Lecture 10 Large projects Trevor A Branch FISH

Lecture 10 Large projects Trevor A. Branch FISH 553 Advanced R School of Aquatic and Fishery Sciences University of Washington

Course evaluations Please come and find me when everyone is finished

Overall aims for R code • Correctness – Have you found all the bugs? • Replicable research – Can someone else repeat your results? – Do you get the same answer every time? • Understandable – Can someone else understand how your code works? – Will you understand your own code in 6 months? 5 years? • Speed of code – Does it run fast enough for your needs?

Speed “We should forget about small efficiencies, say about 97% of the time: premature optimization is the root of all evil” Donald Knuth (author of The Art of Computer Programming, the father of the analysis of computer algorithms, created Te. X in spare time) Photo: Jacob Appelbaum

Big projects The ninety-ninety rule (Tim Cargill): “The first 90% of the code accounts for the first 90% of the development time. The remaining 10% of the code accounts for the other 90% of the development time. ”

Commenting • The most important factor in understanding code is commenting • Each R script should have the following at the top: purpose of the code, original author, date started, any modifications (dated), authors who modified the code • Before every function: a description of what the function does • When doing something unusual, an explanation of what is being done

Different strategies for large projects • Organization of code – – Long scripts Breaking into functions Separate script for each figure or analysis Separate function files with sourcing • Ways to approach programming – Top-down programming – Bottom-up programming

Organization 1: long scripts • Create a single R file, fill it with sequential R commands, no functions, all objects available in the global workspace • Advantage: easy to write • Disadvantage: hard to understand therefore debug, global variables increase the chance of an unexpected outcome, hard to reuse parts of code

Organization 2: lots of functions One long R script with all code for a project Break up your code into individual functions Create functions that call other functions Divide the script into commented sections for each figure or analysis • Advantage: modular, reusable, easier to understand, no global variables • Disadvantage: can be hard to figure out which lines to run to get each result, especially when lots of figures are produced (or analyses run) • •

Organization 3: one R script per task • Each. r file does just one analysis or plot • I almost always use this method • Create a separate. r file for every figure: Fig 1. r, Fig 2. r, etc. Inside, a function plus code to read the data plus code to run the function • Thus running the code in "Fig 1. r" would create Figure 1 in my paper • Advantage: easily replicable, easy to understand, short • Disadvantage: reusing code is harder, what if I want to create four very similar types of figures?
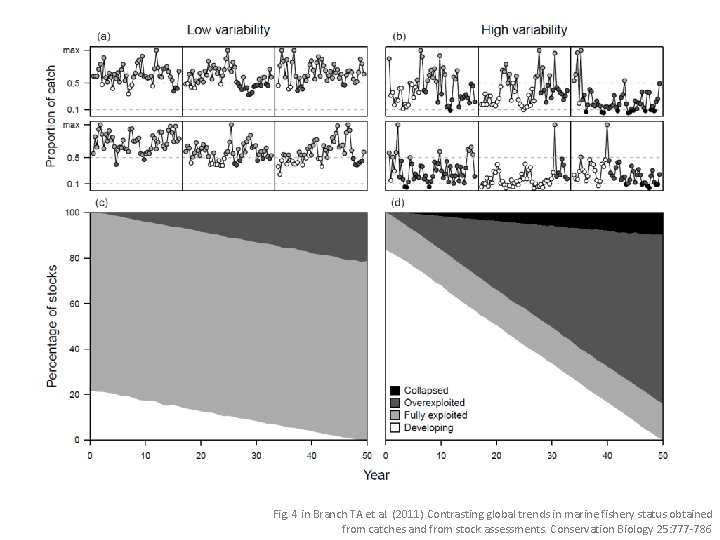
Fig. 4 in Branch TA et al. (2011) Contrasting global trends in marine fishery status obtained from catches and from stock assessments. Conservation Biology 25: 777 -786

To create this figure. . . • All plotting code in one script "Fig 1 sim catch v 1. r" • For major changes I save a copy to directory "old" and increment the filename to "Fig 1 sim catch v 2. r" • The script contains a number of functions: – stackpoly. TB 2() creates the plot – st. autocorrel. catch() simulates 100 -yr of catches – lognorm. catch() plots sample time series (upper panels) – random. status. pauly. 2007() does the lower left plot – simulated. plot 2() master function that calls the other functions to simulate data and plot the results • At the end of the script I call the master function pdf("Fig 1 v 1. pdf", width=8, height=6) simulated. plot 2(autocorrel=0. 5, nsims=2000) dev. off()

In-class exercise 1 • Download the file "10 • Run the code Fig 1. r" from the lectures – Either source the entire file (source button) – Ctrl+A to select all the code and then Ctrl+Enter to run it – Or in the console type source("10 Fig 1. r") • In your working directory, look for the output file "Fig 1 v 1. pdf" • Problem: the code is not replicable because it uses random numbers and I didn’t set a random seed • Modify the code using set. seed() to obtain a repeatable plot

Organization 4: sourcing functions • When each function may be called in multiple ways to produce many different figures • Or you will use the same functions every year to repeat your analysis on a different set of data • Store each major function in a separate. r file that does not include code for calling the function • Create one master file "2013 function calls. r" • The master file will source() the functions and create individual plots/analyses

Example: southern bluefin tuna • Every year I attend CCSBT meetings • My job as consultant to the international panel is to take (very complicated) model output and create (very complicated) plots • I use the same functions every year but the data change • I may use the same function for several plots • I need a record of how I produced each plot

All saved R commands during July 2013 meeting ###################### #Loads all _lab. rep files into one R list #Trevor A. Branch (with Ana Parma) July 2009 ###################### get. all. files <- function(directory) { library(PBSmodelling) files<-dir(directory, pattern="lab. rep") nn = length(files) result <- list() for (i in 1: nn) { print(paste(i, "of", nn)) result[[i]] <- read. List(paste( directory, "\", files[i], sep="")) names(result)[i] <- files[i] } return(result) } Each file contains one function

Inside "2013_08 Portland meeting. r" #=================================== #List of r code used in Portland ME meeting of CCSBT, 23 -26 July 2013. #Written by Trevor A. Branch tbranch@uw. edu #=================================== #=====Required packages======= require(PBSmodelling) #Read in all data from the. lab rep files R script contains function get. all. files() source("get. all. files. r") data. base <- get. all. files("arc\base 2010 sqrt ") Run the function #***Figure 1 likelihood plot comparison source("Table. Likelihood. Components v 2. r") nll. table. base <- likelihood. table 2(data. objects = data. base ) source("Plot. NLLComponents. r") R script contains function plot. NLL. by. steepness() pdf("figs\Fig 1 NLLs base. pdf") plot. NLL. by. steepness(nll. table. base , caption="base 2010 sqrt") dev. off() #***Figure 2 plot the MSY, Fmsy, Bmsy R script contains function MSY. vals() source("MSY values. r") pdf(file="figs\Fig 2 MSY. pdf ", width=10, height=8) x <- MSY. vals(data. objects=data. base , label="") dev. off() #***Figure 3 shaded plot data. base source("Shaded. Plots. r") R script contains function plot. lev() jpeg(file="figs\Fig 3 Shaded base 2010 sqrt. jpg", width=600 , height=600) plot. lev("levfiles\base 2010 sqrt. lev") dev. off()

After the meeting • I get a request: can you please redo Figure 34 and add a dashed line and a legend? • Go to "2013_08 Portland meeting. r", find the code under #***Figure 34 • Copy that section of code to the end of the meeting script, and change the comments and pdf file name to the first available figure number e. g. #***Figure 65 • Find the R script that was sourced, change it and rerun the code • Send the new "Fig 65. pdf" back

In-class exercise 2 • Take the file "10 Fig 1. r" and organize it so that it runs by sourcing functions • Every original function will go to its own. r file • Create a new file called "overall script. r" that uses source() to make each function available and then runs code to create the figure • Add code to the overall script to create four figures, calling the "10 Fig 1. r" code with autocorrel=0. 05, 0. 2, 0. 5, and 0. 8 • Use set. seed() to ensure each figure is comparable
- Slides: 19