Assembly Scaffolding using String Graphs and In Silico

Assembly Scaffolding using String Graphs and In Silico Chromosome Assignment Zemin Ning The Wellcome Trust Sanger Institute
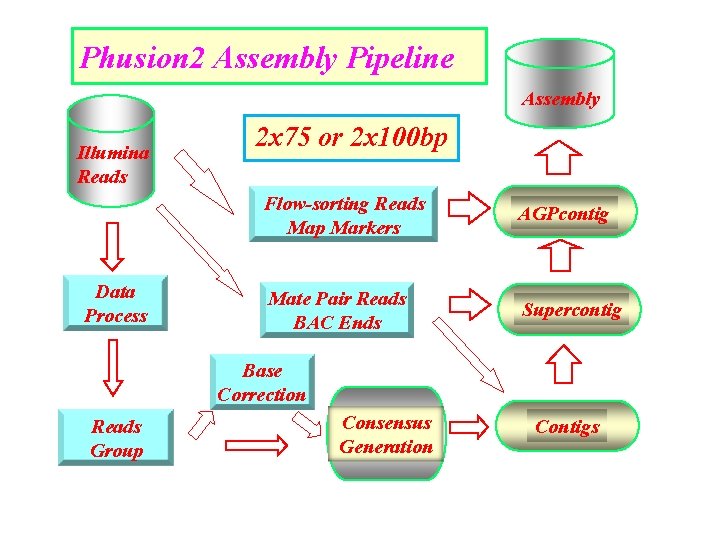
Phusion 2 Assembly Pipeline Assembly Illumina Reads Data Process 2 x 75 or 2 x 100 bp Flow-sorting Reads Map Markers AGPcontig Mate Pair Reads BAC Ends Supercontig Base Correction Reads Group Consensus Generation Contigs
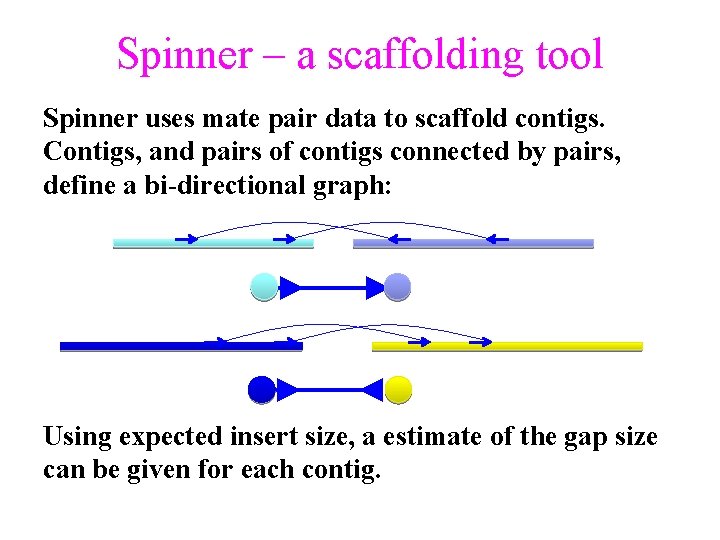
Spinner – a scaffolding tool Spinner uses mate pair data to scaffold contigs. Contigs, and pairs of contigs connected by pairs, define a bi-directional graph: Using expected insert size, a estimate of the gap size can be given for each contig.
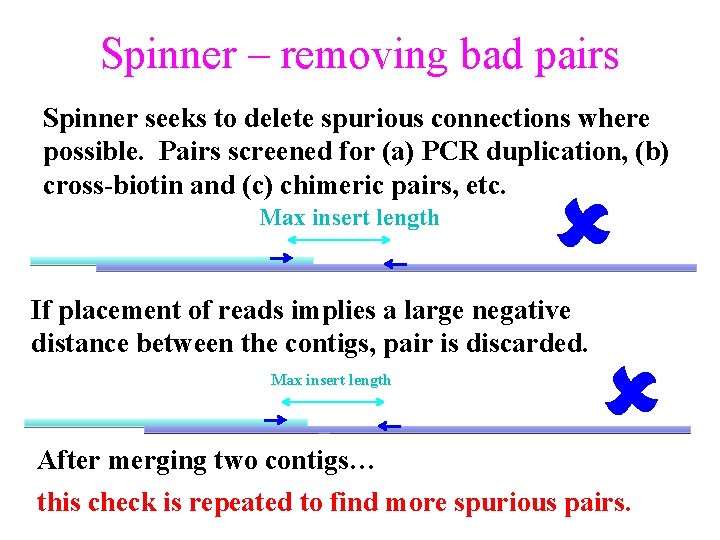
Spinner – removing bad pairs Spinner seeks to delete spurious connections where possible. Pairs screened for (a) PCR duplication, (b) cross-biotin and (c) chimeric pairs, etc. Max insert length If placement of reads implies a large negative distance between the contigs, pair is discarded. Max insert length After merging two contigs… this check is repeated to find more spurious pairs.
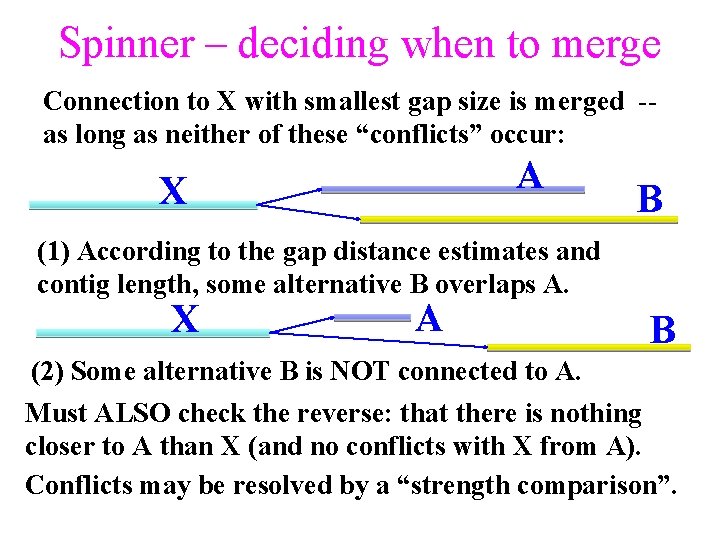
Spinner – deciding when to merge Connection to X with smallest gap size is merged -as long as neither of these “conflicts” occur: A X B (1) According to the gap distance estimates and contig length, some alternative B overlaps A. X A B (2) Some alternative B is NOT connected to A. Must ALSO check the reverse: that there is nothing closer to A than X (and no conflicts with X from A). Conflicts may be resolved by a “strength comparison”.
Spinner – still to do These techniques alone produces useful results. Further stages will be used to resolve repeats pairs that “jump over” repeats, and graph flow concepts.
Remove Heterozygosity Contigs
Pipeline of Contig Gap Closure
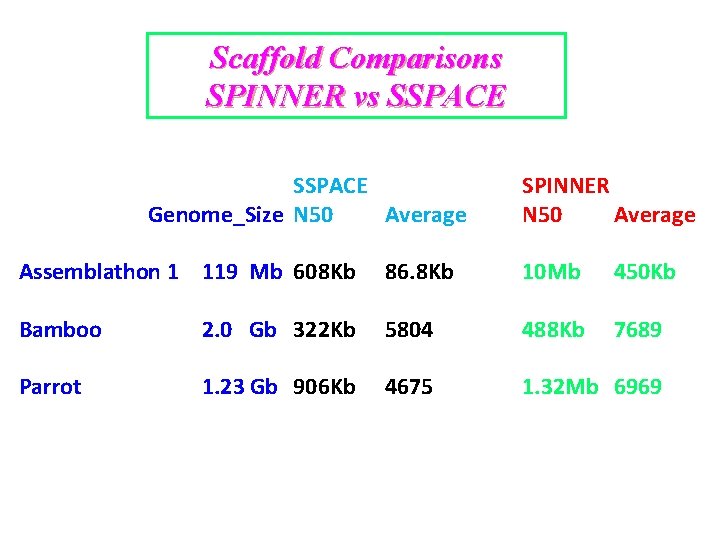
Scaffold Comparisons SPINNER vs SSPACE SSPACE Genome_Size N 50 Average SPINNER N 50 Average Assemblathon 1 119 Mb 608 Kb 86. 8 Kb Bamboo 2. 0 Gb 322 Kb 5804 Parrot 1. 23 Gb 906 Kb 4675 10 Mb 450 Kb 488 Kb 7689 1. 32 Mb 6969
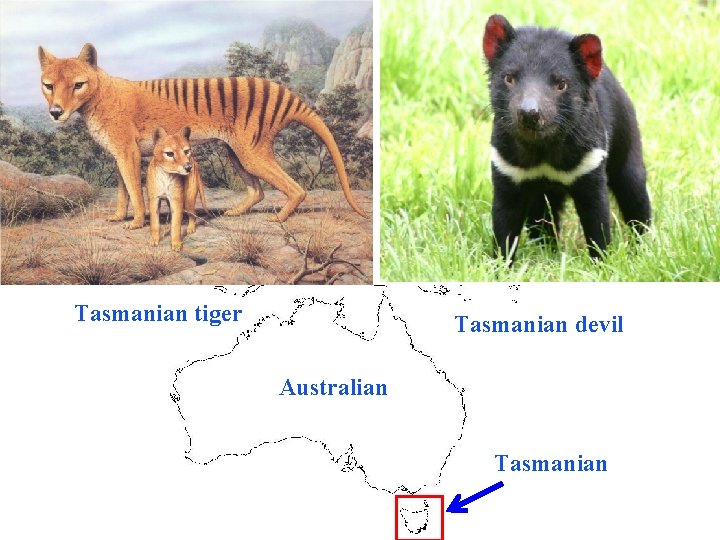
Tasmanian tiger Tasmanian devil Australian Tasmanian
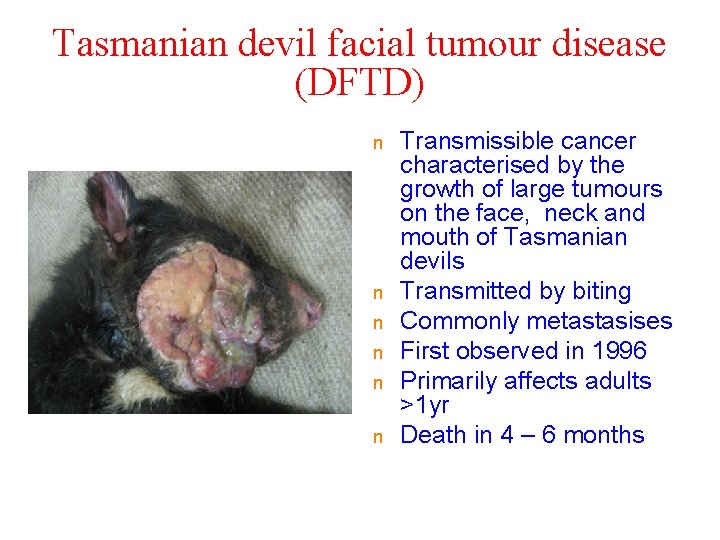
Tasmanian devil facial tumour disease (DFTD) n n n Transmissible cancer characterised by the growth of large tumours on the face, neck and mouth of Tasmanian devils Transmitted by biting Commonly metastasises First observed in 1996 Primarily affects adults >1 yr Death in 4 – 6 months
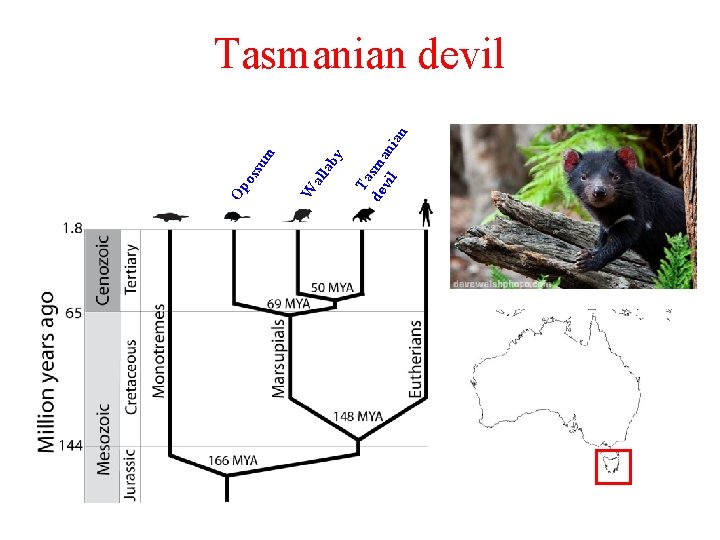
Ta de sma vil n ian ab y all W su m Op os Tasmanian devil
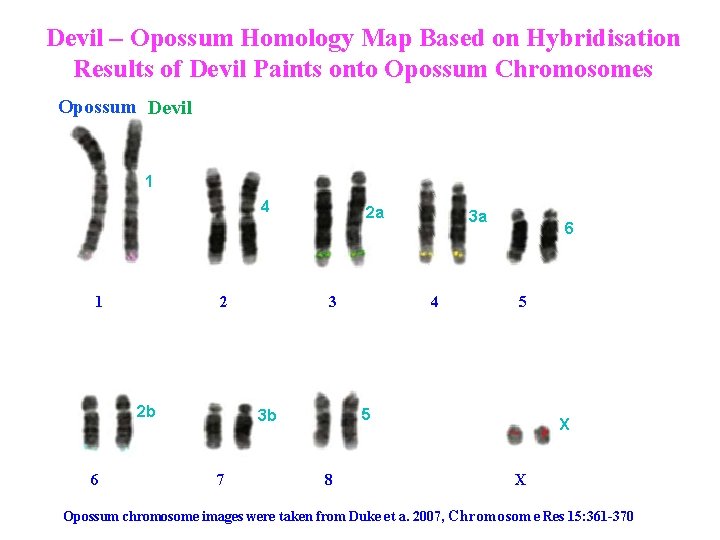
Devil – Opossum Homology Map Based on Hybridisation Results of Devil Paints onto Opossum Chromosomes Opossum Devil 1 4 1 2 2 b 6 2 a 3 4 6 5 5 3 b 7 3 a 8 X X Opossum chromosome images were taken from Duke et a. 2007, Chromosome Res 15: 361 -370
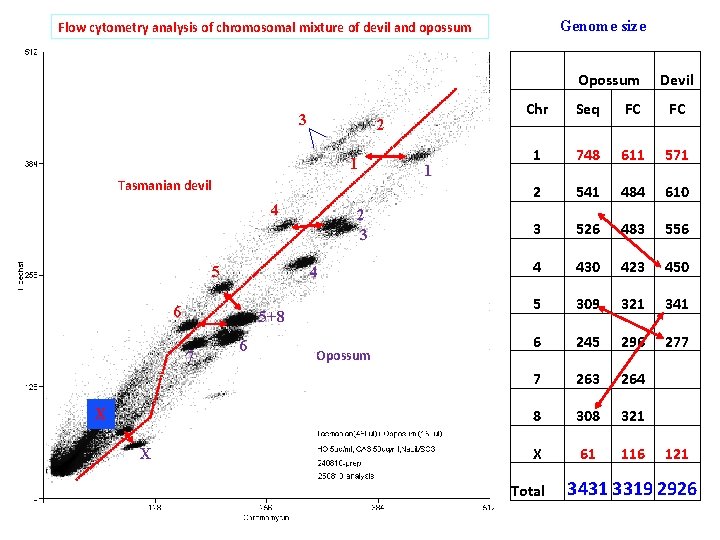
Genome size Flow cytometry analysis of chromosomal mixture of devil and opossum 3 2 1 Tasmanian devil 4 5 4 6 5+8 7 X X 2 3 6 Opossum 1 Opossum Devil Chr Seq FC FC 1 748 611 571 2 541 484 610 3 526 483 556 4 430 423 450 5 309 321 341 6 245 296 277 7 263 264 8 308 321 X 61 116 121 Total 3431 3319 2926

Table 1 Run ID, Template names, Number of reads and Chromosome size 4972_1 chr 1 IL 20_4972: 1 19. 8 571 4967_1 chr 2 IL 21_4967: 1 20. 0 610 4971_1 chr 3 IL 30_4971: 1 21. 7 556 4964_1 chr 4 IL 14_4964: 1 7. 26 450 4969_1 chr 5 IL 17_4969: 1 7. 06 341 4969_2 chr 6 IL 17_4969: 2 8. 59 277 4969_3 chrx IL 17_4969: 3 9. 43 122 Read mapping coefficient: e = Size_of_Chr/Num_reads_in_lane
Perfect - Reads from the same library were mapped to the contig
Acceptable - Majority of the reads were from the same library, but there were reads from other libraries
Bad – mis-assembly error Majority of the reads in one region were from one library. But there is a transition from which we see a new library, i. e. switch to another chromosome.
Unassigned contigs were placed by supercontigs using mate pairs
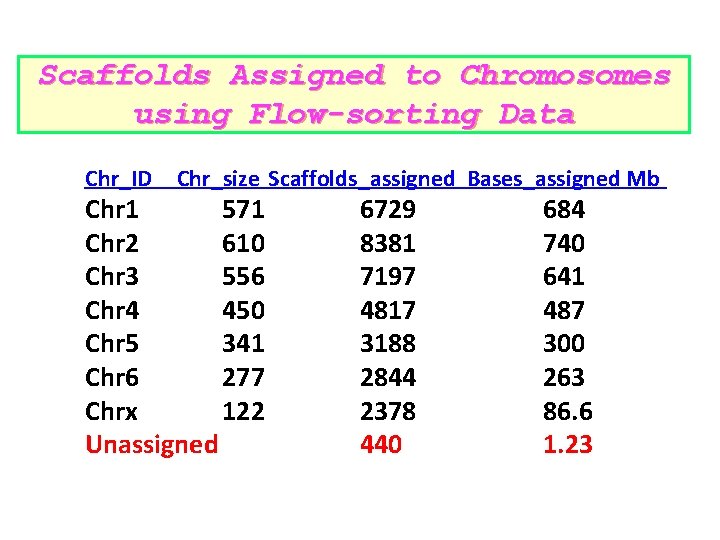
Scaffolds Assigned to Chromosomes using Flow-sorting Data Chr_ID Chr_size Scaffolds_assigned Bases_assigned Mb Chr 1 571 Chr 2 610 Chr 3 556 Chr 4 450 Chr 5 341 Chr 6 277 Chrx 122 Unassigned 6729 8381 7197 4817 3188 2844 2378 440 684 740 641 487 300 263 86. 6 1. 23
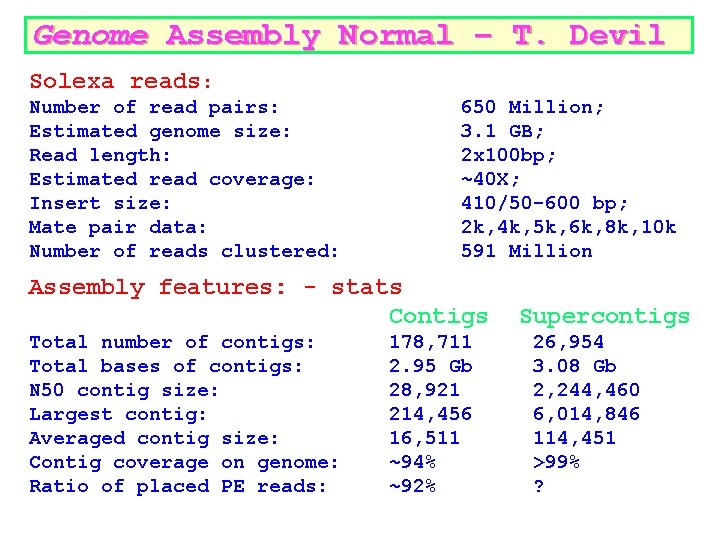
Genome Assembly Normal – T. Devil Solexa reads: Number of read pairs: Estimated genome size: Read length: Estimated read coverage: Insert size: Mate pair data: Number of reads clustered: 650 Million; 3. 1 GB; 2 x 100 bp; ~40 X; 410/50 -600 bp; 2 k, 4 k, 5 k, 6 k, 8 k, 10 k 591 Million Assembly features: - stats Contigs Total number of contigs: Total bases of contigs: N 50 contig size: Largest contig: Averaged contig size: Contig coverage on genome: Ratio of placed PE reads: 178, 711 2. 95 Gb 28, 921 214, 456 16, 511 ~94% ~92% Supercontigs 26, 954 3. 08 Gb 2, 244, 460 6, 014, 846 114, 451 >99% ?
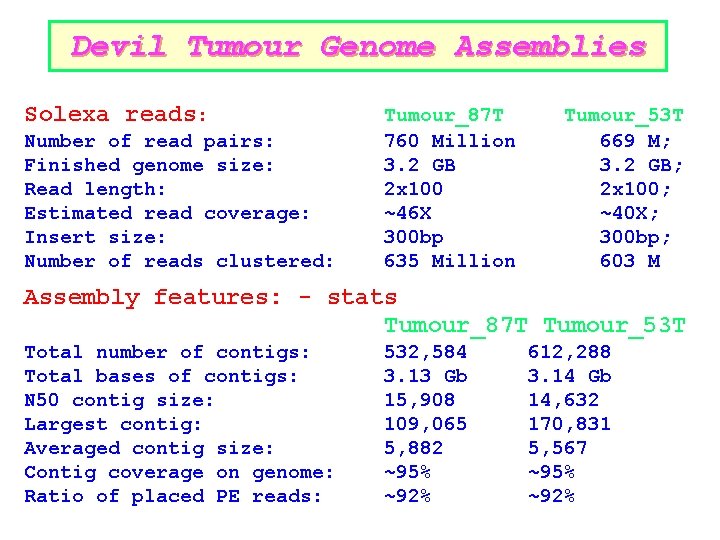
Devil Tumour Genome Assemblies Solexa reads: Number of read pairs: Finished genome size: Read length: Estimated read coverage: Insert size: Number of reads clustered: Tumour_87 T 760 Million 3. 2 GB 2 x 100 ~46 X 300 bp 635 Million Tumour_53 T 669 M; 3. 2 GB; 2 x 100; ~40 X; 300 bp; 603 M Assembly features: - stats Tumour_87 T Tumour_53 T Total number of contigs: Total bases of contigs: N 50 contig size: Largest contig: Averaged contig size: Contig coverage on genome: Ratio of placed PE reads: 532, 584 3. 13 Gb 15, 908 109, 065 5, 882 ~95% ~92% 612, 288 3. 14 Gb 14, 632 170, 831 5, 567 ~95% ~92%
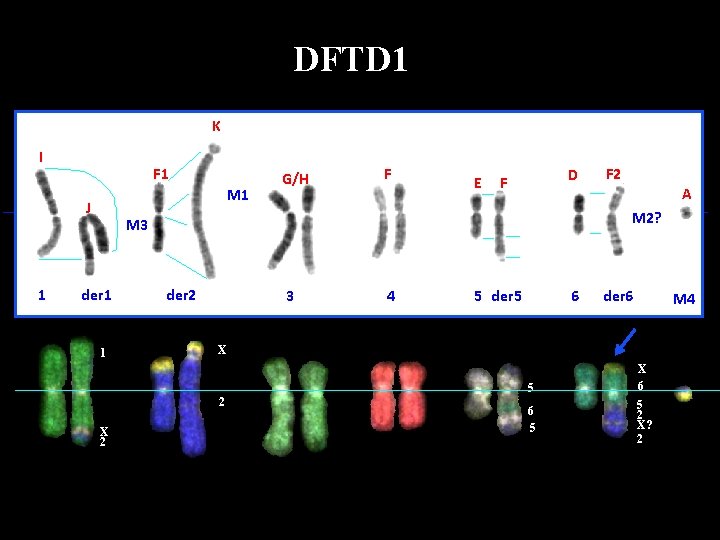
DFTD 1 K I F 1 M 1 J 1 G/H F E D F 1 der 2 3 4 5 der 5 6 der 6 M 4 X 5 2 X 2 A M 2? M 3 der 1 F 2 6 5 X 6 5 2 X? 2
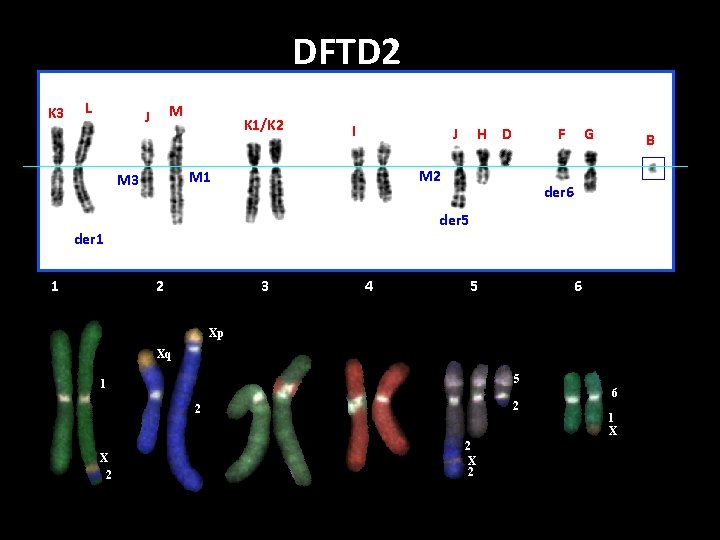
DFTD 2 K 3 L M J K 1/K 2 I G B der 6 der 5 der 1 1 F M 2 M 1 M 3 H D J 2 3 4 5 6 Xp Xq 5 1 2 2 X 2 6 1 X

Acknowledgements: q q q q q Joe Henson Elizabeth Murchuson David Mc. Bride Yong Gu Fengtang Yang Mike Stratton Ole Schulz-Trieglaff Dirk Evers David Bentley
- Slides: 25