Real TimePCR Results Whats Wrong With Agarose Gels









- Slides: 19

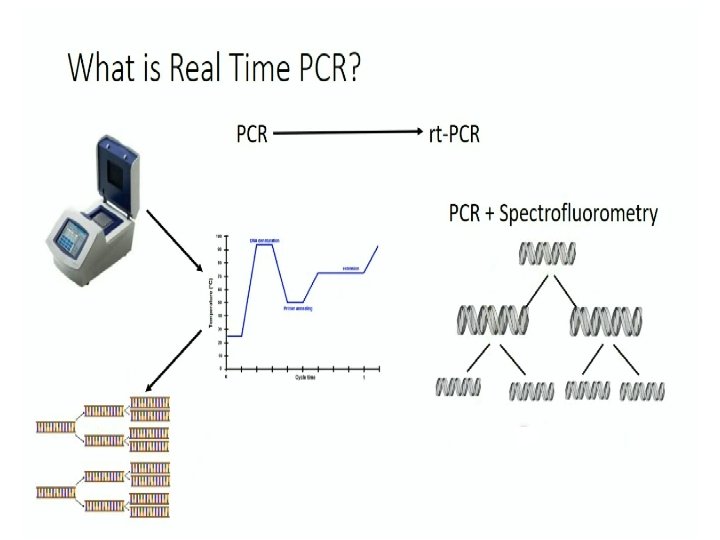
Real. Time-PCR


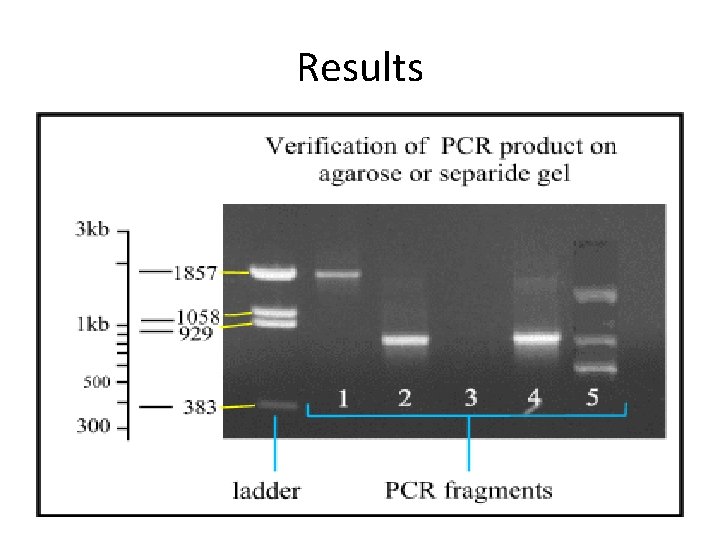
Results

What’s Wrong With Agarose Gels? § § § Low sensitivity Non-automated End point analysis

Working of QPCR? • Real Time PCR is a technique in which probes bind to specific target regions of amplicons (GOI) to produce fluorescence during PCR. • The fluorescence, measured in Real Time, is detected in a PCR cycler with an inbuilt filter flurometer. • In molecular biology, a hybridization probe is a fragment of DNA or RNA of variable length (usually 20– 30 bases long) which can be radioactively labeled. It can then be used in DNA or RNA samples to detect the presence of nucleotide sequences (the DNA target) that are complementary to the sequence in the probe.

Probe types & Design

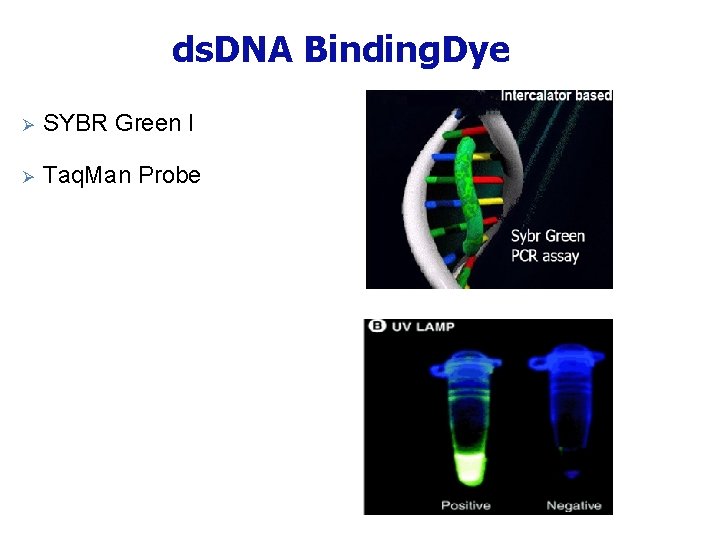
ds. DNA Binding. Dye Ø SYBR Green I Ø Taq. Man Probe

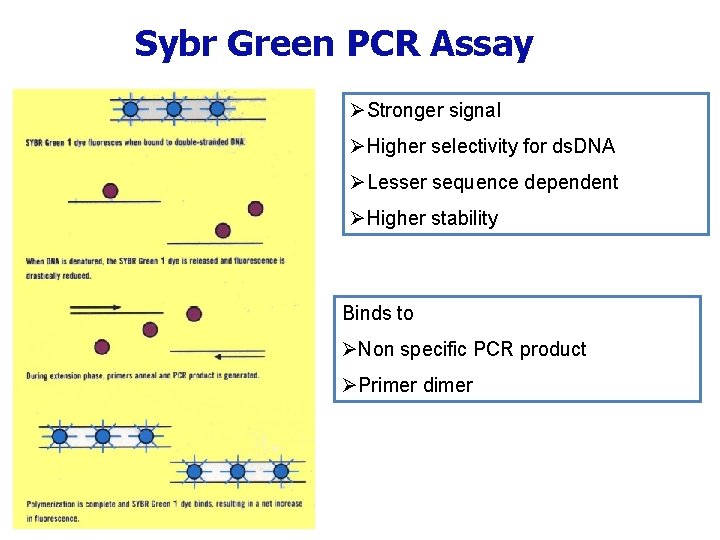
Sybr Green PCR Assay ØStronger signal ØHigher selectivity for ds. DNA ØLesser sequence dependent ØHigher stability Binds to ØNon specific PCR product ØPrimer dimer

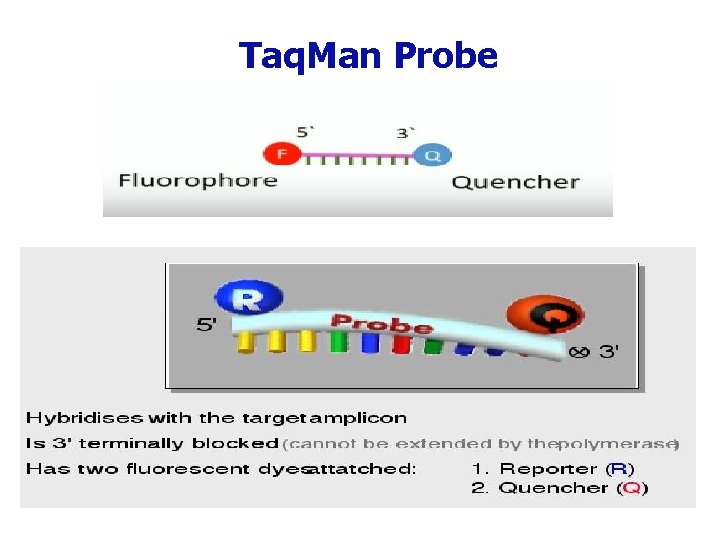
Taq. Man Probe

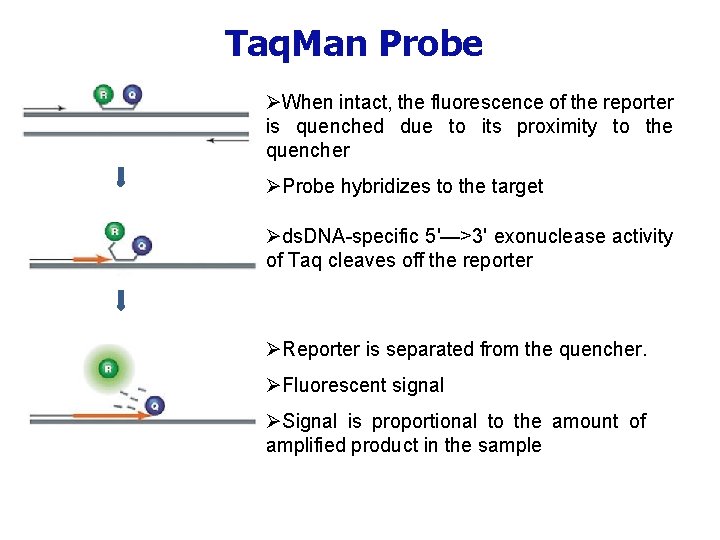
Taq. Man Probe ØWhen intact, the fluorescence of the reporter is quenched due to its proximity to the quencher ØProbe hybridizes to the target Øds. DNA-specific 5'—>3' exonuclease activity of Taq cleaves off the reporter ØReporter is separated from the quencher. ØFluorescent signal ØSignal is proportional to the amount of amplified product in the sample

Taq. Man Probe Advantages Ø Highly fluorogenic Ø Easy PCR setup Ø Sequence-specific detection Disadvantages Ø Expensive Ø Probe design and positioning challenging Ø Similar conditions for primers and probes

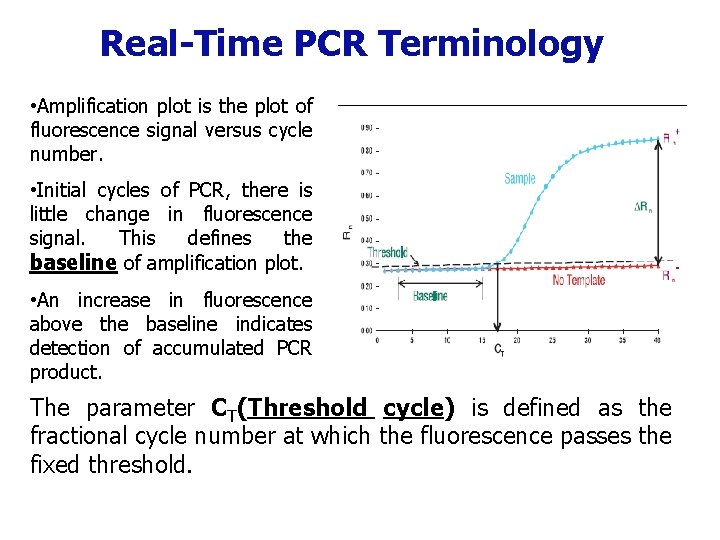
Real-Time PCR Terminology • Amplification plot is the plot of fluorescence signal versus cycle number. • Initial cycles of PCR, there is little change in fluorescence signal. This defines the baseline of amplification plot. • An increase in fluorescence above the baseline indicates detection of accumulated PCR product. The parameter CT(Threshold cycle) is defined as the fractional cycle number at which the fluorescence passes the fixed threshold.

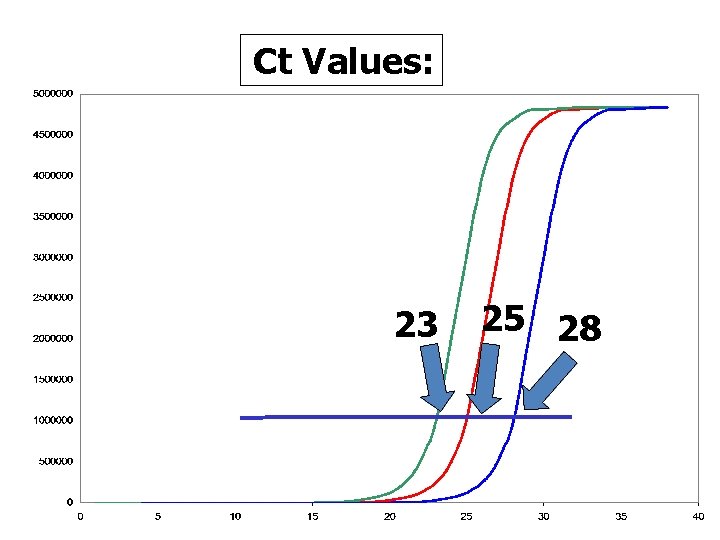
Ct Values: 23 25 28

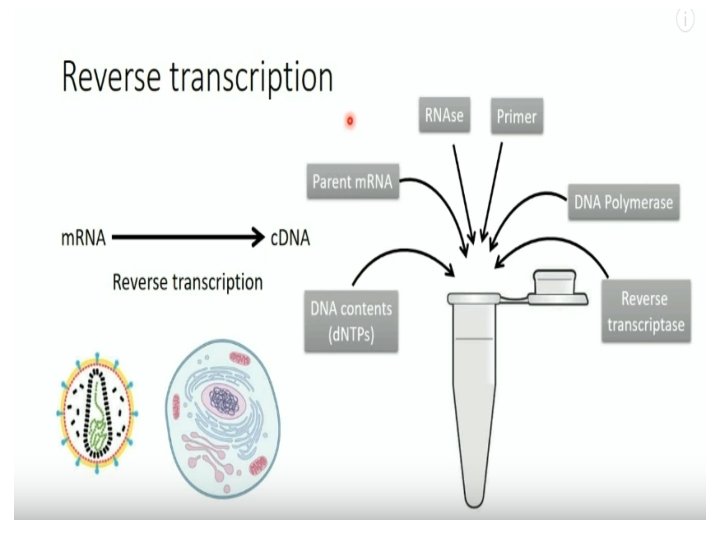
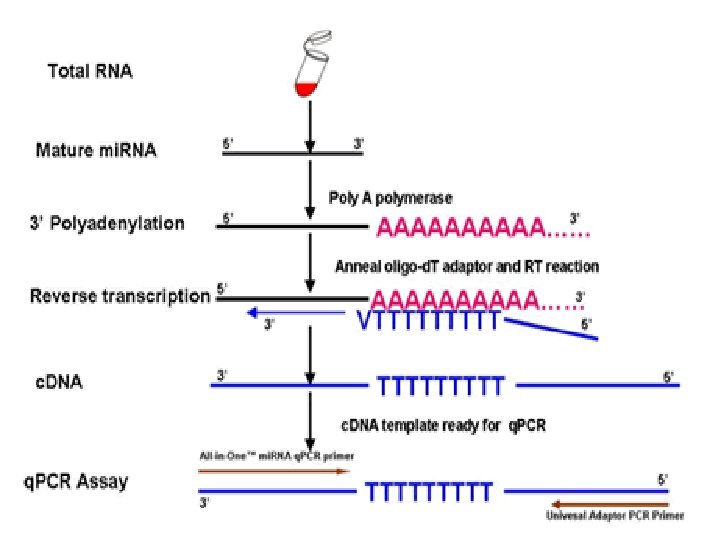
Reverse Transcription (RT) PCR • RT-PCR : a highly sensitive technique for the detection and quantitation of m. RNA (messenger RNA). The technique consists of two parts: The synthesis of c. DNA (complementary DNA) from RNA by reverse transcription (RT) and. The amplification of a specific c. DNA by the polymerase chain reaction (PCR).



Relative Quantification Housekeeping gene: Abundantly and constantly expressed gene. Expression level of these genes remains constant. eg 18 S r. RNA, GAPDH, β Actin Normalization: To accurately quantify gene expression, the measured amount of RNA from the gene of interest is divided by the amount of RNA from a housekeeping gene measured in the sample to normalize for possible variation in the amount and quality of RNA between different samples.

REALTIME-PCR APPLICATIONS

Application in Molecular Diagnostics ØClinical microbiology and Food microbiology ØGene expression Øviral quantitation ØSingle Nucleotide Polymorphism (SNP) analysis ØCancer ØAnalysis of cellular immune response in peripheral blood ØChromosome aberrations