DNA sequencing Determination of nucleotide sequence the determination









- Slides: 11

DNA sequencing ü Determination of nucleotide sequence Ø the determination of the precise sequence of nucleotides in a sample of DNA Two similar methods: 1. Maxam and Gilbert method 2. Sanger method ü They depend on the production of a mixture of oligonucleotides labeled either radioactively or fluorescein, with one common end and differing in length by a single nucleotide at the other end ü This mixture of oligonucleotides is separated by high resolution electrophoresis on polyacrilamide gels and the position of the bands determined ü

Maxam-Gilbert Walter Gilbert ◦ Harvard physicist ◦ Knew James Watson ◦ Became intrigued with the biological side ◦ Became a biophysicist Allan Maxam

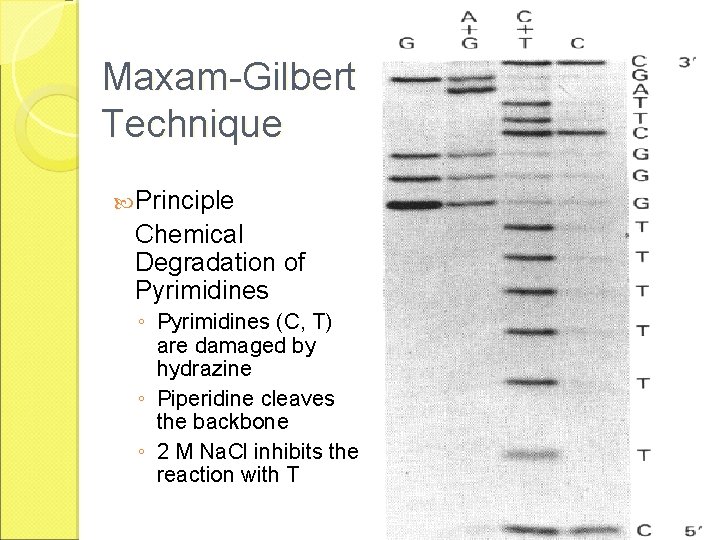
The Maxam-Gilbert Technique Principle - Chemical Degradation of Purines ◦ Purines (A, G) damaged by dimethylsulfate ◦ Methylation of base ◦ Heat releases base ◦ Alkali cleaves G ◦ Dilute acid cleave A>G

Maxam-Gilbert Technique Principle Chemical Degradation of Pyrimidines ◦ Pyrimidines (C, T) are damaged by hydrazine ◦ Piperidine cleaves the backbone ◦ 2 M Na. Cl inhibits the reaction with T

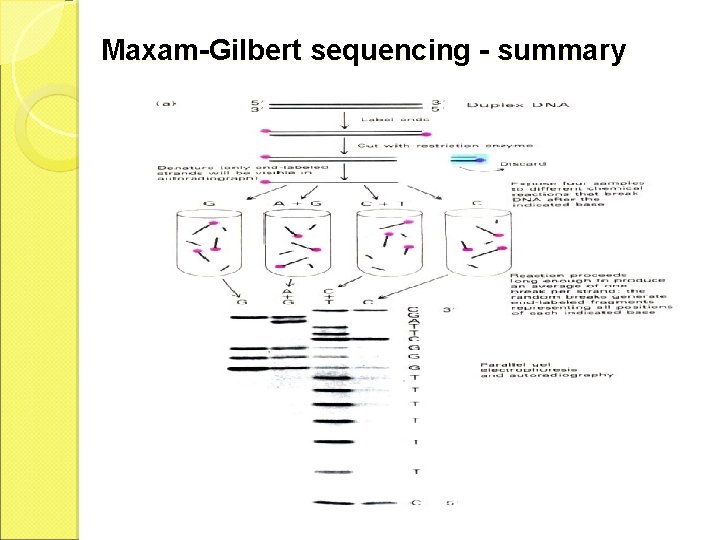
Maxam and Gilbert Method ü Chemical degradation of purified fragments (chemical degradation) ü The single stranded DNA fragment to be sequenced is endlabeled by treatment with alkaline phosphatase to remove the 5’phosphate ü It is then followed by reaction with P-labeled ATP in the presence of polynucleotide kinase, which attaches P labeled to the 5’terminal ü The labeled DNA fragment is then divided into four aliquots, each of which is treated with a reagent which modifies a specific base 1. Aliquot A + dimethyl sulphate, which methylates guanine residue 2. Aliquot B + formic acid, which modifies adenine and guanine residues 3. Aliquot C + Hydrazine, which modifies thymine + cytosine residues 4. Aliquot D + Hydrazine + 5 mol/l Na. Cl, which makes the reaction specific for cytosine ü The four are incubated with piperidine which cleaves the sugar phosphate backbone of DNA next to the residue that

Maxam-Gilbert sequencing - summary

Advantages/disadvantages Maxam-Gilbert sequencing Requires lots of purified DNA, and many intermediate purification steps ü Relatively short readings ü Automation not available (sequencers) ü Remaining use for ‘footprinting’ (partial protection against DNA modification when proteins bind to specific regions, and that produce ‘holes’ in the sequence ladder) ü In contrast, the Sanger sequencing methodology requires little if any DNA purification, no restriction digests, and no labeling of the DNA sequencing template

Sanger Method ü Fred Sanger, 1958 ◦ Was originally a protein chemist ◦ Made his first mark in sequencing proteins ◦ Made his second mark in sequencing RNA ü 1980 dideoxy sequencing

Original Sanger Method ü Random incorporation of a dideoxynucleoside triphosphate into a growing strand of DNA ü Requires DNA polymerase I ü Requires a cloning vector with initial primer (M 13, high yield bacteriophage, modified by adding: beta-galactosidase screening, polylinker) ü Uses 32 P-deoxynucleoside triphosphates

Sanger Method ü in-vitro DNA synthesis using ‘terminators’, use of dideoxi- nucleotides that do not permit chain elongation after their integration ü DNA synthesis using deoxy- and dideoxynucleotides that results in termination of synthesis at specific nucleotides ü Requires a primer, DNA polymerase, a template, a mixture of nucleotides, and detection system ü Incorporation of di-deoxynucleotides into growing strand terminates synthesis ü Synthesized strand sizes are determined for each di-deoxynucleotide by using gel or capillary electrophoresis ü Enzymatic methods

The principles Partial copies of DNA fragments made with DNA polymerase Collection of DNA fragments that terminate with A, C, G or T using dd. NTP Separate by gel electrophoresis Read DNA sequence