Amino Acids When you understand the amino acids










- Slides: 13

Amino Acids “When you understand the amino acids, you understand everything ” ©CMBI 2006

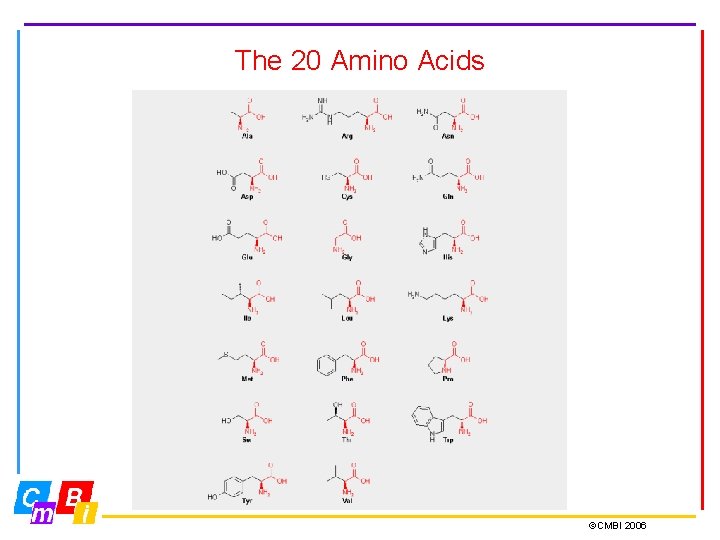
The 20 Amino Acids ©CMBI 2006

Secondary Structure Preference Amino acids form chains, the sequence or primary structure. These chains fold in -helices, b-strands, b-turns, and loops (or for short, helix, strand, turn and loop), the secondary structure. These secondary structure elements fold further to make whole proteins, but more about that later. There are relations between the physico-chemical characteristics of the amino acids and their secondary structure preference. I. e. , the b- branched residues (Ile, Thr, Val) like to sit in b-strands. ©CMBI 2006

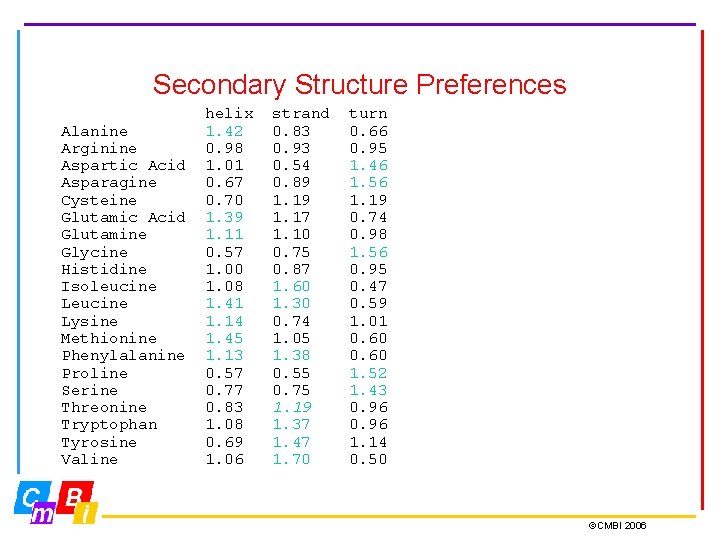
Secondary Structure Preferences Alanine Arginine Aspartic Acid Asparagine Cysteine Glutamic Acid Glutamine Glycine Histidine Isoleucine Lysine Methionine Phenylalanine Proline Serine Threonine Tryptophan Tyrosine Valine helix 1. 42 0. 98 1. 01 0. 67 0. 70 1. 39 1. 11 0. 57 1. 00 1. 08 1. 41 1. 14 1. 45 1. 13 0. 57 0. 77 0. 83 1. 08 0. 69 1. 06 strand 0. 83 0. 93 0. 54 0. 89 1. 17 1. 10 0. 75 0. 87 1. 60 1. 30 0. 74 1. 05 1. 38 0. 55 0. 75 1. 19 1. 37 1. 47 1. 70 turn 0. 66 0. 95 1. 46 1. 56 1. 19 0. 74 0. 98 1. 56 0. 95 0. 47 0. 59 1. 01 0. 60 1. 52 1. 43 0. 96 1. 14 0. 50 ©CMBI 2006

Chou Fasman parameters Say your dataset is 1000 amino acids and 350 of them are in alpha-helix conformation. This is 35%. There are 50 Alanines in your set and 25 of them are in alpha-helix conformation. This is 50%. The helix preference parameter P for Ala is 50/35=1, 43 ©CMBI 2006

Chou Fasman parameters Take home message: Preference parameter > 1. 0 specific residue has a preference for the specific secondary structure. Preference parameter = 1. 0 specific residue does not have a preference for, nor dislikes the specific secondary structure Preference parameter < 1. 0 specific residue dislikes the specific secondary structure. ©CMBI 2006

Secondary Structure Preferences Alanine Glutamic Acid Glutamine Leucine Lysine Methionine Phenylalanine helix 1. 42 1. 39 1. 11 1. 41 1. 14 1. 45 1. 13 strand 0. 83 1. 17 1. 10 1. 30 0. 74 1. 05 1. 38 turn 0. 66 0. 74 0. 98 0. 59 1. 01 0. 60 Subset of helix-lovers. If we forget alanine (I don’t understand that things affair with the helix at all), they share the presence of a (hydrophobic) C-b, C-g and C-d (S-d in Met). These hydrophobic atoms pack on top of each other in the helix. That creates a hydrophobic effect. ©CMBI 2006

Helix ©CMBI 2006

Secondary Structure Preferences Isoleucine Leucine Phenylalanine Threonine Tryptophan Tyrosine Valine helix 1. 08 1. 41 1. 13 0. 83 1. 08 0. 69 1. 06 strand 1. 60 1. 38 1. 19 1. 37 1. 47 1. 70 turn 0. 47 0. 59 0. 60 0. 96 1. 14 0. 50 Subset of strand-lovers. These residues either have in common their b-branched nature (Ile, Thr, Val) or their large and hydrophobic character (rest). ©CMBI 2006

Secondary Structure Preferences Aspartic Acid Asparagine Glycine Proline Serine helix 1. 01 0. 67 0. 57 0. 77 strand 0. 54 0. 89 0. 75 0. 55 0. 75 turn 1. 46 1. 56 1. 52 1. 43 Subset of turn-lovers. Glycine is special because it is so flexible, so it can easily make the sharp turns and bends needed in a b-turn. Proline is special because it is so rigid; you could say that it is prebend for the b-turn. Aspartic acid, asparagine, and serine have in common that they have short side chains that can form hydrogen bonds with the own backbone. These hydrogen bonds compensate the energy loss caused by bending the chain into a b-turn. ©CMBI 2006

Sequence Alignment Don’t forget that we still want to gather information about an unknown protein for which we determined the sequence. To gather that information, we will need databases and sequence alignments. To do these sequence alignments, we need to know everything about the amino acids. And that is what we are working on now. ©CMBI 2006

HYDROPHOBICITY ©CMBI 2006

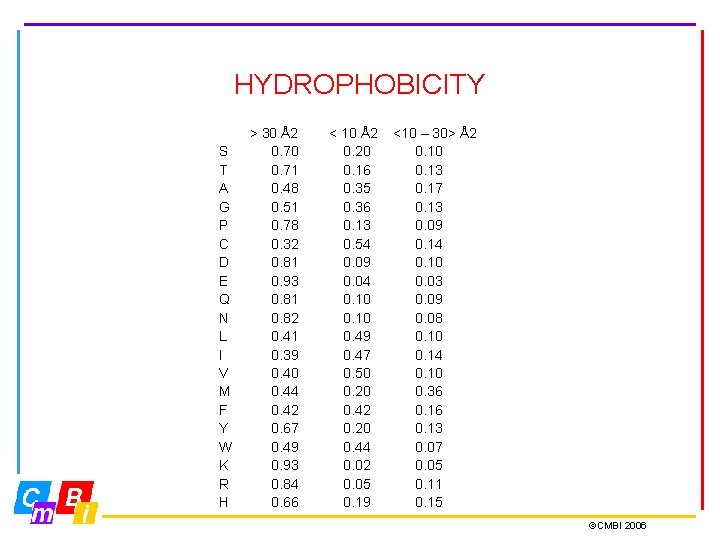
HYDROPHOBICITY S T A G P C D E Q N L I V M F Y W K R H > 30 Å2 0. 70 0. 71 0. 48 0. 51 0. 78 0. 32 0. 81 0. 93 0. 81 0. 82 0. 41 0. 39 0. 40 0. 44 0. 42 0. 67 0. 49 0. 93 0. 84 0. 66 < 10 Å2 0. 20 0. 16 0. 35 0. 36 0. 13 0. 54 0. 09 0. 04 0. 10 0. 49 0. 47 0. 50 0. 20 0. 42 0. 20 0. 44 0. 02 0. 05 0. 19 <10 – 30> Å2 0. 10 0. 13 0. 17 0. 13 0. 09 0. 14 0. 10 0. 03 0. 09 0. 08 0. 10 0. 14 0. 10 0. 36 0. 13 0. 07 0. 05 0. 11 0. 15 ©CMBI 2006