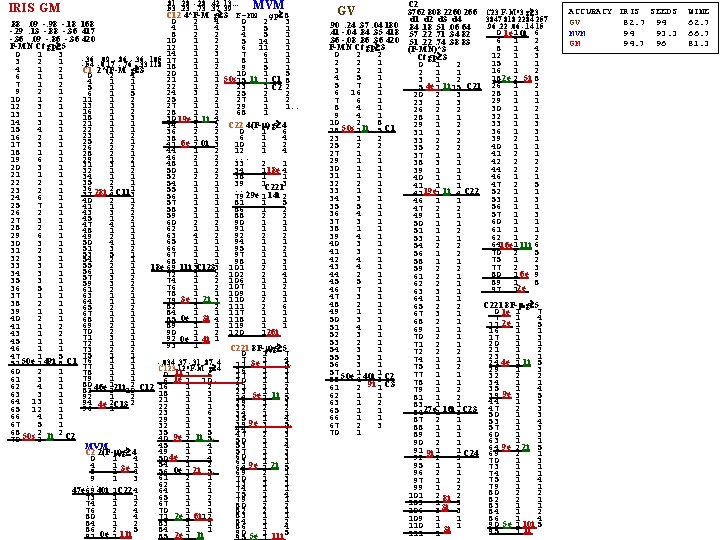
FAUST DI K-class training set, TK, and a given d (e. g. , from D≡Mean. TK Med. TK): Let mi≡mean. Ci s. t. dom 1 dom 2 . . . dom. K Mni≡Min{do. Ci} Mxi≡Max{do. Ci} Mn>i≡Minj>i{Mnj} Mx<i≡Maxj<i{Mxj} Definitei = ( Mx<i, Mn>i ) Indefinitei, i+1 = [ Mn>i, Mx<i+1 ] Then recurse on each Indefinite. For IRIS 15 records were extracted from each Class for Testing. The rest are the Training Set, TK. D=MEANs MEANe Definite_____ Indefinite__ s-Mean 50. 49 34. 74 14. 74 2. 43 s -1 25 e-Mean 63. 50 30. 00 44. 00 13. 50 e 10 37 se 25 10 empty i-Mean 61. 00 31. 50 55. 50 21. 50 i 48 128 ei 37 48 F < 18 setosa (35 seto) 1 ST ROUND D=Means Meane 18 < F < 37 versicolor (15 vers) 37 F 48 Indefinite. Set 2 (20 vers, 10 virg) 48 < F virginica (25 virg) F < 7 versicolor (17 vers. 0 virg) Indef. Set 2 ROUND D=Meane Meani 7 F 10 Indef. Set 3 ( 3 vers, 5 virg) 10 < F virginica ( 0 vers, 5 virg) F < 3 versicolor ( 2 vers. 0 virg) Indef. Set 3 ROUND D=Meane Meani 3 F 7 Indef. Set 4 ( 2 vers, 1 virg) Here we will assign 0 F 7 versicolor 7 < F virginica ( 0 vers, 3 virg) 7 < F virginica Test: F < 15 setosa (15 seto) 1 ST ROUND D=Means Meane 15 < F < 15 versicolor ( 0 vers, 0 virg) 15 F 41 Indefinite. Set 2 (15 vers, 1 virg) 41 < F virginica ( 14 virg) F < 20 versicolor (15 vers. 0 virg) Indef. Set 2 ROUND D=Meane Meani 20 < F virginica ( 0 vers, 1 virg) 100% accuracy. Option-1: The sequence of D's is: Mean(Classk) Mean(Classk+1) k=1. . . (and Mean could be replaced by VOM or? ) Option-2: The sequence of D's is: Mean(Classk) Mean( h=k+1. . n. Classh) k=1. . . (and Mean could be replaced by VOM or? ) Option-3: D seq: Mean(Classk) Mean( h not used yet. Classh) where k is the Class with max count in subcluster (Vo. M instead? ) Option-2: D seq. : Mean(Classk) Mean( h=k+1. . n. Classh) (VOM? ) where k is Class with max count in subcluster. Option-4: D seq. : always pick the means pair which are furthest separated from each other. Option-5: D Start with Median-to-Mean of Indefinite. Set, then means pair corresp to max separation of F(meani), F(meanj) Option-6: D Always use Median-to-Mean of Indefinite. Set, IS. (initially, IS=X)

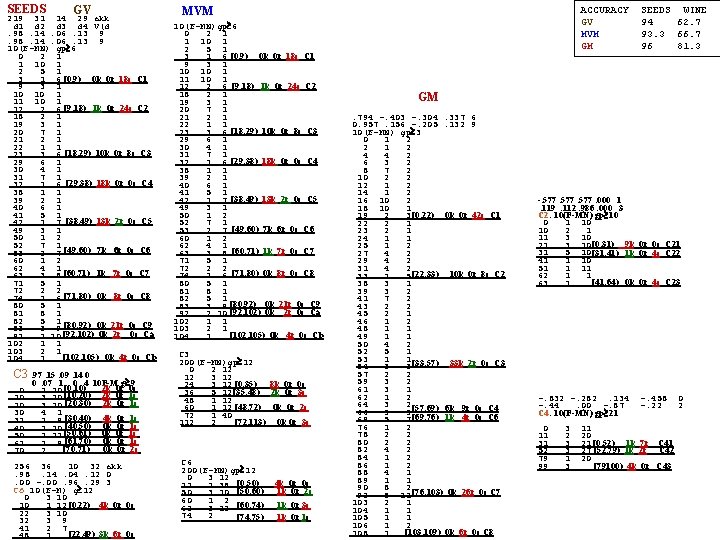
FAUST DI sequential For SEEDS 15 records were extracted from each Class for Testing. Option-4, means pair most separated in X. m 1 14. 4 5. 6 2. 7 5. 1 4. 4 d(m 1, m 2) DEFINITE INDEFINITE m 2 18. 6 6. 2 3. 7 6. 0 3. 4 d(m 1, m 3) 2 -inf 0 m 3 11. 8 5. 0 4. 7 5. 0 7. 0 d(m 2, m 3) 1 106 0 12 0 106 0 F 106, 3 106 inf 23 0 106 so totally non-productive! Option-6: D Median-to-Mean of Indef. Set (initially IS=X) m 1 14. 4 5. 6 2. 7 5. 1 37. 3 mean. F 1 DEFINITE Cl=1 2 3 INDEFINITE m 2 18. 6 6. 2 3. 7 6. 0 71. 2 mean. F 2 def 3[ -inf 21) 0 0 32 m 3 11. 8 5. 0 4. 7 5. 0 `2. 0 mean. F 3 def 1[ 28 49) 22 0 0 ind 1[ 21 28 ) On whole TR def 2[ 58 inf) 0 30 0 ind 2[ 49 58 ) Cl=1 2 3 m 1 13. 0 5. 1 3. 7 5. 0 30 av. F 1 DEFINITE INDEFINITE def 3[ -inf 0 ) 0 0 0 1 0 0 m 3 13. 0 5. 0 4. 0 5. 0 27 av. F 3 def 1[ 37 inf ) in 11[ 0 37 ) On Indef-1 Cls 1 outlier(F= Cl=1 2 3 m 1 13. 2 5. 2 4. 0 5. 0 9 av. F 1 DEFINITE INDEFINITE 54) def 3[ -inf 0 ) 0 0 0 1 0 0 m 3 13. 0 5. 0 4. 0 5. 0 6 av. F 3 def 1[ 13 inf ) in 11[ 0 13 ) On Indef-11 Cls 1 outlier Cl=1 2 3 m 1 13. 0 5. 2 3. 6 5. 0 13 av. F 1 DEFINITE INDEFINITE (F=29) def 3[ -inf 9 ) 0 0 0 1 m 3 13. 0 5. 0 4. 0 5. 0 9 av. F 3 def 1[ 19 inf ) in 111[ 9 19 ) On Indef-111 Cls 3 outlier Cl=1 (F=0) 2 3 m 1 13. 0 5. 2 3. 6 5. 0 13 av. F 1 DEFINITE INDEFINITE def 3[ -inf 9 ) 0 0 0 m 3 13. 0 5. 0 4. 0 5. 0 9 av. F 3 def 1[ 19 inf ) in 1111[ 9 19 ) On Indef-1111 done! declare Class=1 Cl=1 2 3 6 0 3 Cl=1 2 3 5 0 2

FAUST DI sequential For SEEDS 15 records were extracted from each Class for Testing. Option-6: D Median-to-Mean of X m 1 14. 4 5. 6 2. 7 5. 1 37. 3 mean. F 1 DEFINITE Cl=1 2 3 INDEFINITE m 2 18. 6 6. 2 3. 7 6. 0 71. 2 mean. F 2 def 3[ -inf 21) 0 0 32 m 3 11. 8 5. 0 4. 7 5. 0 `2. 0 mean. F 3 def 1[ 28 49) 22 0 0 ind 31[ 21 28 ) On whole TR def 2[ 58 inf) 0 30 0 ind 12[ 49 58 ) D Mean(lo. F)-to-Mean(hi. F) of Indef. Set 31 Cl=1 2 3 m 1 13. 0 5. 1 3. 7 5. 0 30 av. F 1 DEFINITE INDEFINITE m 3 13. 0 5. 0 4. 0 5. 0 27 av. F 3 def 1[-inf 18 ) 1 0 0 0. def 3[ 55 inf ) in 1313[ 18 55 ) Cl=1 2 3 . 6 0 3 D Mean(lo. F)-to-Mean(hi. F) of Indef. Set 1313 Cl=1 2 3 m 1 12. 8 5. 2 3. 2 5. 0 18 av. F 1 DEFINITE INDEFINITE m 3 13. 0 5. 0 4. 0 5. 0 10 av. F 3 def 3[ -inf 10 ) 0 0 1 1 0 0. 5 0 2. def 1[ 20 inf ) in 313131[ 10 20 ) . D Mean(lo. F)-to-Mean(hi. F) of Indef. Set 313131 (d repeats after this so=C 1 Cl=1 2 3 m 1 13. 0 5. 2 3. 6 5. 0 4 av. F 1 DEFINITE INDEFINITE 0 0 0 m 3 13. 0 5. 0 3. 5 5. 0 2 av. F 3 def 1[ -inf 0 ) 1 0 0 def 3[ 5 inf ) C 1= [ 0 5 ) The rest, Class=1 Cl=1 2 3 . 4 0 2 Cl=1 2 3 m 1 16. 2 6. 0 1. 8 5. 2 5. 8 av. F 1 DEFINITE INDEFINITE 5 0 0 m 2 16. 6 6. 0 4. 6 6. 0 6. 2 av. F 2 def 1[ -inf 2 ) 0 5 0 def 2[ 15 inf ) in 1212[ 2 15 ) Cl=1 2 3 . 0 0 0 D Mean(lo. F)-to-Mean(hi. F) of Indef. Set 12 [-inf, 21)class=3 [28, 49)class=2 [58. inf) class=3 d=(. , 9, -, 1, -. 2) [21, 28)ind 31 d=(-. 9, -. 1, . 14, -. 1) [49, 58)ind 12 d=(0, . 31, -. 9, 0) [-inf, 18)def [49, 58)ind 23

FAUST CLUSTERING Xod=Fd(X)=DPP V(d)≡Var. DPPd(X)= (Xod)2 - (Xod)2 d(X) x 1 od d 1 2 x 1 1 ( Xj dj )2 = ( j=1. . n xi, jdj) - x 2 od x j=1. . n N i=1. . N 2 = : dn x. Nod 1 ( x d ) = - ( X d ) ( X d ) ( k xi, kdk) x i, j j k k N j N i j k 2 x x d d 2 2 1 x 2 d 2 + = N i j i, j j N j<k i, j i, k j k - j. Xj dj +2 j<k Xj. Xkdjdk Use DPPd(x), but which unit vector, d *, provides the best gap(s)? 1. DPPd exhaustively searches a grid of d's for the best gap provider. 2. Use some heuristic to choose a good d? GV: Gradient-optimized Variance MM: Use the d that maximizes |Median. F(X)-Mean(F(X))|. We have Avg as a function of d. Median? (Can you do it? ) HMM: Use a heuristic for Median. F(X): F(Vector. Of. Medians)=VOMod MVM: Use D=MEAN(X) VOM(X), d=D/|D| do=ek s. t. akk is max or d 0 =akk k V(d)= 2 a 11 d 1 + j 1 a 1 jdj 2 a 22 d 2 + j 2 a 2 jdj : 2 anndn + j nanjdj +2 j<k. Xj. Xkdjdk - " = Xj 2 dj 2 j = j=1. . n d. T o VX o d 1 . . . dn consecutive 1 0 5 0 0 0 differences 1 0 0 2 0 0 1 0 0 0 3 0 0 0 1 0 0 2 0 6 0 0 1 0 0 0 9 0 1 0 0 2 3 0 0 10 1 0 0 0 0 1 0 0 2 0 3 0 0 1 0 0 0 3 0 0 0 1 10 5 2 1 1 1 0 avg. CD 1. 00 1. 00 max. CD 1. 00 10. 00 5. 00 2. 00 3. 00 6. 00 9. 00 10. 00 ||mean-VOM| 0. 00 0. 91 0. 00 0. 55 1. 18 1. 27 4. 00 4. 55 dn V(d)= jajjdj 2 + j kajkdjdk GRADIENT(V) = 2 A o d 0 0 0 0 1 0 5 0 0 0 2 0 5 2 0 0 3 0 5 2 3 0 0 0 4 0 5 4 3 6 0 0 median 5 0 5 4 3 6 9 0 6 0 5 6 6 6 9 10 7 0 5 6 6 6 9 10 8 0 5 8 6 9 9 10 9 0 5 8 9 9 9 10 10 10 std 3. 16 2. 87 2. 13 3. 20 3. 35 3. 82 4. 57 4. 98 variance 10. 0 8. 3 4. 5 10. 2 11. 2 14. 6 20. 9 24. 8 Avg 5. 00 0. 91 5. 00 4. 55 4. 18 4. 73 5. 00 4. 55 d = Var. DPPd. X≡V subject to i=1. . ndi 2=1 d 1 : V i Xi. Xj-Xi. X, j d 1≡ (V(d 0)) d 2≡ (V(d 1)) til F(dk) 2 a 11 2 a 12. . . 2 a 1 n 2 a 21 2 a 22. . . 2 a 2 n : ' 2 an 1 . . . 2 ann +(2 j=1. . n<k=1. . n (Xj. Xk - (Xj 2 - Xj 2)dj 2 + Xj. Xk)djdk ) d 1 : di : dn : V(d) = ijaijdidj Maximize variance - is it wise? MEDIAN picks out last 2 sequences which have best gaps (discounting outlier gaps at the extremes) and it discards 1, 3, 4 which are not so good. Finding good unit vector, d, for Dot Prod functional, DPP. to maximize gaps Maximize wrt d, |Mean(DPPd(X)) - Median(DPPd(X)| sub to i di 2=1 = Xjdj Mean(DPPd. X)=(1/N) j=1. . n xi, jdj = j (1/N i xi, j ) dj j=1. . n i=1. . N Compute Median(DPPd(X)? Want to use only p. Tree processing. Want a formula in d and numbers only (like the one above for the mean (involves only the vector d and the numbers X 1 , . . . , Xn )
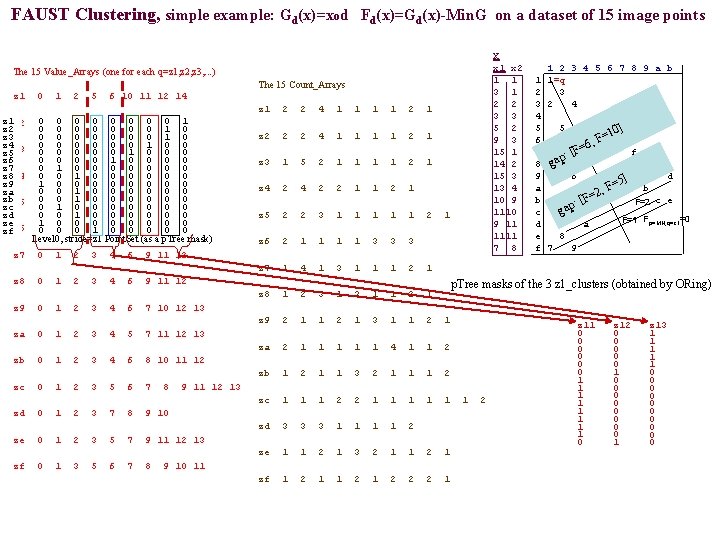
FAUST Clustering, simple example: Gd(x)=xod Fd(x)=Gd(x)-Min. G on a dataset of 15 image points X x 1 x 2 1 2 3 4 5 6 7 8 9 a b 1 1=q 3 1 2 3 2 4 3 3 4 5 2 5 5 10] = 9 3 6 F 6, F= 15 1 7 f [ : 14 2 8 gap 15 3 9 6 p d =5] 13 4 a b F , =2 10 9 b c e F=2 [F : p 1110 c ga F=1 Fp=MN, q=z 1=0 9 11 d a 1111 e 8 7 8 f 7 9 The 15 Value_Arrays (one for each q=z 1, z 2, z 3, . . . ) The 15 Count_Arrays z 1 0 1 2 5 6 10 11 12 14 z 1 2 2 4 1 1 2 1 0 0 0 0 1 z 1 z 2 0 1 2 5 6 10 11 12 14 0 0 0 0 1 0 z 2 0 0 0 0 1 0 z 3 0 0 0 1 0 0 z 4 z 3 0 1 2 5 6 10 11 12 14 0 0 0 1 0 0 0 z 5 0 0 1 0 0 z 6 0 1 0 0 0 0 z 7 0 0 1 0 0 0 z 8 z 4 0 1 3 6 10 11 12 14 1 0 0 0 0 z 9 0 0 1 0 0 0 za 0 0 1 0 0 0 zbz 5 0 1 2 3 5 6 10 11 12 14 0 1 0 0 0 0 zc 0 0 1 0 0 0 zd 1 0 0 0 0 ze z 6 0 1 2 3 7 8 9 10 0 1 0 0 0 zf Level 0, stride=z 1 Point. Set (as a p. Tree mask) z 2 2 2 4 1 1 2 1 z 3 1 5 2 1 1 2 1 z 4 2 2 1 1 2 1 z 5 2 2 3 1 1 1 2 1 z 6 2 1 1 3 3 3 z 7 0 1 2 3 4 6 9 11 12 z 7 1 4 1 3 1 1 1 2 1 z 8 0 1 2 3 4 6 9 11 12 z 8 1 2 3 1 1 2 1 p. Tree masks of the 3 z 1_clusters (obtained by ORing) z 9 0 1 2 3 4 6 7 10 12 13 z 9 2 1 1 2 1 3 1 1 2 1 za 0 1 2 3 4 5 7 11 12 13 za 2 1 1 1 4 1 1 2 zb 0 1 2 3 4 6 8 10 11 12 zb 1 2 1 1 3 2 1 1 1 2 zc 0 1 2 3 5 6 7 8 9 11 12 13 zc 1 1 1 2 2 1 1 1 2 zd 0 1 2 3 7 8 9 10 zd 3 3 3 1 1 2 ze 0 1 2 3 5 7 9 11 12 13 ze 1 1 2 1 3 2 1 1 2 1 zf 0 1 3 5 6 7 8 9 10 11 zf 1 2 1 2 2 2 1 z 11 0 0 0 1 1 1 1 0 z 12 0 0 0 1 0 0 0 0 1 z 13 1 1 1 0 0 0 0 0

What have we learned? What is the DPPd FAUST CLUSTER algorithm? X 2=Sub. Cluster 2 D=Median Mean, d 1≡D/|D| is a good start. But first, Variance-Gradient hill-climb it. (Median means Vector of Medians). For X 2=Sub. Cluster 2 use a d 2 which is perpendicular to d 1? In high dimensions, there are many perpendicular directions. GV hill-climb d 2=D 2/|D 2| (D 2=Median. X 2 -Mean. X 2) constrained to be to d 1, i. e. , constrained to d 2 od 1=0 (in addition to d 2 od 2=1. We may not want to constrain this second hill-climb to unit vectors perpendicular to d 1. It might be the case that the gap gets wider using a d 2 which is not perpendicular to d 1? Sub. Cluster 1 GMP: Gradient hill-climb (wrt d) Variance. DPPd starting at d 2=D 2/|D 2| where d 2≡Unitized( Vom{x-xod 1|x X 2} - Mean{x-xod 1|x X 2} ) Variance-Gradient hillclimbed subject only to dod=1 (We shouldn't constrain the 2 nd hill-climb to d 1 od 2=0 and subsequent hill-climbs to dkodh=0, h=2. . . k-1. (gap could be larger). So the 2 nd round starts at d 2≡Unitized( Vom{x-xod 1|x X 2} - Mean{x-xod 1|x X 2} ) and hill-climbs subject only to dod=1) GCCP: Gradient hill-climb (wrt d) Variance. DPPd starting at d 2=D 2/|D 2| where D 2=CCi(X 2)-CCj(X 2), and hill-climbs subject to dod=1, where the CCs are two of the Circumscribing rectangle's Corners (the CCs may be a faster calculations than Mean and Vom). Taking all edges and diagonals of CCR(X) (the Coordinate-wise Circumscribing Rectangle of X) provides a grid of unit vectors. It is an equispaced grid iff we use a CCC(X) (Coordinate-wise Circumscribing Cube of X). Note that there may be many CCC(X)s. A canonical one is the one that is furthest from the origin (take the longest side first. Extend each other side the same distance from the origin side of that edge. A good choice may be to always take the longest side of CR(X) as D, D≡LSCR(X). Should outliers on the (n-1)-dim-faces at the ends of LSCR(X) be removed first? So remove all LSCR(X)-endface outliers until after removal the same side is still the LSCR(X). Then use that LSCR(X) as D.
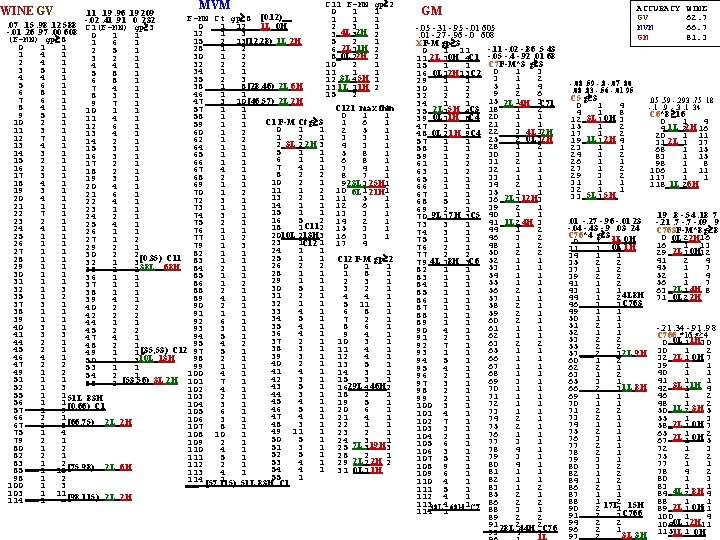
C 11 F-MN gp 2 MVM 0 1 1. 19. 96. 19 209 F-MN Ct gp 8 [0. 12) 1 1 1. 41. 91 0 232. 07. 15. 98. 12 588 -. 02 1 L 0 H 0 1 12 2 3 1 C 1(F-MN) gp 3 -. 01. 26. 97. 00 608 0 _4 L 2 H 12 1 3 3 3 2 1 1 (F-MN) gp 8 15 2 13 ___ _ [12, 28) 1 L 2 H 5 2 1 1 6 1 0 1 1 _2 L 1 H 28 1 2 6 1 2 2 5 1 1 4 1 _0 L 2 H 30 1 2 8 2 2 3 2 1 2 4 1 32 2 2 10 2 1 4 4 1 3 5 1 34 1 1 1 5 8 1 4 4 1 35 2 3 12 3 L 45 H 1 6 8 1 5 6 1 _8 [28, 46) 2 L 6 H 38 ___ 1 13 1 L 11 H 2 7 4 1 6 8 1 46 1 1 15 2 8 3 1 7 6 1 _ [46, 57) 2 L 2 H 47 ___ 3 10 9 7 1 8 4 1 C 121 max thin 57 1 1 10 1 1 0 1 1 9 5 1 58 1 1 11 4 1 C 1 F-M Ct g 3 1 6 1 10 2 1 59 1 1 12 6 1 0 1 1 2 5 1 11 3 1 60 1 2 13 4 1 1 2 1 3 3 1 12 7 1 62 1 2 14 2 1 2 2 3 4 3 1 3 L 2 H 13 4 1 64 1 1 15 3 1 5 1 1 5 8 1 14 3 1 65 1 1 16 3 1 6 1 1 6 8 1 15 2 1 66 1 1 17 2 1 7 4 1 7 4 1 16 2 1 67 4 1 18 2 1 8 2 2 8 7 1 17 3 1 68 2 1 19 3 1 10 2 1 9 3 1 18 4 1 23 L 25 H 69 1 1 20 4 1 11 1 2 10 1 1 19 3 1 6 L 21 H 70 1 2 21 6 1 13 2 1 11 5 1 20 4 1 72 3 1 22 4 1 1 12 6 1 21 1 1 73 1 1 23 1 1 15 1 1 13 3 1 22 7 1 74 3 1 24 2 1 16 5 2 14 2 1 23 2 1 75 2 1 25 4 1 18 1 C 112 15 3 1 24 4 1 76 1 1 2010 L 213 H 3 16 3 1 25 1 1 77 1 2 23 1 C 12 1 17 4 26 1 1 79 1 3 29 2 1 24 1 1 27 1 1 82 1 1 30 2 2 25 1 1 C 12 F-M gp 2 [0. 35) C 11 83 28 1 1 32 1 3 26 2 2 0 1 1 29 1 1 38 L 68 H 84 2 1 35 1 1 28 1 1 1 8 1 30 1 1 85 1 1 36 1 1 29 1 1 2 3 1 31 1 1 86 1 2 37 1 1 30 5 1 3 2 1 3 88 2 1 38 1 1 31 2 1 4 4 1 35 1 2 89 4 1 32 1 1 5 11 1 37 3 1 90 2 1 40 2 2 33 4 1 6 8 1 38 1 1 91 1 1 42 2 2 34 5 1 7 2 1 39 1 1 92 6 1 44 1 1 35 4 1 8 6 1 40 3 1 93 3 1 45 2 2 36 4 1 9 4 1 41 3 3 94 5 1 47 4 1 37 2 1 10 3 1 44 2 1 95 4 2 48 2 1 38 3 1 11 4 1 45 2 1 5 1 49 1 1 [35, 53) C 12 97 39 3 1 12 4 1 46 4 1 10 L 13 H 98 2 1 50 1 3 40 2 1 13 5 1 47 2 2 99 1 1 53 1 1 41 4 1 14 3 1 49 1 2 100 4 1 54 2 1 42 3 1 15 3 1 51 1 1 ___ 2 [53, 56) 3 L 2 H 101 7 1 55 43 5 1 16 29 L 4 46 H 2 52 1 3 102 4 1 44 3 1 18 2 1 55 1 1 51 L 83 H 103 2 1 45 4 1 19 5 1 56 1 1 [0. 66) C 1 104 3 1 46 5 1 20 6 1 57 1 9 105 6 1 47 4 1 21 4 1 66 2 1 106 3 1 _8 [66, 75) 2 L 2 H 48 3 1 22 1 1 67 ___ 2 107 8 1 49 11 1 23 2 1 75 1 4 108 10 1 50 5 1 24 3 1 79 2 1 109 2 1 7 L 19 H 51 3 1 25 3 3 80 1 2 110 4 1 52 5 1 28 2 1 82 2 1 111 5 1 53 4 1 29 2 L 2 2 H 2 83 ___ 1 2 [75, 98) 2 L 6 H 112 2 1 _ 54 4 1 31 0 L 1 1 H 85 1 13 113 4 1 55 1 98 1 2 114 [57, 115) 1 ___ 51 L 83 H C 1 100 1 3 103 ___ 1 11 _ [98, 115) 2 L 2 H 114 1 WINE GV ACCURACY WINE GV 62. 7 MVM 66. 7 GM 81. 3 GM -. 05 -. 31 -. 95 -. 01 605 . 01 -. 27 -. 96 -. 0 608 XF-M gp 3 -. 11 -. 02 -. 86. 5 43 0 1 11 11 2 L 10 H 4 C 1 -. 05 -. 4 -. 92. 01 68 C 7 F-M*3 g 3 15 1 1 16 _0 L 12 H 13 C 2 0 1 3 3 1 2 29 1 1 5 1 4 30 1 2 9 2 6 32 2 2 15 1 3 _2 L 4 H C 71 34 1 1 18 1 2 _2 L 5 H C 3 35 2 4 20 1 1 39 _0 L 11 H 8 C 4 21 1 1 47 2 1 ___ 4 L 2 H 48 _0 L 2 1 H 9 C 4 22 3 3 25 2 3 ___ 0 L 2 H 57 1 1 28 1 2 58 1 1 30 3 1 59 1 2 31 2 1 61 1 2 32 1 1 63 1 2 33 1 1 65 1 1 34 2 1 66 1 1 35 1 1 67 1 1 _2 L 12 H 36 3 3 68 5 1 39 2 1 69 2 1 70 _9 L 1 7 H 3 C 5 40 1 1 _1 L 4 H 41 2 3 73 3 1 44 1 2 74 3 1 46 3 2 75 1 1 48 1 2 76 2 1 50 2 2 77 2 2 79 4 L 18 H 3 C 6 52 1 1 53 1 1 82 1 1 54 1 1 83 1 1 55 1 1 84 1 1 56 2 1 85 1 1 57 1 1 86 1 1 58 2 1 87 1 1 59 2 1 88 1 1 60 2 1 89 1 1 61 1 1 90 4 1 62 1 1 91 2 1 63 2 2 92 7 1 65 1 1 93 1 1 66 1 1 94 5 1 67 1 1 95 4 1 68 1 1 96 2 1 69 3 1 97 3 1 70 1 1 98 2 1 71 1 1 99 2 1 72 1 1 100 3 1 73 1 1 101 4 1 74 2 1 102 7 1 75 2 1 103 3 1 76 1 1 104 2 1 77 3 1 105 6 1 78 4 1 106 3 1 79 3 1 107 5 1 80 4 1 108 9 1 81 1 1 109 6 1 82 1 1 110 4 1 83 1 2 111 5 1 85 2 1 112 4 1 11338 L 4 68 H 1 C 7 86 2 2 88 3 1 114 1 89 2 2 91 2 2 ___ 28 L 44 H C 76 93 4 3 1 L -. 08. 59 -. 8 -. 07 80. 08. 83 -. 56 -. 01 95 C 5 g 3 0 1 4 4 1 8 _3 L 0 H 12 1 3 15 1 2 17 1 2 _1 L 2 H 19 1 4 23 1 1 24 1 2 26 1 1 27 1 2 29 3 2 31 1 1 32 1 1 5 L 5 H 33 1 . 01 -. 27 -. 96 -. 01 23 -. 04 -. 43 -. 9. 03 24 C 76*4 g 3 ___ _1 L 0 H 0 1 31 ___ _0 L 1 H 31 1 3 34 1 1 35 2 2 37 1 2 39 2 2 41 1 2 43 1 1 4 L 8 H 44 1 2 46 3 3 C 763 49 1 1 50 1 1 51 2 1 52 1 1 53 2 2 55 2 2 ___ _ 2 L 9 H 57 2 3 60 1 2 62 2 1 63 1 2 65 3 1 ___ _ 1 L 8 H 66 2 3 69 1 1 70 1 1 71 2 2 73 2 1 74 1 1 75 2 1 76 3 1 77 2 1 78 2 1 79 3 1 80 3 2 82 1 2 84 1 2 86 2 1 87 1 1 88 1 2 17 L 15 H 90 2 1 91 2 3 C 766 94 2 2 96 2 1 ___ _ 3 L 3 H 97 2 . 05. 59 -. 293. 75 18 -. 1 . 9 -. 3. 1 34 C 6*8 16 0 1 4 _1 L 2 H 4 2 16 20 1 11 _2 L 31 1 37 68 1 15 83 1 15 98 1 8 106 1 11 117 1 1 _1 L 6 H 118 2 . 19. 8 -. 54. 18 7 -. 21. 7 -. 09 9 C 763 F-M*8 g 8 0 2 16 0 L 2 H 16 1 13 _2 L 0 H 29 1 12 41 2 4 45 1 7 52 1 4 56 1 7 _2 L 4 H 63 1 8 _0 L 2 H 71 2 -. 21. 34 -. 91. 9 8 C 766 *16 g 4 _0 L 1 H 0 1 30 30 1 2 _2 L 0 H 32 1 7 39 1 1 40 1 1 41 1 1 _3 L 1 H 42 1 4 46 1 2 48 1 2 _1 L 3 H 50 2 5 55 1 3 _2 L 0 H 58 1 7 65 1 2 _2 L 0 H 67 1 5 72 1 3 75 2 2 77 1 1 78 4 2 80 1 3 83 1 1 _4 L 8 H 84 2 4 88 1 1 _2 L 0 H 89 1 11 100 1 4 _0 L 2 H 104 1 11 115 1 _1 L 0 H
SEEDS GV 219 31 14 29 akk d 1 d 2 d 3 d 4 V(d. 98. 14. 06. 13 9 10(F-MN) gp 6 0 2 1 1 10 1 2 5 1 1___6 [0, 9) 0 k 0 r 18 c C 1 ___3 9 3 1 10 10 1 11 10 1 ___ 12 2___6 [9, 18) 1 k 0 r 24 c C 2 18 2 1 19 3 1 20 7 1 21 2 1 22 1 1 ___ 23 3___6 [18, 29) 10 k 0 r 8 c C 3 29 6 1 30 4 1 31 7 1 ___ 32 1 ___6 [29, 38) 18 k 0 r 0 c C 4 38 1 1 39 2 1 40 6 1 41 5 1 ___ 42 1 ___7 [38, 49) 13 k 2 r 0 c C 5 49 3 1 50 1 2 52 7 1 ___ 53 2___7 [49, 60) 7 k 6 r 0 c C 6 60 1 2 62 4 1 ___ 63 3___8 [60, 71) 1 k 7 r 0 c C 7 71 5 1 72 2 2 ___ 74 1___6 [71, 80) 0 k 8 r 0 c C 8 80 5 1 81 8 1 82 5 1 ___ 83 3___9 [80, 92) 0 k 21 r 0 c C 9 ___ 92 2___ 10 [92, 102) 0 k 2 r 0 c Ca 102 1 1 103 2 1 ___ 104 1___ [102, 105) 0 k 4 r 0 c Cb C 3. 97. 15. 09. 14 0 ___ 10 ___ 20 30 ___ 31 ___ 40 ___ 50 ___ 61 ___ 70 0. 07 1 0 4 10 F-M g 9 2 k 0 r 0 c 2___ 10 [0, 10) 3___ 10 [10, 20) 2 k 0 r 1 c 3___ 10 [20, 30) 2 k 0 r 1 c 4 1 1___9 [30, 40) 4 k 0 r 1 c 1___ 10 [40, 50) 0 k 0 r 1 c 1___ 11 [50, 61) 0 k 0 r 1 c ___ 1 9 [61, 70) 0 k 0 r 1 c 0 k 0 r 2 c 2___ [70, 71) 256 36 10 32 akk. 98. 14. 04. 12 0. 00 -. 00. 96. 29 3 C 6 10(F-M) g 12 0 3 10 10 1___ 12 [0, 22) 4 k 0 r 0 c ___ 22 3 10 32 3 9 41 2 7 ___ 48 1___ [22, 49) 3 k 6 r 0 c MVM 10(F-MN)gp 6 0 2 1 1 10 1 2 5 1 ___3 1___6 [0, 9) 0 k 0 r 18 c C 1 9 3 1 10 10 1 11 10 1 ___ 12 2 ___6 [9, 18) 1 k 0 r 24 c C 2 18 2 1 19 3 1 20 7 1 21 2 1 22 1 1 ___ 23 3___6 [18, 29) 10 k 0 r 8 c C 3 29 6 1 30 4 1 31 7 1 ___ 32 1___6 [29, 38) 18 k 0 r 0 c C 4 38 1 1 39 2 1 40 6 1 41 5 1 ___ 42 1___7 [38, 49) 13 k 2 r 0 c C 5 49 3 1 50 1 2 52 7 1 ___ 53 2 ___7 [49, 60) 7 k 6 r 0 c C 6 60 1 2 62 4 1 ___ 63 3 ___8 [60, 71) 1 k 7 r 0 c C 7 71 5 1 72 2 2 ___ 74 1 ___6 [71, 80) 0 k 8 r 0 c C 8 80 5 1 81 8 1 82 5 1 ___ 83 3 ___9 [80, 92) 0 k 21 r 0 c C 9 ___ 92 2 ___ 10 [92, 102) 0 k 2 r 0 c Ca 102 1 1 103 2 1 ___ 104 1 ___ [102, 105) 0 k 4 r 0 c Cb C 3 200(F-MN)gp 12 0 2 12 12 3 12 ___ 8 k 0 r 0 c 24 3___ 12 [0, 35) ___ 36 5___ 12 [35, 48) 2 k 0 r 3 c 48 1 12 60 1 ___ 12 [48, 72) 0 k 0 r 2 c ___ 72 1 40 ___ 112 2 ___ [72, 113) 0 k 0 r 3 c C 6 200(F-MN)gp 12 0 3 12 ___ 12 1 ___ 38 [0, 50) ___ 50 3 ___ 10 [50, 60) 60 1 2 ___ 62 3 ___ 12 [60, 74) 74 2 ___ [74, 75) ___ 4 k 0 r 0 c 1 k 0 r 2 c 1 k 0 r 3 c 1 k 0 r 1 c ACCURACY SEEDS WINE GV 94 62. 7 MVM 93. 3 66. 7 GM 96 81. 3 GM. 794 -. 403 -. 304. 337 6 0. 957. 156 -. 205. 132 9 10(F-MN) gp 3 0 1 2 2 1 2 4 4 2 6 3 2 8 7 2 10 2 2 12 1 2 14 1 2 16 10 2 18 10 1 ___ 19 2 ___3 [0, 22) 0 k 0 r 42 c C 1 22 2 1 23 2 1 24 1 1 25 1 2 27 4 2 29 4 2 31 4 2 ___ 33 2 ___5 [22, 33) 10 k 0 r 8 c C 2 38 3 1 39 3 2 41 7 2 43 2 2 45 2 1 46 1 2 48 1 1 49 1 1 50 4 2 52 5 1 53 1 ___1 [33, 57) 33 k 2 r 0 c C 3 ___ 54 3 3 57 2 2 59 3 2 61 3 1 62 1 2 64 3 ___2 [57, 69) 6 k 9 r 0 c C 4 ___ 66 3 3 ___ 69 5 ___7 [69, 76) 1 k 4 r 0 c C 6 76 1 2 78 2 2 80 2 2 82 4 2 84 1 2 86 1 2 88 4 1 89 1 1 90 8 2 ___ 92 5 11 [76, 103) 0 k 26 r 0 c C 7 103 2 1 104 1 1 105 1 1 106 1 2 ___ 108 1___ [103, 109) 0 k 6 r 0 c C 8 -. 577. 000 1. 119. 112. 986. 000 3 C 2: 10(F-MN) gp 10 0 1 10 10 2 1 11 3 10 ___ 21 3 ___ 10 [0, 31) 9 k 0 r 0 c C 21 31 5 ___ 10 [31, 41) 1 k 0 r 4 c C 22 ___ 41 1 10 51 1 11 62 1 1 ___ 63 1 ___ [41, 64) 0 k 0 r 4 c C 23 -. 832 -. 282. 134 -. 44. 00 -. 87 C 4: 10(F-MN) gp 21 0 11 ___ 31 ___ 52 79 99 ___ -. 458 -. 22 3 11 2 20 3 ___ 21 [0, 52) 1 k 7 r C 41 3 ___ 27 [52, 79) 1 k 2 r C 42 1 20 3 ___ [79100) 4 k 0 r C 43 0 2
IRIS GM . 81. 28 -. 28. 42 13. . 53. 23. 73. 37 39 MVM C 12 4*F-M g 3 F-MN gp 8 0 2 4 0 2 3. 88. 09 -. 98 -. 18 168 4 1 4 3 5 1 -. 29. 13 -. 88 -. 36 417 8 2 2 4 5 1 -. 36. 09 -. 86 -. 36 420 10 1 2 5 14 1 F-MN Ct gp 5 12 1 2 6 11 1 14 1 3 0 1 3 7 6 1 -. 36. 09 -. 86 -. 36 105 17 1 1 8 1 1 3 2 1 -. 54 -0. 17 -. 76 -. 33 118 18 1 2 9 5 1 4 1 2 C 1 2*(F-M g 3 20 1 1 10 1 5 0 2 4 6 1 1 50 s 1 i C 1 21 1 1 15 1 8 4 1 1 7 1 2 22 1 2 23 1 C 2 2 5 1 1 9 2 1 24 3 1 25 2 2 6 1 5 25 1 2 10 1 2 11 1 2 27 1 2 13 1 3 27 1 1 29 1 1. . 12 3 1 16 1 2 28 1 2 68 1 13 1 1 18 1 3 30 1 4 ___ 19 e 1 i 14 3 1 21 1 1 34 1 2 C 22 4(F- ) g 4 22 1 1 15 4 1 1 6 36 2 2 0 23 1 2 16 2 1 6 1 4 38 1 3 25 2 1 ___ 6 e 0 i 10 1 2 41 2 3 17 3 1 26 2 2 12 1 4 44 1 2 18 1 1 28 2 1. . . 46 2 2 29 1 2 19 6 1 33 2 1 48 1 2 31 3 1 20 3 1 34 1 18 e 4 ___ 50 1 2 32 1 2 21 1 1 34 1 1 38 1 1 52 2 2 22 2 1 35 2 1 39 1 C 2213 54 1 1 36 2 1. . . 55 1 1 23 2 1 ___28 i 37 4 C 113 79 29 e 1 14 i 2 56 1 1 24 6 1 40 2 1 81 1 ___5 57 1 1 ___ 25 7 1 41 1 2 86 1 2 58 3 1 43 3 2 26 2 1 88 2 2 59 1 1 45 1 2 27 3 1 90 1 1 60 2 2 47 4 1 28 2 1 91 1 1 62 1 1 48 1 1 92 2 2 63 4 2 29 6 1 49 2 1 94 1 1 65 1 1 50 4 1 30 3 1 51 3 2 95 1 2 66 1 1 31 2 1 53 5 1 97 1 1 67 1 1 32 3 1 54 2 1 98 1 3 68 1 1 33 3 1 55 2 1 18 e 11 i C 123 101 2 1 69 3 3 56 1 1 34 3 1 2 4 72 1 2 102 57 3 2 106 1 1 74 1 2 35 3 1 59 3 2 107 1 2 76 1 2 36 5 1 61 2 2 109 1 1 78 1 1 63 1 1 37 1 1 ___ 3 e 2 i 110 2 1 79 1 3 64 1 1 38 2 1 2 6 82 1 2 111 65 2 2 39 1 1 84 1 1 117 67 1 1 40 2 1 ___ 68 1 1 85 0 e 1 3 i 4 118 69 2 1 1 1 89 1 1 119 41 1 2 70 2 1 120 1 ___ 26 i 90 1 2 43 1 2 71 3 1 92 1 1 ___ 0 e 4 i 45 1 1 72 1 1 93 1 C 221 8 F- )g 5 46 1 1 73 2 2 0 1 7 75 1 1 47 1 5 7 1 4. 76 1 1 ___50 e 49 i C 1 -. 034. 37 -. 31. 87 4 ___ 52 1 8 11 3 e 1 5 77 1 1 C 123 12*F-M g 4 16 1 1 60 2 1 __ 1 i. 78 1 1 0 1 6 17 1 3 79 1 1 61 3 1 ___ 6 1 e 1 10. 20 1 1 80 1 2 21 1 2 62 4 1 16 1 2 _46 e 21 i C 12 82 2 10 23 1 18 1 3 63 3 1 ___ 5 e 1 1 i 1 92 1 2 24 5 21 1 1 64 13 1 29 1 3 94 4 e 2 C 13 2 ___ 22 1 1 32 2 2 96 1 65 12 1 23 1 6 34 1 1 66 4 1 35 1 4 29 1 3 ___9 e 39 3 5. 67 5 1 32 1 3 44 1 3 68 2 2 35 1 5 47 2 3 ___ 50 s 1 i C 2 ___ 9 e 1 i. 40 2 5 70 2 50 1 3 45 1 4 53 1 4 MVM 57 1 3 49 1 1 C 2 2(F- )g 4 60 1 3 _50 4 e 2 4. 0 1 4 63 1 2 i 1 54 1 2 4 1 1 __9 e 64 2 5 ___ 3 e __ 56 0 e 1 2 i 5. 5 1 4 69 2 1 61 2 1 9 1 3 70 1 3 73 1 1 62 1 2. . . 74 1 1 47 e 69 40 i 1 C 22 4 64 1 1 75 1 4 65 1 2 73 1 1 79 1 1 67 1 3 74 1 2 80 2 2 70 1 1 76 2 4 82 2 1 83 1 1 ___ 2 e 1 6 i 12. 71 80 1 4 84 1 2 83 1 1 84 1 2 86 1 4 84 1 1 86 2 5 90 1 ___ 2 e 1 1 i 85 91 0 e 1 11 i ___ 5 e 11 i 5 GV. 90. 24. 37. 04 180. 41 -. 04. 84. 35 418. 36 -. 08. 86. 36 420 F-MN Ct gp 3 0 2 2 1 3 2 1 4 5 1 5 7 1 6 16 1 7 6 1 8 4 1 9 4 1 10 2 8 ___50 s 1 i 18 1 5 C 1 23 1 2 25 2 2 27 1 2 29 1 1 30 1 1 31 1 1 32 2 1 33 1 1 34 3 1 35 5 1 36 4 1 37 3 1 38 1 1 39 4 1 40 3 1 41 3 1 42 4 1 43 4 1 44 2 1 45 5 1 46 7 1 47 3 1 48 2 1 49 1 1 50 3 1 51 4 1 52 3 1 53 2 1 54 3 1 55 3 1 56 3 1 57 50 e 1 40 i 1 C 2 ___ 58 4 9 i 3 C 3 61 2 1 62 1 1 63 1 2 65 1 1 66 1 1 67 2 3 70 1 C 2 3762 808 2260 266 C 23 F-M*3 g 3 3847 818 2284 257 d 1 d 2 d 3 d 4. 96. 22. 06 -. 14 15. 84. 18. 51. 06 64 ___0 1 e 1 0 i 6. 57. 22. 71. 34 82 6 1 2. 51. 22. 74. 38 83 8 1 4 (F-MN)*3 12 1 3 Ct gp 3 15 1 1 0 1 2 16 1 2 2 1 1 18 __2 e 2 5 i 8 3 1 2 2 ___ 5 4 e 1 1 i 15 C 21 26 1 28 1 1 20 2 3 29 1 1 23 1 3 30 1 2 26 2 2 32 1 1 28 1 1 33 1 3 29 1 2 36 1 3 31 1 2 39 2 1 33 2 2 40 1 1 35 2 2 41 2 1 37 1 1 42 2 2 38 3 1 44 2 2 39 1 1 46 1 1 40 1 1 47 2 5 41 1 ___4219 e 1 1 i 4 C 22 52 1 53 1 3 46 1 1 56 1 1 47 2 2 57 1 3 49 1 1 60 1 1 50 1 1 61 1 1 51 1 2 62 1 2 53 1 1 64 1 ___ 16 e 11 i 6 54 2 2 70 2 5 56 1 2 75 1 2 58 1 1 77 2 3 59 2 2 ___ 80 1 6 e 9 61 2 1 89 1 8 62 2 1 ___97 12 e 63 3 1 64 1 1 C 221 8 F- g 5 65 2 2 0 7 ___1 e 1 67 3 1 7 1 4 68 2 1 ___ 11 2 e 1 5 69 1 1 16 1 1 70 2 1 17 1 3 20 1 1 71 2 1 21 1 2 72 2 2 23 1 1 74 1 1 __ 4 e 1 i 24 1 5 75 1 2 29 1 3 77 1 1 32 2 2 78 1 1 34 1 1 35 1 4 79 1 2 ___9 e 3 39 5 81 1 2 44 1 3 83 1 1 47 2 3 ___8427 e 1 16 i 3 C 23 50 1 3 87 2 1 53 1 4 88 1 1 57 1 3 60 1 3 89 1 1 63 1 1 90 2 1 64 2 5 __9 e 2 i 91 9 i 1 1 C 24 69 ___ 2 1 92 2 3 70 1 3 95 1 1 73 1 1 96 2 1 74 1 1 75 1 4 97 1 2 79 1 1 99 1 2 80 2 2 101 2 8 i 2 ___ 82 2 1 103 1 3 83 1 1 ___ 106 3 3 i 3 84 1 2 109 1 1 86 1 4 ___ 5 e 10 i 90 1 5 110 1_3 i 1 ___ 1 1 i 95 111 1 ACCURACY IRIS SEEDS WINE GV 82. 7 94 62. 7 MVM 94 93. 3 66. 7 GM 94. 7 96 81. 3
MVM C 11 F- /4 g 4 GM 0 4 2 MVM 2 1 2 F-m/8 g 4 C 2 4 4 2 (F- )/4 gp 4 0 1 2 6 25 2 2 1 1 0 1 1 8 2 1 3 1 2 ___ 1 1 2 M 4 5 2 3 9 7 1 5 1 1 8 1 2 10 4 1. . . 1 s 10 1 1 46 4 3 11 9 2 C 2111 0 L 1 8 M 5 0 H 13 C 149 43 L 133 M 755 H 3 1 16 1 2 56 1 2 6 1 C 2218 1 2 M 5 0 H 14 58 1 3 15 4 1 1 1 61 1 4 C 2323 65 1 1 24 1 1 16 1 3 66 1 3 25 2 1 19 5 4 69 1 2 26 2 1 C 111 23 3 L 2 23 M 3 49 H GV ___ 7 M C 2 71 1 6 27 1 2 26 5 1 77 1 3 29 4 1 80 1 3 27 4 1 2 1 g 4 F-MN/8 30 83 1 3 28 9 1 2 1 ___ 4 M 1 C 314 0 1 2 31 86 1 1 29 5 2 100 1 3 2 1 2 32 3 2 103 1 2 31 6 1 4 1 2 33. . . 1 s 105 1 3 32 5 3 6 1 1 65 1 108 2 35 6 5 ___6 M 1 C 4 4 112 7 1 1 ___ 40 30 L 2 1 M 4 H 8 1 2 C 231 g 4 F-M/8 10 1 1 0 1 7 11 1 1. . . 1 s 12 1 1 12 1 2 13 1 3 14 6 1 16 2 3 15 7 4 19 1 2 C 1 F- /4 g 4 19 1 1 ___14 M 21 1 0 H 5 C 1 0 1 1 20 3 1 1 1 7 26 1 1 C 2 8 1 4 21 3 1 27 1 1 12 1 4. ___ 4 M 22 2 1 28 2 1 C 23 g 3 F-M/8 16 1 2 18 1 2 23 1 2 29 2 1 0 2 2 20 2 1 25 1 2 30 1 2 2 21 2 2 1 1 27 1 2. . . 1 s+2 s 32 5 1 3 3 1 71 2 2 29 1 1 33 2 1 4 3 1 73 1 1 30 1 1 34 2 1 5 74 1 2 1 1 76 2 2 31 1 2 35 1 1 6 1 1 78 2 4 53 H C 11 43 L 23 M _30 L 8 H_. 33 1 6 36 3 1 7 82 2 2 6 1 39 3 L 1 2 M 3 37 3 1 8 84 1 6 1 1 42 1 4 38 3 1 9 8 1 90 2 8 46 1 10 39 5 1 10 2 1 98 1 9 56 2 40 3 1 11 6 1 107 1 16 C 232 g 2 F-M/8 41 7 1 12 2 1 ___6 M 1 2 H 123 0 1 1 42 6 1 13 5 1 1 43 3 1 14 2 1 2 2 1 44 5 1 15 0 L 2 32 M 3 13 H 3 2 L 1 2 M 2 1 H 45 1 1 18 11 L 1 13 M 1 54 H 5 2 1 46 3 1 19 6 1 1 7 2 1 47 3 1 20 1 1 8 2 1 48 4 1 21 3 1 C 111 9 1 4 M 7 3 H ___1 L 49 7 1 22 1 1 F- /4 g 4 16 1 1 50 4 1 23 2 1 17 3 1 0 ___ 1 16 __1 L 51 6 1 24 ___1 L 1 M 4 H 18 2 2 4 1 16 3 1 52 10 1 25 20 7 1 1 2 17 2 1 21 8 1 53 3 1 27 8 1 18 9 1 22 7 1 54 4 1 28 9 1 19 3 2 ___ 23 3 L 1 2 M 2 18 H 55 8 1 29 4 2 21 1 L 5 6 21 M 25 2 1 56 5 1 31 2 1 26 3 1 27 3 1 57 3 1 32 27 2 1 1 1 28 5 1 58 7 1 33 28 3 1 29 14 1 29 1 1 59 2 1 34 3 2 __ 30 1 L 12 M 8 20 H 30 1 1 60 2 1 36 7 1 38 2 2 ___4 L 31 22 M 2 8 H 61 1 1 37 12 2 40 15 1 33 1 1 62 2 2 39 1 1 34 4 1 41 3 4 64 1 1 40 35 3 3 8 H 1 1 ___ 45 3 2 65 2 1 41 38 3 1 1 1 47 2 19 39 8 2 M 11 9 H 66 1 1 ___ 42 6___ 66 3 21 50 2 1 1 H 67 2 48 1 2 2 M ___1 L ___ 87 1 __ 31 H 51 1 2 H CONCRETE C 2 -. 6 . 2 -. 07. 771 6882. . -. 72 . 19 -. 40 . 54 9251. 38 . 14 -. 79 . 46 11781 X g 4 (F-MN)/8 0 2 2 2 1 2 4 2 1 5 1 3 8 2 3 11 1 1 12 3 2 14 4 1 15 3 1 16 3 1 17 2 1 18 3 1 19 6 1 20 3 1 21 3 1 22 2 1 23 5 1 24 4 1 25 3 1 26 6 1 27 3 1 28 1 1 29 6 1 30 3 1 31 2 1 32 3 1 33 3 1 34 1 2 36 3 1 37 1 1 38 2 1 39 3 1 40 5 1 41 1 1 42 6 1 43 1 1 44 3 2 46 5 1 47 1 1 48 3 1 49 1 1 50 2 1 51 1 1 52 1 1 53 1 1 54 1 1 55 1 1 56 3 1 57 3 2 59 1 2 61 1 1 62 3 3 65 2 9 43 L 38 M 55 H C 2 74 1 4 0 L 14 M 0 H C 1 78 1 3 81 1 2 83 1 3 86 1 2 88 1 2 90 1 1 91 1 4 95 1 2 97 1 1 98 1 2 100 1 4 104 1 3 ACCURACY CONCRETE IRIS SEEDS WINE GV 76 82. 7 94 62. 7 MVM 78. 8 94 93. 3 66. 7 GM 83 94. 7 96 81. 3 C 2 gp 8 (F-MN)/5 0 2 2 2 1 2 4 2 1 5 1 3 8 2 3 11 1 1 12 2 2 14 4 1 15 3 1 16 3 1 17 2 1 18 3 1 19 6 1 20 3 1 21 3 1 22 1 1 23 5 1 24 3 1 25 3 1 26 6 1 27 3 1 28 1 1 29 6 1 30 3 1 31 2 1 32 1 1 33 3 1 34 1 2 36 3 2 38 2 1 39 2 1 40 5 1 41 1 1 42 6 1 43 1 1 44 3 2 46 5 1 47 1 1 48 1 1 49 1 1 50 2 1 51 1 1 52 1 1 53 1 1 54 1 1 55 1 1 56 3 1 57 2 2 59 1 2 61 1 1 62 3 3 43 L 65 2 9 28 M 55 H C 21 74 1 4 78 1 8 86 1 2 88 1 2 90 1 5 95 1 2 97 1 1 98 1 2 0 L 100 1 4 10 M 0 H C 22 104 1 C 21 g 4 F-M/4 0 1 1 3 4 1 3 7 2 1 8 2 1 9 1 2 11 1 2 13 4 1 14 2 1 15 4 1 16 1 2 18 2 1 19 3 1 20 1 1 21 2 1 22 6 2 24 2 1 25 3 1 26 1 2 28 2 2 30 1 1 31 1 2 C 21133 1 4 32 L 37 1 1 13 M 0 H 38 2 1 39 2 1 40 1 1 41 1 1 42 1 1 43 2 1 44 1 1 45 2 1 46 1 1 47 1 1 C 21248 1 1 7 L 49 2 2 3 M 10 H 51 2 4 55 1 1 56 8 1 57 4 1 58 4 1 59 2 1 60 1 1 61 1 2 63 5 2 65 1 2 67 2 1 68 1 3 71 1 1 72 4 1 C 21373 8 1 4 L 74 5 1 7 M 38 H 75 1 8 83 3 1 84 3 1 C 214 85 2 1 0 L 86 1 5 M 7 H 99 3 C 211 g 5 F-M)/4 0 1 6 6 2 1 7 2 5 ___5 L. 12 1 1 13 4 1 14 1 1 15 4 2 17 1 1 18 2 1 19 2 2 21 2 1 22 3 1 23 1 1 24 3 4 __20 L 5 M. 28 1 14 42 1 2 44 1 1 45 1 3 48 2 2 ___ 5 L 1 M. 50 1 5 55 1 2 57 1 1 ___ 2 L 1 M. 58 1 5 63 1 1 64 1 7 71 1 11 82 1 16 ___5 L 1 M 98 2 C 212 g 5 F-M/3 0 1 20 20 1 8 28 1 1 29 2 9 38 1 11 49 1 5 54 1 11 __6 L 3 M. 65 1 10 75 2 3 78 1 11 __1 L 2 H 89 1 7 96 1 2 98 1 2 100 1 11 111 2 1 ___ 8 H 112 1
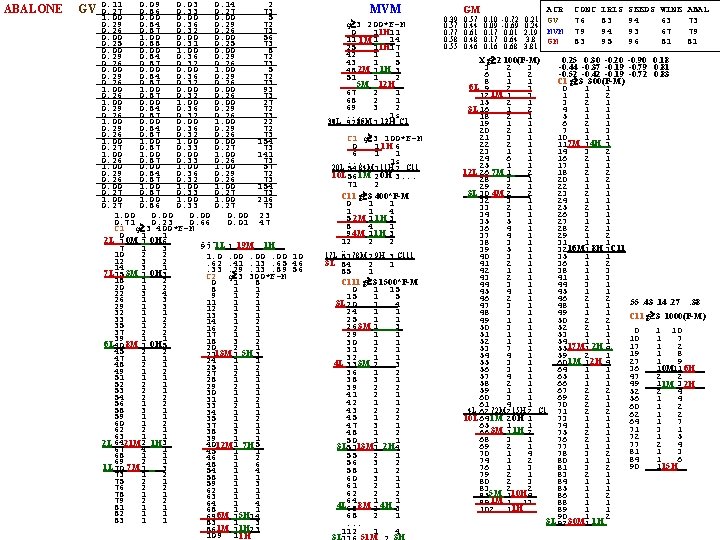
ABALONE GV 0. 11 0. 09 0. 03 0. 14 2 0. 27 0. 86 0. 33 0. 27 73 1. 00 0. 00 5 0. 29 0. 84 0. 36 0. 29 72 0. 26 0. 87 0. 32 0. 26 73 0. 00 1. 00 0. 00 56 0. 25 0. 88 0. 31 0. 25 73 0. 00 1. 00 0. 00 8 0. 29 0. 84 0. 36 0. 29 72 0. 26 0. 87 0. 32 0. 26 73 0. 00 1. 00 5 0. 29 0. 84 0. 36 0. 29 72 0. 26 0. 87 0. 32 0. 26 73 1. 00 0. 00 93 0. 26 0. 87 0. 32 0. 26 73 1. 00 0. 00 27 0. 29 0. 84 0. 36 0. 29 72 0. 26 0. 87 0. 32 0. 26 73 1. 00 0. 00 1. 00 22 0. 29 0. 84 0. 36 0. 29 72 0. 26 0. 87 0. 32 0. 26 73 1. 00 0. 00 154 0. 27 0. 87 0. 33 0. 27 73 1. 00 0. 00 141 0. 26 0. 87 0. 33 0. 26 73 1. 00 0. 00 1. 00 57 0. 29 0. 84 0. 36 0. 29 72 0. 26 0. 87 0. 32 0. 26 73 0. 00 154 0. 27 0. 87 0. 33 0. 27 73 1. 00 216 0. 27 0. 86 0. 33 0. 27 73 1. 00 0. 00 23 0. 71 0. 23 0. 66 0. 01 47 C 1 g 3 400*F-M 0 1 1. . . 2 L 1 0 M 1 0 H 6_ 1 L 19 M 1 H _ 97 1 7 1 3 10 2 2 1. 0. 00. 00 10 12 3 2. 62. 41. 13. 65 46 14 3 1. 33. 29. 13. 89 56 7 L 15 3 M 1 0 H 3_ C 2 g 3 300*F-M 18 1 2 0 1 8 20 1 2 8 1 1 22 3 4 9 1 2 26 1 3 11 1 1 29 1 3 12 1 1 32 1 1 13 3 1 33 1 2 14 1 2 35 1 2 16 2 1 37 2 2 17 1 1 39 1 1 18 3 2 6 L 40 8 M 1 0 H 5_ 20 2 1 45 2 2 2113 M 1 5 H 3_ 47 1 1 24 1 1 48 2 1 25 1 2 49 1 2 27 2 1 51 1 1 28 1 1 52 2 1 29 2 1 53 2 1 30 1 1 54 2 2 31 1 2 56 1 2 33 2 1 58 3 1 34 1 1 59 1 1 35 1 2 60 1 2 37 1 1 62 2 1 38 3 1 63 1 1 39 1 1 2 L 6421 M 2 1 H 3_ 40 1 12 M 7 H 5_ 67 4 1 45 1 1 68 1 1 46 1 2 69 2 1 48 1 6 1 L 70 7 M 1 _3 54 1 4 73 1 2 58 1 1 75 2 1 59 1 3 76 2 2 62 1 1 78 1 1 63 1 1 79 2 2 64 1 4 81 1 1 68 1 1 82 1 1 6 M 5 H _ 69 1 14 83 1 1 83 1 3 _ 861 M 11 H 23 109 1 1 H MVM g 3 200*F-M 0 1 1 H 11 11 1 M 1 14 _ 25 1 1 H 17 42 1 1 43 1 5 2 M 1 H 48 1 3_ 51 1 2 5 M 12 H _. . . 67 2 1 68 2 1 69 3 2. . . 1 s 30 L 92 85 M 1 12 H C 1 g 3 100*F-M 0 1 1 H 6 6 1 1. . . 1 s 20 L 84 M 11 H C 11 54 1 2 10 L 56 1 M 2 0 H 3. . . 71 2 C 11 g 3 400*F-M 0 1 1 4 5 2 M 1 1 H 3_ 8 4 1 9 4 M 1 1 H 3_ 12 2 2. . 17 L 78 M 9 H C 111 81 2 3 3 L 84 2 1 85 1 C 111 g 3 1500*F-M 0 1 15 15 1 5 3 L 20 _4 1 24 1 1 25 1 1 26 3 M 1 3_ 29 1 1 30 1 1 31 2 1 32 1 1 4 L 33 3 M 2 _3 36 1 2 38 3 1 39 2 2 41 2 1 42 1 1 43 2 2 45 1 2 47 3 1 48 1 2 50 1 1 3 L 5113 M 1 2 H 4 55 2 1 56 3 2 58 1 2 60 3 1 61 2 1 62 2 2 64 1 1 4 L 65 8 M 2 4 H 3 68 2 1. . . 112 1 4 3 L 51 M 3 H GM 0. 39 0. 57 0. 10 -0. 72 0. 21 0. 57 0. 44 0. 09 -0. 69 0. 24 0. 77 0. 61 0. 17 0. 01 2. 19 0. 58 0. 48 0. 17 0. 64 3. 8 0. 55 0. 46 0. 16 0. 68 3. 81 ACR GV MVM GM CONC 76 79 83 IRIS 83 94 95 SEEDS 94 93 96 WINE 63 67 81 ABAL 73 79 81 0. 25 0. 30 -0. 20 -0. 90 0. 18 X g 2 100(F-M) 3 2 3 -0. 44 -0. 37 -0. 19 -0. 79 0. 81 6 1 2 -0. 52 -0. 42 -0. 19 -0. 72 0. 83 8 1 1 C 1 g 3 300(F-M) 6 L 9 2 3. 0 1 1 1 M _ 12 1 3 1 1 2 15 2 1 3 2 1 16 1 2 4 1 1 3 L. 18 2 1 5 1 1 19 1 1 6 2 1 20 2 1 7 1 3 21 3 1 10 1 1 7 M 4 H. 22 2 1 11 1 3 23 1 1 14 3 2 24 6 1 16 2 1 25 1 1 17 1 1 12 L 26 1 2 7 M _ 18 2 2 28 3 1 20 1 2 29 2 1 22 1 1 3 L 30 2 2 4 M _ 23 2 1 32 3 1 24 1 1 33 2 1 25 2 1 34 3 1 26 3 1 35 5 1 27 1 1 36 4 1 28 2 1 37 4 1 29 1 2 38 3 1 31 1 1 16 M 8 H C 11 39 5 1 32 1 3 40 3 1 35 1 1 41 2 1 36 1 2 42 1 1 38 1 3 43 2 1 41 1 3 44 3 1 45 4 1 45 1 1 46 2 2. 55. 43. 14. 27. 38 47 3 1 48 1 1 48 3 1 49 1 1 C 11 g 3 1000(F-M) 49 1 1 50 2 2 50 3 1 52 2 1 0 1 10 51 1 1 53 1 1 10 1 7 52 1 1 54 1 1 17 1 2 17 M 2 H. 53 7 1 55 1 4 19 1 8 54 4 1 59 2 1 27 1 9 1 M 2 H _ 55 3 1 60 1 4 0 M 6 H _ 36 1 11 56 3 1 64 1 1 57 4 1 65 1 1 47 2 2 58 2 1 66 1 1 49 1 3 1 M 2 H _ 59 1 1 67 2 2 52 2 4 60 3 1 69 2 1 56 1 4 61 4 1 70 2 1 60 1 2 4 L 72 M 15 H C 1 62 2 2 71 2 2 62 1 2 73 1 1 10 L 64 2 1 1 M 0 H 64 1 7 65 1 1 74 1 1 71 3 1 3 M 1 H _ 66 1 2 75 2 1 72 1 5 68 3 1 76 2 1 77 2 4 69 2 1 77 1 1 81 1 3 70 1 4 78 3 2 84 1 6 74 1 2 80 1 1 15 H _ 90 1 76 1 3 81 3 2 79 2 1 83 2 1 80 2 3 84 1 1 83 2 2 85 1 1 5 M 10 H _ 85 1 4 86 1 2 1 M _ 89 1 13 88 1 1 1 H 102 1 89 1 1 90 1 2 3 L 92 1 30 M 1 H
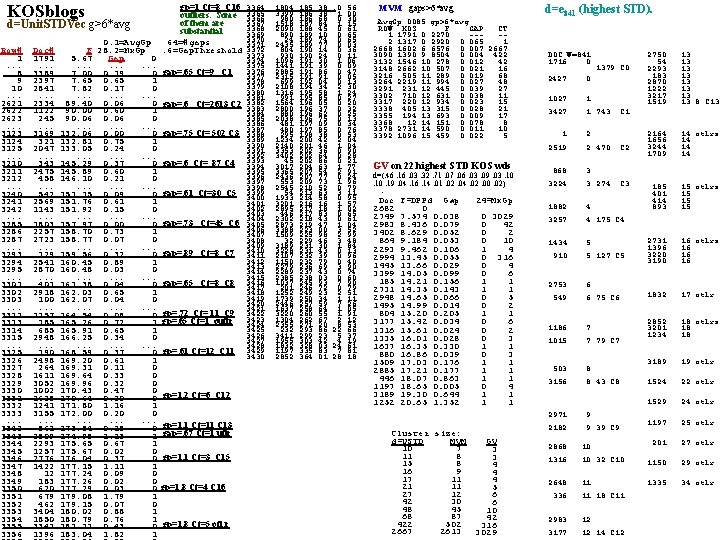
KOSblogs d=Unit. STDVec g>6*avg Row# 1. . . ___8 9 10. . . 2621 ___ 2622 2623. . . 3123 ___ 3124 3125. . . 3210 ___ 3211 3212. . . ___ 3240 3241 3242. . . ___ 3285 3286 3287. . . ___ 3293 3294 3295. . . ___ 3301 3302 3303. . . ___ 3312 ___ 3313 3314 3315. . . ___ 3325 3326 3327 3328 3329 3330 ___ 3331 3332 3333. . . ___ 3342 ___ 3343 3344 3345 ___ 3346 3347 3348 3349 ___ 3350 3351 3352 3353 3354 ___ 3355 3356 Doc# 1791. . . 3389 2397 2841. . . 2334 1122 245. . . 3169 321 2047. . . 343 2475 458. . . 542 2569 1143. . . 1803 2257 2723. . . 129 2541 2870. . . 401 2918 100. . . 1157 185 685 2948. . . 190 2498 264 1611 3052 1002 1628 1241 3155. . . 861 2509 2293 1257 2776 1422 12 183 620 679 462 3404 1850 3342 1396 gp=1 Ct=8 C 16 outliers. Some of them are substantial 64=#gaps. 6=Gap. Threshold 3364 3365 3366 3367 3368 3369 3370 3371 3372 3373 3374 3375 3376 gap=. 65 Ct=9 C 1 3377 3378 3379 3380 3381 gap=. 6 Ct=2613 C 2 3383 3384 3385 3386 3387 gap=. 75 Ct= 502 C 3 3388 3389 3390 3391 3392 3393 gap=. 6 Ct= 87 C 4 3395 3396 3397 3398 gap=. 61 Ct=30 C 5 3399 3400 3401 3402 3403 3404 gap=. 73 Ct=45 C 6 3405 3406 3407 3408 3409 gap=. 89 Ct=8 C 7 3410 3411 3412 3413 3414 3415 gap=. 65 Ct=8 C 8 3416 3417 3418 3419 3420 gp=. 72 Ct= 11 C 9 3421 3422 gp=. 65 Ct=1 outlr 3423 3424 3425 3426 3427 3428 gp=. 61 Ct=12 C 11 3429 3430 0. 1=Avg. Gp F 28. 2=Mx. Gp 5. 67 Gap 0. . ___ 7. 00 0. 19 0 7. 65 0. 65 1 7. 82 0. 17 0. . 89. 40 0. 06 0 ___ 90. 00 0. 60 1 90. 06 0. . 132. 06 0. 00 0 ___ 132. 81 0. 75 1 133. 05 0. 24 0. . 145. 29 0. 37 0 ___ 145. 89 0. 60 1 146. 10 0. 21 0. . ___ 151. 15 0. 09 0 151. 76 0. 61 1 151. 92 0. 15 0. . ___ 157. 97 0. 00 0 158. 70 0. 73 1 158. 77 0. 07 0. . ___ 159. 56 0. 32 0 160. 45 0. 89 1 160. 48 0. 03 0. . ___ 161. 38 0. 04 0 162. 03 0. 65 1 162. 07 0. 04 0. . ___ 164. 54 0. 08 0 ___ 165. 26 0. 72 1 165. 91 0. 65 1 166. 25 0. 34 0. . ___ 168. 59 0. 37 0 169. 20 0. 61 1 169. 31 0. 11 0 169. 64 0. 33 0 169. 96 0. 32 0 170. 43 0. 47 0 ___ 170. 64 0. 20 0 gp=1. 2 Ct=6 C 12 171. 80 1. 16 1 172. 00 0. 20 0. . ___ 173. 84 0. 15 0 gp=1. 1 Ct=11 C 13 ___ 174. 98 1. 13 1 gap=. 67 Ct=1 utlr 175. 65 0. 67 1 175. 67 0. 02 0 ___ 176. 04 0. 37 0 gp=1. 1 Ct=3 C 15 177. 15 1. 11 1 177. 24 0. 09 0 177. 26 0. 02 0 ___ 177. 29 0. 03 0 gp=1. 8 Ct=4 C 16 179. 08 1. 79 1 179. 15 0. 07 0 180. 02 0. 88 1 180. 79 0. 76 1 ___ 181. 21 0. 43 0 gp=1. 8 Ct=5 otl; r 183. 04 1. 82 1 1804 3399 980 1518 2090 890 24 2435 804 930 1096 1441 2885 2315 699 2108 1316 991 1564 2800 880 2038 481 480 295 1234 2140 3353 3402 45 3017 3365 2436 553 2545 54 1933 3201 2895 446 2302 2873 3388 1509 32 3189 3228 2107 1150 2279 2289 2385 1037 201 1252 1739 2446 1637 3220 1304 2355 232 3411 1955 1832 1197 2852 185. 38 186. 68 187. 84 188. 45 189. 10 189. 74 189. 77 190. 14 190. 24 191. 30 191. 39 191. 86 191. 91 192. 04 194. 34 195. 58 195. 85 196. 05 196. 37 196. 62 196. 75 197. 09 197. 85 198. 38 200. 42 201. 46 202. 36 202. 64 202. 86 204. 63 207. 54 207. 77 209. 73 210. 52 213. 63 214. 58 216. 16 217. 18 217. 83 218. 43 219. 47 223. 00 225. 98 229. 46 231. 30 231. 43 232. 39 232. 79 236. 69 237. 43 238. 03 245. 93 246. 72 249. 23 250. 34 257. 59 258. 64 260. 55 262. 67 271. 20 293. 86 299. 23 303. 42 328. 03 335. 83 364. 01 . 0. 56 1. 00 0. 30 1. 15 0. 61 0. 65 0. 03 0. 36 0. 11 1. 06 0. 09 0. 47 0. 05 0. 13 2. 30 1. 24 0. 27 0. 20 0. 32 0. 25 0. 13 0. 34 0. 76 0. 53 2. 04 1. 04 0. 90 0. 28 0. 21 1. 77 2. 91 0. 24 1. 96 0. 79 3. 11 0. 95 1. 57 1. 02 0. 65 0. 61 1. 04 3. 52 2. 99 3. 48 1. 84 0. 13 0. 96 0. 40 3. 90 0. 74 0. 60 7. 90 0. 79 2. 51 1. 11 7. 26 1. 05 1. 91 2. 12 8. 53 22. 66 5. 37 4. 19 24. 61 7. 81 28. 18 0 MVM gaps>6*avg 1 0 Avg. Gp. 0085 gp>6*avg 1 ROW KOS F GAP CT 1 1791 0. 2270 ---1 2 1317 0. 2920 0. 065 1 0 2668 1602 6. 6576 0. 007 2667 0 3090 1390 9. 8504 0. 004 422 0 1 3132 1546 10. 278 0. 012 42 0 3148 2662 10. 507 0. 021 16 0 3216 505 11. 289 0. 019 68 0 0 3264 2219 11. 994 0. 027 48 1 3291 231 12. 445 0. 039 27 1 3302 710 12. 631 0. 038 11 0 3317 220 12. 934 0. 023 15 0 0 3338 405 13. 315 0. 028 21 0 3355 194 13. 693 0. 009 17 0 3368 12 14. 151 0. 078 8 0 3378 2731 14. 590 0. 011 10 1 0 3392 1096 15. 459 0. 022 5 1 1 1 0 0 1 on 22 highest STD KOS wds GV 1 d=(. 46. 16. 03. 32. 71. 07. 06. 03. 09. 03. 10 0 1. 10. 19. 04. 16. 14. 01. 02. 04. 02. 00. 02) 1 1 1 Doc F=DPPd Gap 24=Mx. Gp 1 1 2682 0 1 2749 7. 574 0. 038 0 3029 1 1 2983 8. 436 0. 079 0 42 1 3402 8. 629 0. 052 0 2 1 1 864 9. 184 0. 053 0 10 1 2293 9. 462 0. 106 1 4 0 1 2994 13. 45 0. 055 0 316 0 1445 13. 66 0. 029 0 4 1 1 3399 14. 05 0. 099 0 6 0 185 14. 21 0. 156 1 1 1 1 2731 14. 35 0. 143 1 1 1 2948 14. 65 0. 066 0 5 1 1 1495 14. 99 0. 014 0 2 1 804 15. 20 0. 205 1 1 1 1 3177 15. 42 0. 034 0 6 1 1 1316 15. 61 0. 024 0 2 1 1335 16. 01 0. 028 0 3 1 1 1637 16. 35 0. 330 1 1 1 880 16. 86 0. 039 0 3 1 1509 17. 03 0. 176 1 1 2885 17. 21 0. 177 1 1 446 18. 07 0. 863 1 1 1197 18. 65 0. 005 0 4 3189 19. 30 0. 644 1 1 1252 20. 65 1. 352 1 1 Cluster size: d=USTD MVM 10 7 11 8 15 8 16 9 17 11 21 11 27 12 42 30 48 45 68 87 422 502 2667 2613 GV 3 3 4 4 4 5 6 6 10 42 316 3029 d=e 841 (highest STD). DOC W=841 1716 0. . . 1379 C 0 2427 0 2750 54 2293 183 2870 1222 3217 1519 13 13 8 C 13 2164 1656 3244 1709 14 otlrs 14 14 14 185 401 414 893 15 otlrs 15 15 15 1027. . . 3427 1. . . 1 743 1. . . 2519 2. . . 2 470 C 2 868. . . 3224 3. . . 3 274 C 3 1882. . . 3257 4. . . 4 175 C 4 1434. . . 910 5. . . 5 127 C 5 2731 1396 3220 3190 16 otlrs 16 16 16 2753. . . 549 6. . . 6 75 C 6 1832 17 otlr 1186. . . 1015 7. . . 7 79 C 7 2852 3201 1234 18 otlrs 18 18 503. . . 3156 8. . . 8 43 C 8 3189 19 otlr 1524 22 otlr 1529 24 otlr 2971. . . 2182 9. . . 9 39 C 9 1197 25 otlr 2868. . . 1316 10. . . 10 32 C 10 201 27 otlr 1150 29 otlr 2648. . . 336 11. . . 11 18 C 11 1335 34 otlr 2983. . . 3177 12. . . 12 14 C 12 C 1
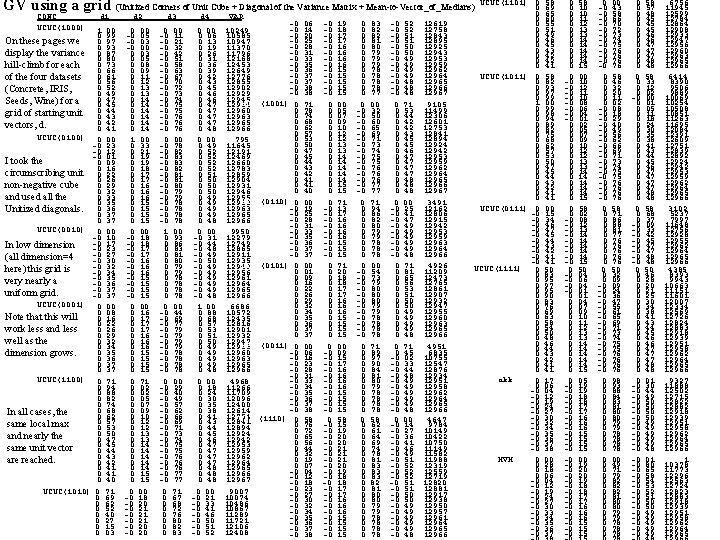
GV using a grid (Unitized Corners of Unit Cube + Diagonal of the Variance Matrix + Mean-to-Vector_of_Medians) CONC d 1 UCUC(1000) 1. 00 0. 99 On these pages we 0. 97 0. 93 display the variance 0. 87 0. 80 hill-climb for each 0. 73 0. 66 of the four datasets 0. 61 0. 56 0. 52 (Concrete, IRIS, 0. 49 Seeds, Wine) for a 0. 47 0. 45 grid of starting unit 0. 44 0. 43 0. 42 vectors, d. 0. 41 UCUC(0100) 0. 00 -0. 23 -0. 12 -0. 01 I took the 0. 09 circumscribing unit 0. 16 0. 22 0. 26 non-negative cube 0. 29 0. 32 0. 33 and used all the 0. 35 Unitized diagonals. 0. 36 0. 37 UCUC(0010) 0. 00 -0. 10 In low dimension -0. 17 -0. 23 -0. 27 (all dimension=4 -0. 30 -0. 32 here) this grid is -0. 34 -0. 35 very nearly a -0. 36 -0. 37 uniform grid. -0. 37 UCUC(0001) 0. 00 0. 08 Note that this will 0. 16 0. 22 work less and less 0. 26 0. 29 0. 32 well as the 0. 34 0. 35 dimension grows. 0. 36 0. 37 UCUC(1100) 0. 71 0. 94 0. 88 0. 82 0. 74 0. 68 In all cases, the 0. 62 0. 57 same local max 0. 53 0. 50 and nearly the 0. 47 0. 45 same unit vector 0. 44 0. 43 are reached. 0. 42 0. 41 0. 40 UCUC(1010) 0. 71 0. 69 0. 62 0. 52 0. 40 0. 27 0. 15 0. 03 d 2 d 3 d 4 0. 00 -0. 05 -0. 03 -0. 00 0. 03 0. 05 0. 08 0. 09 0. 11 0. 12 0. 13 0. 14 1. 00 0. 33 0. 21 0. 19 0. 18 0. 17 0. 16 0. 15 0. 00 -0. 18 -0. 17 -0. 16 -0. 15 0. 00 0. 16 0. 17 0. 16 0. 15 0. 71 0. 02 0. 05 0. 07 0. 09 0. 10 0. 12 0. 13 0. 14 0. 15 0. 00 -0. 11 -0. 21 -0. 32 -0. 42 -0. 51 -0. 58 -0. 63 -0. 67 -0. 70 -0. 72 -0. 73 -0. 74 -0. 75 -0. 76 0. 00 -0. 78 -0. 82 -0. 83 -0. 82 -0. 81 -0. 80 -0. 79 -0. 78 1. 00 0. 93 0. 86 0. 83 0. 81 0. 80 0. 79 0. 78 0. 00 -0. 44 -0. 69 -0. 77 -0. 79 -0. 78 0. 00 -0. 29 -0. 40 -0. 49 -0. 57 -0. 62 -0. 66 -0. 69 -0. 71 -0. 73 -0. 74 -0. 75 -0. 76 -0. 77 0. 00 0. 06 0. 13 0. 19 0. 26 0. 31 0. 36 0. 39 0. 42 0. 43 0. 45 0. 46 0. 47 0. 48 0. 00 0. 49 0. 52 0. 51 0. 50 0. 49 0. 48 0. 00 -0. 31 -0. 44 -0. 48 -0. 49 -0. 50 -0. 49 -0. 48 1. 00 0. 88 0. 68 0. 57 0. 53 0. 51 0. 50 0. 49 0. 48 0. 00 0. 18 0. 24 0. 30 0. 35 0. 38 0. 41 0. 43 0. 44 0. 45 0. 46 0. 47 0. 48 0. 00 -0. 18 -0. 20 -0. 21 -0. 20 0. 71 0. 67 0. 68 0. 72 0. 76 0. 80 0. 82 0. 83 0. 00 -0. 21 -0. 33 -0. 41 -0. 46 -0. 50 -0. 51 -0. 52 VAR -0. 06 -0. 14 10249 -0. 20 10585 -0. 25 10947 -0. 28 11370 -0. 31 11796 -0. 33 12168 -0. 35 12453 -0. 36 12649 -0. 37 12776 -0. 37 12855 -0. 38 12902 -0. 38 12929 12945 UCUC(1001) 0. 71 12954 0. 78 12960 0. 74 12963 0. 68 12965 0. 62 12966 0. 57 0. 53 795 0. 50 11645 0. 47 12191 0. 45 12469 0. 44 12660 0. 43 12783 0. 42 12859 0. 41 12904 0. 41 12931 0. 40 12946 12955 UCUC(0110) 0. 00 12960 -0. 19 12963 -0. 25 12965 -0. 28 12966 -0. 31 -0. 33 9950 -0. 35 12279 -0. 36 12749 -0. 37 12865 -0. 37 12911 12935 UCUC(0101) 0. 00 12949 0. 01 12956 0. 09 12961 0. 16 12964 0. 22 12965 0. 26 12966 0. 29 0. 32 6686 0. 34 10572 0. 35 12435 0. 36 12816 0. 37 12901 0. 37 12932 12947 UCUC(0011) 0. 00 12955 -0. 06 12960 -0. 16 12963 -0. 23 12965 -0. 28 12966 -0. 31 -0. 33 4968 -0. 34 11266 -0. 35 11709 -0. 36 12096 -0. 37 12400 -0. 38 12614 12754 UCUC(1110) 0. 58 12841 0. 76 12894 0. 72 12924 0. 65 12942 0. 56 12953 0. 44 12959 0. 32 12962 0. 19 12964 0. 07 12965 -0. 04 12966 -0. 12 12967 -0. 18 -0. 23 9007 -0. 27 10074 -0. 30 10486 -0. 32 10867 -0. 34 11289 -0. 35 11721 -0. 36 12106 -0. 37 12408 -0. 38 -0. 19 -0. 18 -0. 17 -0. 16 -0. 15 0. 00 0. 05 0. 07 0. 09 0. 10 0. 12 0. 13 0. 14 0. 15 0. 71 -0. 13 -0. 17 -0. 16 -0. 15 0. 71 0. 20 0. 18 0. 17 0. 16 0. 15 0. 00 -0. 09 -0. 15 -0. 17 -0. 16 -0. 15 0. 58 -0. 15 -0. 19 -0. 20 -0. 21 -0. 20 -0. 19 -0. 18 -0. 17 -0. 16 -0. 15 0. 83 0. 82 0. 81 0. 80 0. 79 0. 78 0. 77 0. 00 -0. 32 -0. 50 -0. 65 -0. 69 -0. 71 -0. 73 -0. 74 -0. 75 -0. 76 -0. 77 0. 71 0. 94 0. 86 0. 82 0. 80 0. 79 0. 78 0. 00 -0. 54 -0. 73 -0. 79 -0. 80 -0. 79 -0. 78 0. 71 0. 89 0. 97 0. 90 0. 84 0. 81 0. 80 0. 79 0. 78 0. 58 0. 62 0. 61 0. 64 0. 69 0. 74 0. 78 0. 81 0. 83 0. 82 0. 81 0. 80 0. 79 0. 78 -0. 52 -0. 51 -0. 50 -0. 49 -0. 48 0. 71 0. 53 0. 44 0. 42 0. 43 0. 45 0. 46 0. 47 0. 48 0. 00 -0. 25 -0. 41 -0. 47 -0. 49 -0. 48 0. 71 0. 81 0. 65 0. 56 0. 53 0. 51 0. 50 0. 49 0. 48 0. 71 0. 45 -0. 02 -0. 33 -0. 44 -0. 48 -0. 49 -0. 48 0. 00 -0. 14 -0. 27 -0. 36 -0. 41 -0. 46 -0. 49 -0. 51 -0. 52 -0. 51 -0. 50 -0. 49 -0. 48 12619 12758 12843 12895 12925 12943 12959 12962 12964 12965 12966 12967 9105 11499 12306 12601 12753 12841 12894 12924 12942 12953 12959 12962 12964 12965 12966 12967 3491 12162 12806 12915 12942 12953 12959 12963 12964 12966 4926 11209 12473 12765 12861 12907 12932 12947 12955 12960 12963 12965 12966 4951 6835 10755 12547 12876 12934 12951 12958 12962 12964 12965 12966 4647 9784 10149 10422 10750 11149 11582 11988 12319 12559 12719 12820 12881 12917 12938 12950 12957 12961 12964 12965 12966 UCUC(1101) 0. 58 0. 69 0. 65 0. 60 0. 55 0. 51 0. 49 0. 46 0. 45 0. 43 0. 42 0. 41 UCUC(1011) 0. 58 0. 82 0. 93 0. 97 0. 99 1. 00 0. 99 0. 98 0. 94 0. 89 0. 82 0. 75 0. 68 0. 62 0. 57 0. 53 0. 50 0. 47 0. 45 0. 44 0. 43 0. 42 0. 41 UCUC(0111) 0. 00 -0. 15 -0. 34 -0. 46 -0. 47 -0. 45 -0. 44 -0. 43 -0. 42 -0. 41 UCUC(1111) 0. 50 0. 83 0. 95 0. 97 0. 95 0. 90 0. 83 0. 76 0. 69 0. 63 0. 58 0. 54 0. 50 0. 48 0. 46 0. 44 0. 43 0. 42 0. 41 akk 0. 17 0. 06 -0. 04 -0. 12 -0. 19 -0. 24 -0. 27 -0. 30 -0. 32 -0. 34 -0. 35 -0. 36 -0. 37 -0. 38 MVM 0. 00 0. 28 0. 18 0. 06 -0. 04 -0. 12 -0. 19 -0. 24 -0. 27 -0. 30 -0. 33 -0. 34 -0. 35 -0. 36 -0. 37 0. 58 0. 10 0. 11 0. 12 0. 13 0. 14 0. 15 0. 00 -0. 12 -0. 11 -0. 10 -0. 08 -0. 06 -0. 04 -0. 01 0. 02 0. 05 0. 07 0. 09 0. 10 0. 12 0. 13 0. 14 0. 15 0. 58 0. 02 -0. 08 -0. 12 -0. 13 -0. 14 -0. 15 0. 50 -0. 04 -0. 06 -0. 04 -0. 01 0. 04 0. 07 0. 09 0. 10 0. 11 0. 12 0. 13 0. 14 0. 15 0. 05 -0. 19 -0. 18 -0. 17 -0. 16 -0. 15 -0. 00 -0. 19 -0. 20 -0. 19 -0. 18 -0. 17 -0. 16 -0. 15 0. 00 -0. 43 -0. 58 -0. 66 -0. 70 -0. 72 -0. 73 -0. 74 -0. 75 -0. 76 0. 58 0. 46 0. 32 0. 20 0. 11 0. 02 -0. 08 -0. 18 -0. 29 -0. 40 -0. 49 -0. 56 -0. 62 -0. 66 -0. 69 -0. 71 -0. 73 -0. 74 -0. 75 -0. 76 0. 58 0. 71 0. 86 0. 88 0. 81 0. 77 0. 76 0. 50 0. 32 0. 09 -0. 24 -0. 36 -0. 47 -0. 55 -0. 61 -0. 65 -0. 69 -0. 71 -0. 73 -0. 74 -0. 75 -0. 76 0. 98 0. 93 0. 88 0. 84 0. 83 0. 81 0. 80 0. 79 0. 78 0. 00 0. 49 0. 71 0. 79 0. 82 0. 81 0. 80 0. 79 0. 78 0. 57 0. 48 0. 45 0. 46 0. 47 0. 48 0. 58 0. 33 0. 12 0. 02 -0. 00 0. 01 0. 05 0. 11 0. 18 0. 24 0. 30 0. 35 0. 38 0. 41 0. 43 0. 44 0. 45 0. 46 0. 47 0. 48 0. 58 0. 68 0. 37 -0. 09 -0. 33 -0. 42 -0. 45 -0. 47 -0. 48 0. 50 0. 46 0. 28 0. 20 0. 21 0. 25 0. 30 0. 34 0. 38 0. 41 0. 43 0. 44 0. 45 0. 46 0. 47 0. 48 0. 01 -0. 30 -0. 44 -0. 49 -0. 50 -0. 49 -0. 48 -0. 01 -0. 80 -0. 65 -0. 58 -0. 54 -0. 53 -0. 52 -0. 51 -0. 50 -0. 49 6756 11945 12599 12784 12864 12908 12933 12947 12956 12960 12963 12965 12966 6414 8390 9506 9889 10069 10254 10508 10851 11263 11695 12084 12391 12609 12751 12839 12892 12924 12942 12953 12959 12962 12964 12965 12966 3102 5237 7997 11648 12756 12928 12955 12962 12964 12965 12966 4385 8393 9943 10663 11151 11601 12007 12334 12569 12726 12824 12883 12918 12939 12951 12958 12962 12964 12965 12966 9327 11888 12502 12715 12822 12882 12918 12939 12951 12958 12962 12964 12965 12966 1 10378 11773 12296 12563 12724 12823 12883 12918 12939 12951 12958 12962 12964 12965
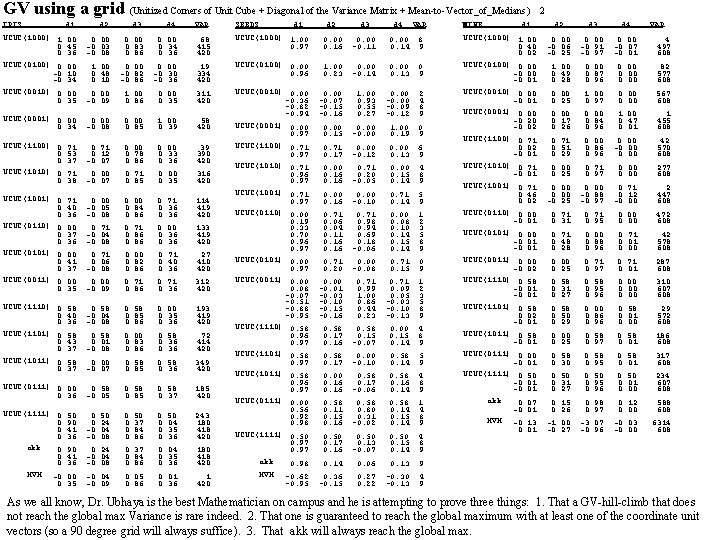
GV using a grid (Unitized Corners of Unit Cube + Diagonal of the Variance Matrix + Mean-to- Vector_of_Medians) 2 IRIS d 1 d 2 d 3 d 4 VAR UCUC(1000) 1. 00 0. 45 0. 36 UCUC(0100) 0. 00 -0. 10 -0. 34 UCUC(0010) 0. 00 0. 35 0. 00 -0. 03 -0. 08 0. 00 0. 83 0. 86 0. 00 0. 34 0. 36 68 415 420 1. 00 0. 48 0. 10 0. 00 -0. 82 -0. 86 0. 00 -0. 36 19 334 420 0. 00 -0. 09 1. 00 0. 86 0. 00 0. 35 311 420 UCUC(0001) 0. 00 0. 34 0. 00 -0. 08 0. 00 0. 85 1. 00 0. 39 58 420 UCUC(1100) 0. 71 0. 53 0. 37 UCUC(1010) 0. 71 0. 38 0. 71 0. 12 -0. 07 0. 00 0. 78 0. 86 0. 00 0. 33 0. 36 39 390 420 0. 00 -0. 07 0. 71 0. 85 0. 00 0. 35 316 420 UCUC(1001) 0. 71 0. 40 0. 36 UCUC(0110) 0. 00 0. 37 0. 36 UCUC(0101) 0. 00 0. 41 0. 37 UCUC(0011) 0. 00 0. 35 0. 00 -0. 05 -0. 08 0. 00 0. 84 0. 86 0. 71 0. 36 114 419 420 0. 71 -0. 04 -0. 08 0. 71 0. 86 0. 00 0. 36 133 419 420 0. 71 0. 06 -0. 08 0. 00 0. 82 0. 86 0. 71 0. 40 0. 36 27 410 420 0. 00 -0. 09 0. 71 0. 86 0. 71 0. 36 312 420 UCUC(1110) 0. 58 0. 40 0. 36 UCUC(1101) 0. 58 0. 43 0. 37 UCUC(1011) 0. 58 0. 37 0. 58 -0. 04 -0. 08 0. 58 0. 85 0. 86 0. 00 0. 35 0. 36 193 419 420 0. 58 0. 01 -0. 08 0. 00 0. 83 0. 86 0. 58 0. 36 72 414 420 0. 00 -0. 07 0. 58 0. 85 0. 58 0. 36 349 420 UCUC(0111) 0. 00 0. 36 0. 58 -0. 05 0. 58 0. 85 0. 58 0. 37 185 420 UCUC(1111) 0. 50 0. 90 0. 41 0. 36 akk 0. 90 0. 41 0. 36 MVM -0. 00 0. 35 0. 50 0. 24 -0. 08 0. 50 0. 37 0. 84 0. 86 0. 50 0. 04 0. 35 0. 36 243 180 418 420 0. 24 -0. 08 0. 37 0. 84 0. 86 0. 04 0. 35 0. 36 180 418 420 -0. 04 -0. 09 0. 05 0. 86 0. 01 0. 36 1 420 SEEDS d 1 d 2 d 3 d 4 VAR UCUC(1000) 1. 00 0. 00 8 0. 97 0. 16 -0. 11 0. 14 9 UCUC(0100) 0. 00 1. 00 0. 00 0 0. 96 0. 23 -0. 14 0. 13 9 UCUC(0010) 0. 00 1. 00 0. 00 2 -0. 36 -0. 07 0. 93 -0. 00 4 -0. 82 -0. 15 0. 55 -0. 09 8 -0. 94 -0. 16 0. 27 -0. 12 9 UCUC(0001) 0. 00 1. 00 0 0. 97 0. 15 -0. 00 0. 19 9 UCUC(1100) 0. 71 0. 00 6 0. 97 0. 17 -0. 12 0. 13 9 UCUC(1010) 0. 71 0. 00 4 0. 96 0. 16 0. 20 0. 15 8 0. 97 0. 16 -0. 05 0. 14 9 UCUC(1001) 0. 71 0. 00 0. 71 5 0. 97 0. 16 -0. 10 0. 14 9 UCUC(0110) 0. 00 0. 71 0. 00 1 0. 19 0. 06 0. 98 0. 08 2 0. 33 0. 04 0. 94 0. 10 3 0. 70 0. 11 0. 69 0. 14 5 0. 96 0. 18 0. 15 8 0. 97 0. 16 -0. 06 0. 14 9 UCUC(0101) 0. 00 0. 71 0 0. 97 0. 20 -0. 08 0. 15 9 UCUC(0011) 0. 00 0. 71 1 0. 08 -0. 01 0. 99 0. 09 2 -0. 07 -0. 03 1. 00 0. 05 3 -0. 51 -0. 10 0. 86 -0. 03 5 -0. 88 -0. 15 0. 44 -0. 10 8 -0. 95 -0. 16 0. 23 -0. 13 9 UCUC(1110) 0. 58 0. 00 4 0. 96 0. 17 0. 15 8 0. 97 0. 16 -0. 07 0. 14 9 UCUC(1101) 0. 58 0. 00 0. 58 5 0. 97 0. 17 -0. 10 0. 14 9 UCUC(1011) 0. 58 0. 00 0. 58 4 0. 96 0. 17 0. 16 8 0. 97 0. 16 -0. 06 0. 14 9 UCUC(0111) 0. 00 0. 58 1 0. 56 0. 11 0. 80 0. 14 4 0. 92 0. 15 0. 31 0. 15 8 0. 98 0. 16 -0. 02 0. 14 9 UCUC(1111) 0. 50 4 0. 97 0. 13 0. 15 8 0. 97 0. 16 -0. 07 0. 14 9 akk 0. 98 0. 14 0. 06 0. 13 9 MVM WINE d 1 UCUC(1000) 1. 00 0. 40 0. 02 UCUC(0100) 0. 00 -0. 01 UCUC(0010) 0. 00 -0. 01 UCUC(0001) 0. 00 -0. 20 -0. 02 UCUC(1100) 0. 71 0. 02 -0. 01 UCUC(1010) 0. 71 -0. 01 UCUC(1001) 0. 71 0. 46 0. 02 UCUC(0110) 0. 00 -0. 01 UCUC(0101) 0. 00 -0. 01 UCUC(0011) 0. 00 -0. 02 UCUC(1110) 0. 58 -0. 01 UCUC(1101) 0. 58 0. 02 -0. 01 UCUC(1011) 0. 58 -0. 01 UCUC(0111) 0. 00 -0. 01 UCUC(1111) 0. 50 -0. 01 akk 0. 07 -0. 01 MVM -0. 13 0. 01 d 2 d 3 d 4 VAR 0. 00 -0. 06 -0. 25 0. 00 -0. 91 -0. 97 0. 00 -0. 07 -0. 01 4 497 608 1. 00 0. 49 0. 28 0. 00 0. 87 0. 96 0. 00 82 577 608 0. 00 0. 25 1. 00 0. 97 0. 00 567 608 0. 00 0. 17 0. 26 0. 00 0. 84 0. 96 1. 00 0. 47 0. 01 1 455 608 0. 71 0. 51 0. 29 0. 00 0. 86 0. 96 0. 00 -0. 00 42 570 608 0. 00 0. 25 0. 71 0. 97 0. 00 277 608 0. 00 -0. 25 0. 00 -0. 88 -0. 97 0. 71 0. 12 -0. 00 2 447 608 0. 71 0. 31 0. 71 0. 95 0. 00 472 608 0. 71 0. 48 0. 28 0. 00 0. 88 0. 96 0. 71 0. 00 42 578 608 0. 00 0. 25 0. 71 0. 97 0. 71 0. 01 287 608 0. 58 0. 31 0. 27 0. 58 0. 95 0. 96 0. 00 310 607 608 0. 50 0. 29 0. 00 0. 86 0. 96 0. 58 0. 01 0. 00 29 572 608 0. 00 0. 25 0. 58 0. 97 0. 58 0. 01 186 608 0. 58 0. 30 0. 58 0. 95 0. 58 0. 01 317 608 0. 50 0. 31 0. 27 0. 50 0. 95 0. 96 0. 50 0. 01 0. 00 234 607 608 0. 15 0. 26 0. 98 0. 97 0. 12 0. 00 588 608 -1. 00 -0. 27 -3. 07 -0. 96 -0. 03 -0. 00 6314 608 -0. 62 0. 36 0. 27 -0. 30 4 -0. 95 -0. 15 0. 22 -0. 13 9 As we all know, Dr. Ubhaya is the best Mathematician on campus and he is attempting to prove three things: 1. That a GV-hill-climb that does not reach the global max Variance is rare indeed. 2. That one is guaranteed to reach the global maximum with at least one of the coordinate unit vectors (so a 90 degree grid will always suffice). 3. That akk will always reach the global max.

Finding round clusters that aren't DPPd separable? (no linear gap) Find the golf ball? Suppose we have a white mask p. Tree. No linear gaps exits to reveal it. Search a grid of d-tubes until a DPPd gap is found in the interior of the tube (Form mask p. Tree for interior of the d-tube. Apply DPPd that mask to reveal interior gaps. ) Look for conical gaps (fix the cone point at the middle of tube) over all cone angles (look for an interval of angles with no points). Notice that this method includes DPPd since a gap for a cone angle of 90 degrees is linear. d

Width 24 so compute all p. Tree combinations down to p 4 and p'4 d=M-p FAUST Gap Revealer Z p= z 1 1 1 z 2 3 1 z 3 2 2 z 4 3 3 z 5 6 2 z 6 9 3 z 7 15 1 z 8 14 2 z 9 15 3 za 13 4 zb 10 9 zc 11 10 zd 9 11 ze 11 11 zf 7 8 F=zod 11 27 23 34 53 80 118 114 125 114 110 121 109 125 83 p 6 0 0 0 1 1 1 1 1 p 5 0 0 0 1 1 1 1 1 0 p 4 0 1 1 1 1 0 1 1 p 3 1 1 0 0 0 1 0 1 1 0 p 2 0 0 1 0 1 0 1 1 0 p 1 1 1 0 0 0 1 p 0 1 1 1 0 0 0 1 1 1 1 p 6' 1 1 1 0 0 0 0 0 p 5' 1 1 1 0 0 0 0 0 1 p 4' 1 0 0 0 0 1 0 0 p 3' 0 0 1 1 1 0 1 0 0 1 p 2' 1 1 0 1 0 1 0 0 1 p 1' 0 0 1 1 1 0 p 0' 0 0 0 1 1 1 0 0 0 0 1 z 2 z 7 2 z 3 z 5 z 8 3 z 4 z 6 z 9 4 za 5 M 6 7 8 zf 9 zb a zc b zd ze c 0 1 2 3 4 5 6 7 8 9 a b c d e f p 4' 0 p 6' 0 0 p 5' 1 p &p &p 6' 5' 4' 0 1 1 0 [000 0000, 000 1111]= [0, 15]=[0, 16) 1 1 4 thinning. 0 0 has 1 point, z 1. This is a 2 0 1 1 0 z 1 o d=11 is only 5 units from the right 0 1 1 0 0 0 1 edge, so z 1 is not declared an outlier) 0 0 0 Next, we check the min dis from the 0 1 0 0 right edge of the next interval to see if 0 0 1 4 (the 1 0 z 1's right-side gap is actually 2 0 0 0 1 calculation of the min is a p. Tree 0 0 1 process - no x looping required!) C=5 C=3 C=1 p 4' p 6' p 5' 0 0 1 1 1 1 0 0 1 [001 0000, 001 1111] = [16, 32). The 1 1 1 [010 0000 , 010 1111] = [32, 48). 0 minimum, z 3 od=23 is 7 units from the 0 1 1 1 1 0 0 z 4 od=34 is within 2 of 32, so z 4 1 1 0 0 0 left edge, 16, so z 1 has only a 5+7=12 0 is not declared an anomaly. 1 1 0 0 unit gap on its right (not a 2 0 4 gap). So z 1 0 1 1 0 0 1 1 4 (and is declared a 2 4 is not declared a 2 1 1 0 0 1 1 1 inlier). 1 0 0 0 0 1 1 1 0 0 1 1 1 0 0 1 C=5 C=2 C=3 C=2 C=1 p 4' p 6 p &p ' 4' p 5' 6 1 5 0 0 1 [100 0000 , 100 1111]= [64, 80). 1 0 0 1 1 4 gap. 1 This is clearly a 2 0 1 0 0 0 1 1 0 0 1 1 0 0 1 1 p 4 p 6 p 5' 0 0 1 1 1 [101 0000 , 101 1111]= [80, 96). 0 0 1 1 0 0 1 0 z 6 od=80, zfod=83 0 1 X 2 d. X 1 X 2 0 1 0 X 1 X 2 d. X 1 X 2 1 z 7 z 8 1. 4 1 z 9 z 10 2. 2 1 0 z 7 z 9 2. 0 1 0 z 9 z 11 7. 8 1 0 z 7 z 10 3. 6 1 0 z 9 z 12 8. 1 1 0 z 7 z 11 9. 4 1 0 z 9 z 13 10. 0 1 0 z 7 z 12 9. 8 1 0 z 9 z 14 8. 9 0 z 7 z 13 11. 7 1 1 0 0 z 11 5. 8 0 z 7 z 14 10. 8 1 0 1 z 10 z 12 6. 3 1 1 0 z 8 z 9 1. 4 0 z 13 8. 1 1 z 8 z 10 2. 2 1 C 10 z 8 z 11 8. 1 z 10 z 14 7. 3 C=2 z 8 z 12 8. 5 z 8 z 13 10. 3 z 8 z 14 9. 5 C 10 C=2 C=0 p 4 p 6' p 5 0 0 1 1 [011 0000, 011 1111] = [ 48, 64). 1 1 0 0 1 1 0 z 5 od=53 is 19 from z 4 0 od=34 (>24) 0 0 1 1 1 but 11 from 64. But the next int 1 1 1 0 [64, 80) is empty z 5 is 27 from its 0 1 1 0 0 right nbr. z 5 is declared an outlier 1 1 0 0 1 and we put a subcluster cut thru z 5 1 0 0 1 1 1 0 0 1 [111 0000 , 1111]= [112, 128) 0 0 1 z 7 od=118 1 z 8 od=114 1 0 0 1 1 z 9 od=125 1 0 0 zaod=114 C=5 zcod=121 zeod=125 C=2 C=1 No 24 gaps. But we can consult Sp. S(d 2(x, y) for actual distances: p 6 p 4' p 5 0 1 1 0 0 0 [110 0000 , 110 1111]= [96, 112). 0 0 zbod=110, zdo 1 d=109. So both 0 0 1 1 {z 6, zf} declared outliers (gap 16 1 1 0 both sides. 0 1 0 0 1 X 2 d. X 1 X 2 1 0 0 1 z 11 z 12 1. 4 1 0 0 1 z 13 2. 2 1 1 z 14 2. 2 1 1 0 0 1 1 z 12 z 13 2. 2 1 1 0 0 z 12 z 14 1. 0 C 10 C=8 C=2 z 13 z 14 2. 0 p 6 p 4 p 5 0 0 1 1 0 0 1 Which reveals that there are 1 0 0 1 1 no 24 gaps in this 1 1 0 0 1 1 subcluster. 1 1 And, incidentally, it reveals 1 1 0 0 a 5. 8 gap between {7, 8, 9, a} 1 1 and {b, c, d, e} but that 1 1 0 0 1 1 analysis is messy and the 1 1 0 gap would be revealed by 0 C 10 the next x o f. M round on this C=8 C=6 sub-cluster anyway.
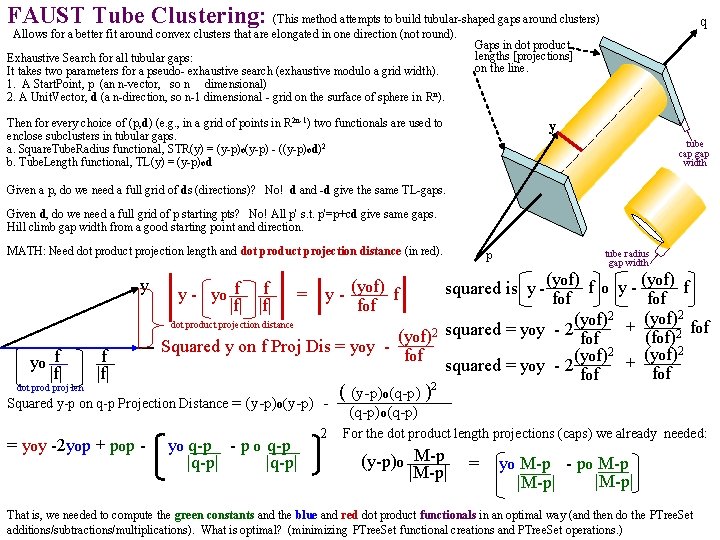
FAUST Tube Clustering: (This method attempts to build tubular-shaped gaps around clusters) Allows for a better fit around convex clusters that are elongated in one direction (not round). Exhaustive Search for all tubular gaps: It takes two parameters for a pseudo- exhaustive search (exhaustive modulo a grid width). 1. A Start. Point, p (an n-vector, so n dimensional) 2. A Unit. Vector, d (a n-direction, so n-1 dimensional - grid on the surface of sphere in Rn). q Gaps in dot product lengths [projections] on the line. Then for every choice of (p, d) (e. g. , in a grid of points in R 2 n-1) two functionals are used to enclose subclusters in tubular gaps. a. Square. Tube. Radius functional, STR(y) = (y-p)o(y-p) - ((y-p)od)2 b. Tube. Length functional, TL(y) = (y-p)od y tube cap gap width Given a p, do we need a full grid of ds (directions)? No! d and -d give the same TL-gaps. Given d, do we need a full grid of p starting pts? No! All p' s. t. p'=p+cd give same gaps. Hill climb gap width from a good starting point and direction. MATH: Need dot product projection length and dot product projection distance (in red). y yo f |f| p tube radius gap width (yof) f o y - (yof) f squared is y - fof (yof)2 + f dot product projection distance (yof)2 squared = yoy - 2 fof (f of)2 Squared y on f Proj Dis = y o y f (yof)2 + fof squared = yoy - 2 fof y - yo f f |f| = y - (yof) f fof ( (y-p)o(q-p) )2 Squared y-p on q-p Projection Distance = (y-p)o(y-p) (q-p)o(q-p) dot prod proj len 1 st = yoy -2 yop + pop - ( yo(q-p) - p o(q-p |q-p| 2 For the dot product length projections (caps) we already needed: (y-p)o M-p = ( yo(M-p) - po M-p ) |M-p| That is, we needed to compute the green constants and the blue and red dot product functionals in an optimal way (and then do the PTree. Set additions/subtractions/multiplications). What is optimal? (minimizing PTree. Set functional creations and PTree. Set operations. )
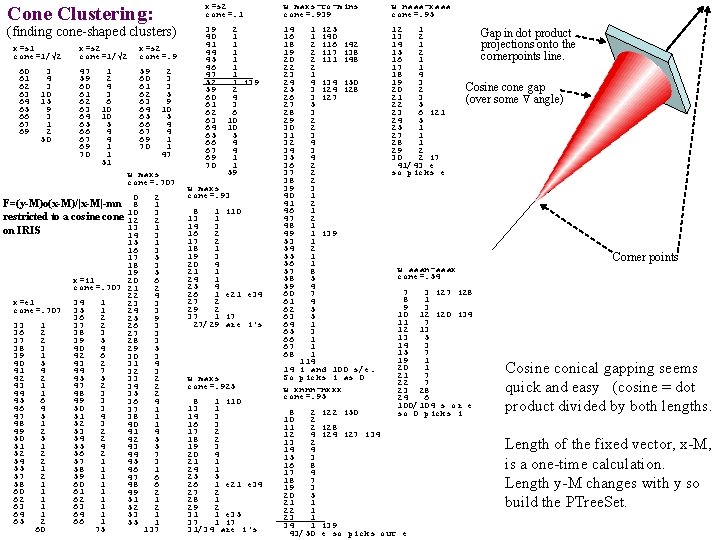
Cone Clustering: (finding cone-shaped clusters) x=s 1 cone=1/√ 2 x=s 2 cone=. 9 60 3 61 4 62 3 63 10 64 15 65 9 66 3 67 1 69 2 50 47 1 59 2 60 4 61 3 62 6 63 10 64 10 65 5 66 4 67 4 69 1 70 1 51 59 2 60 3 61 3 62 5 63 9 64 10 65 5 66 4 67 4 69 1 70 1 47 w maxs cone=. 707 0 2 F=(y-M)o(x-M)/|x-M|-mn 8 1 restricted to a cosine cone 10 3 12 2 13 1 on IRIS 14 3 15 1 16 3 17 5 18 3 19 5 x=i 1 20 6 cone=. 707 21 2 22 4 x=e 1 34 1 23 3 cone=. 707 35 1 24 3 36 2 25 9 33 1 37 2 26 3 36 2 38 3 27 3 37 2 39 5 28 3 38 3 40 4 29 5 39 1 42 6 30 3 40 5 43 2 31 4 44 7 32 3 42 2 45 5 33 2 43 1 47 2 34 2 44 1 48 3 35 2 45 6 49 3 36 4 46 4 50 3 37 1 47 5 51 4 38 1 48 1 52 3 40 1 49 2 53 2 41 4 50 5 54 2 42 5 51 1 55 4 43 5 52 2 56 2 44 7 54 2 57 1 45 3 55 1 58 1 46 1 57 2 59 1 47 6 58 1 60 1 48 6 60 1 61 1 49 2 62 1 51 1 63 1 52 2 64 1 53 1 65 2 66 1 55 1 60 75 137 x=s 2 cone=. 1 w maxs-to-mins cone=. 939 w naaa-xaaa cone=. 95 39 2 40 1 41 1 44 1 45 1 46 1 47 1 52 1 i 39 59 2 60 4 61 3 62 6 63 10 64 10 65 5 66 4 67 4 69 1 70 1 59 14 1 i 25 16 1 i 40 18 2 i 16 i 42 19 2 i 17 i 38 20 2 i 11 i 48 22 2 23 1 24 4 i 34 i 50 25 3 i 24 i 28 26 3 i 27 27 5 28 3 29 2 30 2 31 3 32 4 34 3 35 4 36 2 37 2 38 2 39 3 40 1 41 2 46 1 47 2 48 1 49 1 i 39 53 1 54 2 55 1 56 1 57 8 58 5 59 4 60 7 61 4 62 5 63 5 64 1 65 3 66 1 67 1 68 1 114 14 i and 100 s/e. So picks i as 0 w xnnn-nxxx cone=. 95 12 1 13 2 14 1 15 2 16 1 17 1 18 4 19 3 20 2 21 3 22 5 23 6 i 21 24 5 25 1 27 1 28 1 29 2 30 2 i 7 41/43 e so picks e w maxs cone=. 93 8 1 i 10 13 1 14 3 16 2 17 2 18 1 19 3 20 4 21 1 24 1 25 4 26 1 e 21 e 34 27 2 29 2 37 1 i 7 27/29 are i's w maxs cone=. 925 8 1 i 10 13 1 14 3 16 3 17 2 18 2 19 3 20 4 21 1 24 1 25 5 26 1 e 21 e 34 27 2 28 1 29 2 31 1 e 35 37 1 i 7 31/34 are i's Gap in dot product projections onto the cornerpoints line. Cosine cone gap (over some angle) Corner points w aaan-aaax cone=. 54 7 3 i 27 i 28 8 1 9 3 10 12 i 20 i 34 11 7 12 13 13 5 14 3 15 7 19 1 20 1 21 7 22 7 23 28 24 6 100/104 s or e so 0 picks i 8 2 i 22 i 50 10 2 11 2 i 28 12 4 i 27 i 34 13 2 14 4 15 3 16 8 17 4 18 7 19 3 20 5 21 1 22 1 23 1 34 1 i 39 43/50 e so picks out e Cosine conical gapping seems quick and easy (cosine = dot product divided by both lengths. Length of the fixed vector, x-M, is a one-time calculation. Length y-M changes with y so build the PTree. Set.
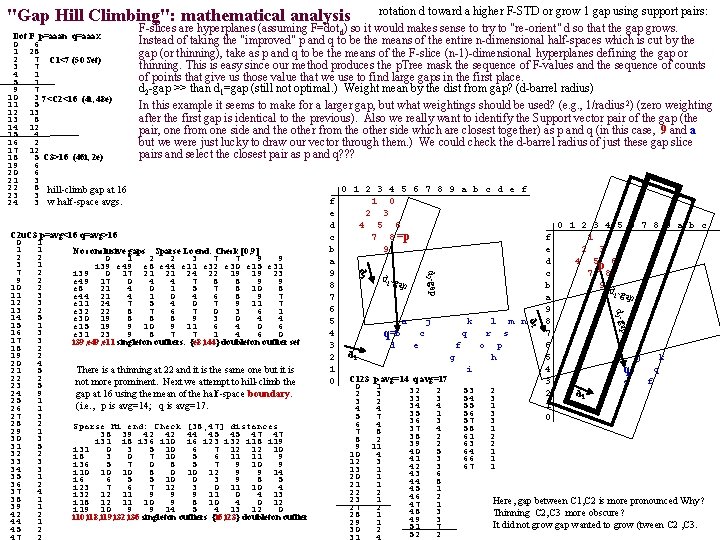
rotation d toward a higher F-STD or grow 1 gap using support pairs: F-slices are hyperplanes (assuming F=dotd) so it would makes sense to try to "re-orient" d so that the gap grows. Instead of taking the "improved" p and q to be the means of the entire n-dimensional half-spaces which is cut by the gap (or thinning), take as p and q to be the means of the F-slice (n-1)-dimensional hyperplanes defining the gap or thinning. This is easy since our method produces the p. Tree mask the sequence of F-values and the sequence of counts of points that give us those value that we use to find large gaps in the first place. d 2 -gap >> than d 1=gap (still not optimal. ) Weight mean by the dist from gap? (d-barrel radius) In this example it seems to make for a larger gap, but what weightings should be used? (e. g. , 1/radius 2) (zero weighting after the first gap is identical to the previous). Also we really want to identify the Support vector pair of the gap (the pair, one from one side and the other from the other side which are closest together) as p and q (in this case, 9 and a but we were just lucky to draw our vector through them. ) We could check the d-barrel radius of just these gap slice pairs and select the closest pair as p and q? ? ? "Gap Hill Climbing": mathematical analysis Dot F p=aaan q=aaax 0 6 1 28 2 7 C 1<7 (50 Set) 3 7 4 1 5 1 9 7 10 3 7<C 2<16 (4 i, 48 e) 11 5 12 13 13 8 14 12 15 4 16 2 17 12 18 5 C 3>16 (46 i, 2 e) 19 6 20 6 21 3 22 8 hill-climb gap at 16 23 3 24 3 w half-space avgs. 0 1 2 3 4 5 6 7 8 9 a b c d e f 1 0 2 3 4 5 6 7 8 =p 9 d 2 -gap d 2 d 1 -g ap j d e C 123 p avg=14 q avg=17 0 1 32 2 2 3 33 3 3 2 34 4 35 1 5 7 36 3 6 4 37 4 7 8 38 2 39 2 9 11 40 5 10 4 41 3 12 3 42 3 13 1 43 6 20 1 44 8 21 1 45 1 22 2 46 2 23 1 47 1 27 2 48 3 28 1 49 3 29 1 51 7 30 2 52 2 31 4 l q m n r f s o g p h i 53 54 55 56 57 58 61 63 64 66 67 2 3 1 3 3 1 2 2 1 1 1 0 1 2 3 4 5 6 7 8 9 a b c 1 2 3 4 5 p 6 7 8 9 d 1 -gap p d 1 k c f e d c b a 9 8 7 6 5 4 3 2 1 0 d 2 -ga a q=b d 2 C 2 u. C 3 p=avg<16 q=avg>16 0 1 1 1 No conclusive gaps Sparse Lo end: Check [0, 9] 2 2 0 1 2 2 3 7 7 9 9 3 1 i 39 e 49 e 8 e 44 e 11 e 32 e 30 e 15 e 31 7 2 i 39 0 17 21 21 24 22 19 19 23 9 2 e 49 17 0 4 4 7 8 8 9 9 10 2 e 8 21 4 0 1 5 7 8 10 8 11 3 e 44 21 4 1 0 4 6 8 9 7 12 3 e 11 24 7 5 4 0 7 9 11 7 13 2 e 32 22 8 7 6 7 0 3 6 1 14 5 e 30 19 8 8 8 9 3 0 4 4 15 1 e 15 19 9 10 9 11 6 4 0 6 16 3 e 31 23 9 8 7 7 1 4 6 0 17 3 i 39, e 49, e 11 singleton outliers. {e 8, i 44} doubleton outlier set 18 2 19 2 20 4 21 5 There is a thinning at 22 and it is the same one but it is 22 2 not more prominent. Next we attempt to hill-climb the 23 5 24 9 gap at 16 using the mean of the half-space boundary. 25 1 (i. e. , p is avg=14; q is avg=17. 26 1 27 3 28 2 Sparse Hi end: Check [38, 47] distances 29 1 38 39 42 42 44 45 45 47 47 30 3 i 31 i 8 i 36 i 10 i 6 i 23 i 32 i 18 i 19 31 5 i 31 0 3 5 10 6 7 12 12 10 32 2 i 8 3 0 7 10 5 6 11 11 9 33 3 i 36 5 7 0 8 5 7 9 10 9 34 3 i 10 10 10 8 0 10 12 9 9 14 35 1 i 6 6 5 5 10 0 3 9 8 5 36 2 i 23 7 6 7 12 3 0 11 10 4 37 4 i 32 12 11 9 9 9 11 0 4 13 38 1 i 18 12 11 10 9 8 10 4 0 12 39 1 i 19 10 9 9 14 5 4 13 12 0 42 2 i 10, i 18, i 19, i 32, i 36 singleton outliers {i 6, i 23} doubleton outlier 44 1 45 2 47 2 f e d c b a 9 8 7 6 5 4 3 2 1 0 a b d j k qc e q f d 1 Here, gap between C 1, C 2 is more pronounced Why? Thinning C 2, C 3 more obscure? It did not grow gap wanted to grow (tween C 2 , C 3.

CAINE 2013 Call for Papers 26 th International Conference on Computer Applications in Industry and Engineering September 25{27, 2013, Omni Hotel, Los Angles, Califorria, USA Sponsored by the International Society for Computers and Their Applications (ISCA) CAINE{2013 will feature contributed papers as well as workshops and special sessions. Papers will be accepted into oral presentation sessions. The topics will include, but are not limited to, the following areas: Agent-Based Systems Image/Signal Processing Autonomous Systems Information Assurance Big Data Analytics Information Systems/Databases Bioinformatics, Biomedical Systems/Engineering Internet and Web-Based Systems Computer-Aided Design/Manufacturing Knowledge-based Systems Computer Architecture/VLSI Mobile Computing Computer Graphics and Animation Multimedia Applications Computer Modeling/Simulation Neural Networks Computer Security Pattern Recognition/Computer Vision Computers in Education Rough Set and Fuzzy Logic Computers in Healthcare Robotics Computer Networks Fuzzy Logic Control Systems Sensor Networks Data Communication Scientic Computing Data Mining Software Engineering/CASE Distributed Systems Visualization Embedded Systems Wireless Networks and Communication Important Dates: Workshop/special session proposal. . May 2. 5, . 2. 013 Full Paper Submis. . June 5, . 2013. Notice Accept. . July. 5 , 2013. Pre-registration & Camera-Ready Paper Due. . . August 5, 2013. Event Dates. . . Sept 25 -27, 2013 SEDE Conf is interested in gathering researchers and professionals in the domains of SE and DE to present and discuss high-quality research results and outcomes in their fields. SEDE 2013 aims at facilitating cross-fertilization of ideas in Software and Data Engineering, The conference topics include, but not limited to: . Requirements Engineering for Data Intensive Software Systems. Software Verification and Model of Checking. Model-Based Methodologies. Software Quality and Software Metrics. Architecture and Design of Data Intensive Software Systems. Software Testing. Service- and Aspect-Oriented Techniques. Adaptive Software Systems. Information System Development. Software and Data Visualization. Development Tools for Data Intensive. Software Systems. Software Processes. Software Project Mgnt. Applications and Case Studies. Engineering Distributed, Parallel, and Peer-to-Peer Databases. Cloud infrastructure, Mobile, Distributed, and Peer-to-Peer Data Management. Semi-Structured Data and XML Databases. Data Integration, Interoperability, and Metadata. Data Mining: Traditional, Large-Scale, and Parallel. Ubiquitous Data Management and Mobile Databases. Data Privacy and Security. Scientific and Biological Databases and Bioinformatics. Social networks, web, and personal information management. Data Grids, Data Warehousing, OLAP. Temporal, Spatial, Sensor, and Multimedia Databases. Taxonomy and Categorization. Pattern Recognition, Clustering, and Classification. Knowledge Management and Ontologies. Query Processing and Optimization. Database Applications and Experiences. Web Data Mgnt and Deep Web May 23, 2013 Paper Submission Deadline June 30, 2013 Notification of Acceptance July 20, 2013 Registration and Camera-Ready Manuscript Conference Website: http: //theory. utdallas. edu/SEDE 2013/ ACC-2013 provides an international forum for presentation and discussion of research on a variety of aspects of advanced computing and its applications, and communication and networking systems. Important Dates May 5, 2013 - Special Sessions Proposal June 5, 2013 - Full Paper Submission July 5, 2013 - Author Notification Aug. 5, 2013 - Advance Registration & Camera Ready Paper Due CBR International Workshop Case-Based Reasoning CBR-MD 2013 July 19, 2013, New York/USA Topics of interest include (but are not limited to): CBR for signals, images, video, audio and text Similarity assessment Case representation and case mining Retrieval and indexing Conversational CBR Metalearning for model improvement and parameter setting for processing with CBR Incremental model improvement by CBR Case base maintenance for systems Case authoring Life-time of a CBR system Measuring coverage of case bases Ontology learning with CBR Submission Deadline: March 20 th, 2013 Notification Date: April 30 th, 2013 Camera-Ready Deadline: May 12 th, 2013 Workshop on Data Mining in Life Sciences DMLS Discovery of high-level structures, incl e. g. association networks Text mining from biomedical literatur Medical images mining Biomedical signals mining Temporal and sequential data mining Mining heterogeneous data Mining data from molecular biology, genomics, proteomics, pylogenetic classification With regard to different methodologies and case studies: Data mining project development methodology for biomedicine Integration of data mining in the clinic Ontology-driver data mining in life sciences Methodology for mining complex data, e. g. a combination of laboratory test results, images, signals, genomic and proteomic samples Data mining for personal disease management Utility considerations in DMLS, including e. g. cost-sensitive learning Submission Deadline: March 20 th, 2013 Notification Date: April 30 th, 2013 Camera-Ready Deadline: May 12 th, 2013 Workshop date: July 19 th, 2013 Workshop on Data Mining in Marketing DMM'2013 In business environment data warehousing - the practice of creating huge, central stores of customer data that can be used throughout the enterprise - is becoming more and more common practice and, as a consequence, the importance of data mining is growing stronger. Applications in Marketing Methods for User Profiling Mining Insurance Data E-Markteing with Data Mining Logfile Analysis Churn Management Association Rules for Marketing Applications Online Targeting and Controlling Behavioral Targeting Juridical Conditions of E-Marketing, Online Targeting and so one Controll of Online-Marketing Activities New Trends in Online Marketing Aspects of E-Mailing Activities and Newsletter Mailing Submission Deadline: March 20 th, 2013 Notification Date: April 30 th, 2013 Camera-Ready Deadline: May 12 th, 2013 Workshop date: July 19 th, 2013 Workshop Data Mining in Ag DMA 2013 Data Mining on Sensor and Spatial Data from Agricultural Applications Analysis of Remote Sensor Data Feature Selection on Agricultural Data Evaluation of Data Mining Experiments Spatial Autocorrelation in Agricultural Data Submission Deadline: March 20 th, 2013 Notification Date: April 30 th, 2013 Camera-Ready Deadline: May 12 th, 2013 Workshop date: July 19 th, 2013
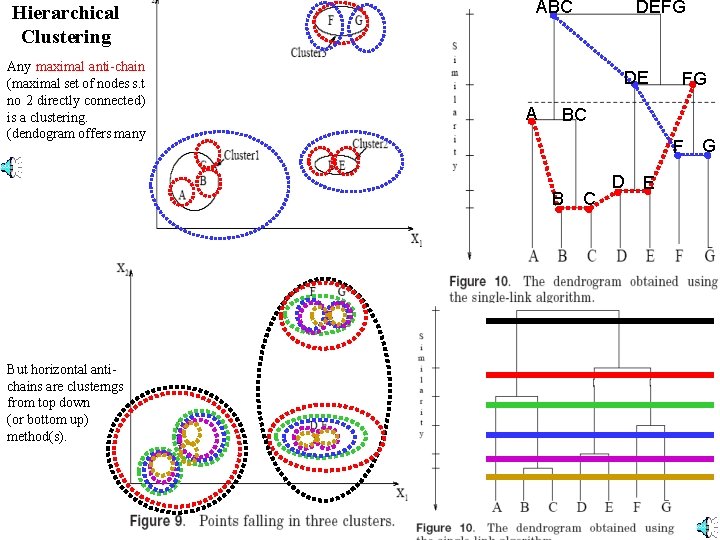
Hierarchical Clustering Any maximal anti-chain (maximal set of nodes s. t no 2 directly connected) is a clustering. (dendogram offers many DE A FG BC F B But horizontal antichains are clusterngs from top down (or bottom up) method(s). DEFG ABC D E C G
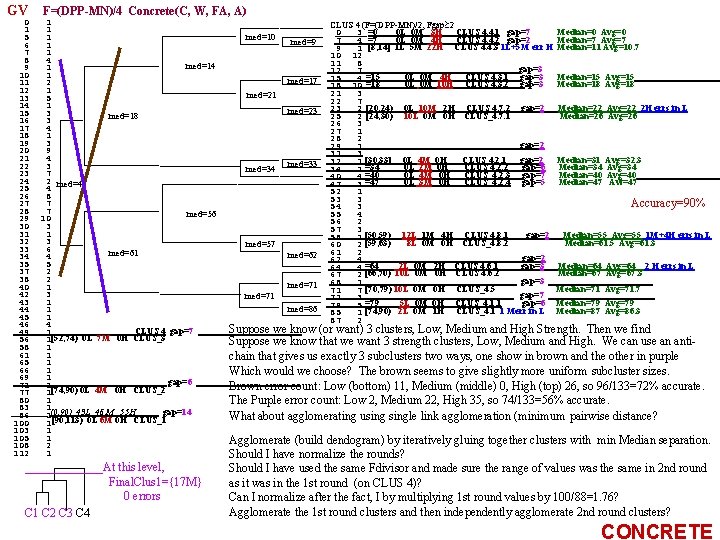
GV F=(DPP-MN)/4 Concrete(C, W, FA, A) 0 1 1 1 5 1 6 1 7 1 8 4 med=14 9 1 10 1 11 2 12 1 13 5 14 1 15 3 med=18 16 3 17 4 18 1 19 3 20 9 21 4 22 3 23 7 24 2 med=40 25 4 26 8 27 7 28 7 med=56 29 10 30 3 31 1 32 3 33 6 med=61 34 4 35 5 37 2 38 2 40 1 42 3 43 1 44 1 45 1 46 4 ______ CLUS 4 gap=7 49 1 56 1 [52, 74) 0 L 7 M 0 H CLUS_3 58 1 61 1 65 1 66 1 69 1 ______ gap=6 71 1 77 1 [74, 90) 0 L 4 M 0 H CLUS_2 80 1 83 1 ____ gap=14 86 1[0. 90) 43 L 46 M 55 H 100 1 [90, 113) 0 L 6 M 0 H CLUS_1 103 1 105 1 108 2 112 1 _______At this level, Final. Clus 1={17 M} 0 errors C 1 C 2 C 3 C 4 med=10 med=9 med=17 med=21 med=23 med=34 med=33 med=57 med=62 med=71 med=86 CLUS 4 (F=(DPP-MN)/2, Fgap 2 _______ 0 L 0 M 3 H CLUS 4. 4. 1 gap=7 0 3 =0 0 L 0 M 4 H CLUS 4. 4. 2 gap=2 7 4 =7 9 1 [8, 14] 1 L 5 M 22 H CLUS 4. 4. 3 1 L+5 M err H 10 12 11 8 gap=3 12 7 ______ 0 L 0 M 4 H CLUS 4. 3. 1 gap=3 15 4 =15 =18 0 L 0 M 10 H CLUS 4. 3. 2 gap=3 18 10 21 3 22 7 ______ 23 2 [20, 24) 0 L 10 M 2 H CLUS 4. 7. 2 gap=2 25 2 [24, 30) 10 L 0 M 0 H CLUS_4. 7. 1 26 3 27 1 28 2 gap=2 29 1 31 3 CLUS 4. 2. 1 gap=2 32 1 [30, 33] 0 L 4 M 0 H 0 L 2 M 0 H CLUS 4. 2. 2 gap=6 34 2 =34 ______ 0 L 4 M 0 H CLUS_4. 2. 3 gap=7 40 4 =40 0 L 3 M 0 H CLUS_4. 2. 4 gap=5 47 3 =47 52 1 53 3 54 3 55 4 56 2 57 3 ______ gap=2 58 1 [50, 59) 12 L 1 M 4 H CLUS 4. 8. 1 8 L 0 M 0 H CLUS_4. 8. 2 60 2 [59, 63) 61 2 gap=2 62 4 ______ 2 L 0 M 2 H CLUS 4. 6. 1 gap=3 64 4 =64 67 2 [66, 70) 10 L 0 M 0 H CLUS 4. 6. 2 gap=3 68 1 71 7 [70, 79) 10 L 0 M 0 H CLUS_4. 5 ______ gap=7 72 3 5 L 0 M 0 H CLUS_4. 1. 1 gap=6 79 5 =79 85 1 [74, 90) 2 L 0 M 1 H CLUS_4. 1 1 Merr in L 87 2 Median=0 Avg=0 Median=7 Avg=7 Median=11 Avg=10. 7 Median=15 Avg=15 Median=18 Avg=18 Median=22 Avg=22 2 H errs in L Median=26 Avg=26 Median=31 Median=34 Median=40 Median=47 Avg=32. 3 Avg=34 Avg=40 Avt=47 Accuracy=90% Median=55 Avg=55 1 M+4 H errs in L Median=61. 5 Avg=61. 3 Median=64 Avg=64 2 H errs in L Median=67 Avg=67. 3 Median=71 Avg=71. 7 Median=79 Avg=79 Median=87 Avg=86. 3 Suppose we know (or want) 3 clusters, Low, Medium and High Strength. Then we find Suppose we know that we want 3 strength clusters, Low, Medium and High. We can use an antichain that gives us exactly 3 subclusters two ways, one show in brown and the other in purple Which would we choose? The brown seems to give slightly more uniform subcluster sizes. Brown error count: Low (bottom) 11, Medium (middle) 0, High (top) 26, so 96/133=72% accurate. The Purple error count: Low 2, Medium 22, High 35, so 74/133=56% accurate. What about agglomerating usingle link agglomeration (minimum pairwise distance? Agglomerate (build dendogram) by iteratively gluing together clusters with min Median separation. Should I have normalize the rounds? Should I have used the same Fdivisor and made sure the range of values was the same in 2 nd round as it was in the 1 st round (on CLUS 4)? Can I normalize after the fact, I by multiplying 1 st round values by 100/88=1. 76? Agglomerate the 1 st round clusters and then independently agglomerate 2 nd round clusters? CONCRETE
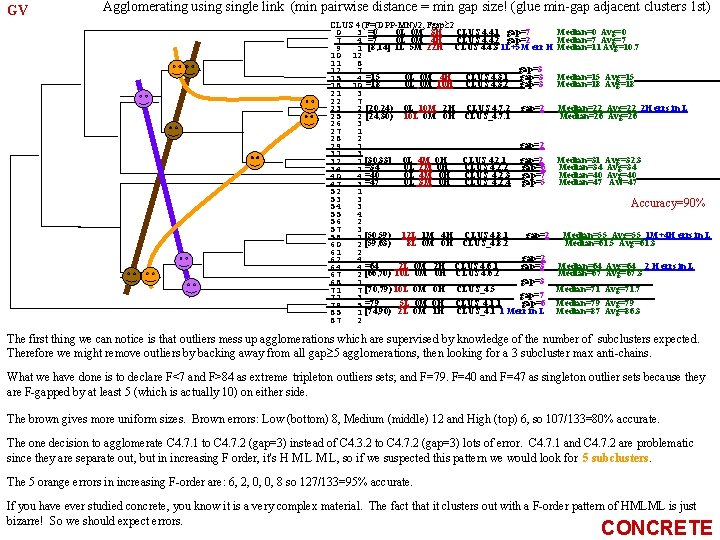
GV Agglomerating usingle link (min pairwise distance = min gap size! (glue min-gap adjacent clusters 1 st) CLUS 4 (F=(DPP-MN)/2, Fgap 2 _______ 0 L 0 M 3 H CLUS 4. 4. 1 gap=7 0 3 =0 0 L 0 M 4 H CLUS 4. 4. 2 gap=2 7 4 =7 9 1 [8, 14] 1 L 5 M 22 H CLUS 4. 4. 3 1 L+5 M err H 10 12 11 8 gap=3 12 7 ______ 0 L 0 M 4 H CLUS 4. 3. 1 gap=3 15 4 =15 =18 0 L 0 M 10 H CLUS 4. 3. 2 gap=3 18 10 21 3 22 7 ______ 23 2 [20, 24) 0 L 10 M 2 H CLUS 4. 7. 2 gap=2 25 2 [24, 30) 10 L 0 M 0 H CLUS_4. 7. 1 26 3 27 1 28 2 gap=2 29 1 31 3 CLUS 4. 2. 1 gap=2 32 1 [30, 33] 0 L 4 M 0 H 0 L 2 M 0 H CLUS 4. 2. 2 gap=6 34 2 =34 ______ 0 L 4 M 0 H CLUS_4. 2. 3 gap=7 40 4 =40 0 L 3 M 0 H CLUS_4. 2. 4 gap=5 47 3 =47 52 1 53 3 54 3 55 4 56 2 57 3 ______ gap=2 58 1 [50, 59) 12 L 1 M 4 H CLUS 4. 8. 1 8 L 0 M 0 H CLUS_4. 8. 2 60 2 [59, 63) 61 2 gap=2 62 4 ______ 2 L 0 M 2 H CLUS 4. 6. 1 gap=3 64 4 =64 67 2 [66, 70) 10 L 0 M 0 H CLUS 4. 6. 2 gap=3 68 1 71 7 [70, 79) 10 L 0 M 0 H CLUS_4. 5 ______ gap=7 72 3 5 L 0 M 0 H CLUS_4. 1. 1 gap=6 79 5 =79 85 1 [74, 90) 2 L 0 M 1 H CLUS_4. 1 1 Merr in L 87 2 Median=0 Avg=0 Median=7 Avg=7 Median=11 Avg=10. 7 Median=15 Avg=15 Median=18 Avg=18 Median=22 Avg=22 2 H errs in L Median=26 Avg=26 Median=31 Median=34 Median=40 Median=47 Avg=32. 3 Avg=34 Avg=40 Avt=47 Accuracy=90% Median=55 Avg=55 1 M+4 H errs in L Median=61. 5 Avg=61. 3 Median=64 Avg=64 2 H errs in L Median=67 Avg=67. 3 Median=71 Avg=71. 7 Median=79 Avg=79 Median=87 Avg=86. 3 The first thing we can notice is that outliers mess up agglomerations which are supervised by knowledge of the number of subclusters expected. Therefore we might remove outliers by backing away from all gap 5 agglomerations, then looking for a 3 subcluster max anti-chains. What we have done is to declare F<7 and F>84 as extreme tripleton outliers sets; and F=79. F=40 and F=47 as singleton outlier sets because they are F-gapped by at least 5 (which is actually 10) on either side. The brown gives more uniform sizes. Brown errors: Low (bottom) 8, Medium (middle) 12 and High (top) 6, so 107/133=80% accurate. The one decision to agglomerate C 4. 7. 1 to C 4. 7. 2 (gap=3) instead of C 4. 3. 2 to C 4. 7. 2 (gap=3) lots of error. C 4. 7. 1 and C 4. 7. 2 are problematic since they are separate out, but in increasing F order, it's H M L, so if we suspected this pattern we would look for 5 subclusters. The 5 orange errors in increasing F-order are: 6, 2, 0, 0, 8 so 127/133=95% accurate. If you have ever studied concrete, you know it is a very complex material. The fact that it clusters out with a F-order pattern of HMLML is just bizarre! So we should expect errors. CONCRETE
- Slides: 35