A DistributionFree Summarization Method for Affymetrix Gene Chip

A Distribution-Free Summarization Method for Affymetrix Gene. Chip Arrays Zhongxue Chen, Monnie Mc. Gee, Qingzhong Liu and Richard Scheuermann Dallas Area Bioinformatics Workshop August 29, 2006 DAB Workshop 2006

A new summarization method • • • Distribution Free Weighted (DFW) Summarization Use information on variability of probe intensities to summarize Affymetrix data Translate variability into weights which allow downweighting of poorly performing probes DAB Workshop 2006 2

Need for Summarization • • • Result of unique Affymetrix array structure Summarization is necessary to obtain one number for each gene All 11 - 20 probes interrogating each gene must be summarized into one expression value DAB Workshop 2006 3

Structure of Affymetrix Arrays • • Probe = sequence of 25 bases Probe pair = perfect match (PM) probe and its corresponding mismatch (MM) Probe set = 11 to 20 probe pairs interrogating one gene or EST Chips contain 6 K to 54 K probe sets Image courtesy of Affymetrix DAB Workshop 2006 4

PM and MM • • • PM = 25 base probe perfectly complementary to a specific region of a gene MM = 25 base probe agreeing with PM apart from middle base Middle base is a transition to Watson-Crick complement (A T, G C) DAB Workshop 2006 5

DFW • Transform probe-level intensities to log 2 scale for all arrays in experiment • • Arrange arrays in N by R matrix • • Stabilizes the variance (larger intensity increased variability N = total number of PM probes R = total number of arrays for entire experiment For each probe set, calculate a weight for each PM probe using Tukey biweight function Multiply weights by each probe intensity and summarize DAB Workshop 2006 6

Calculating Weights • • Calculate range of log intensities for each PM Find median of each range (M) Calculate distance of range to M for each PM (call this distance x) Weighting function: DAB Workshop 2006 7

Probe Weights • Weight for probe i is given by • J = number of probes in the probe set DAB Workshop 2006 8

More Calculations • Weighted Range (WR) • • • Range of weighted intensities Weighted Standard Deviation (WSD) Transformed Intensity Values (TIV) • Standardizes measures between DEGs and non-DEGs m and n should be positive integers DAB Workshop 2006 9
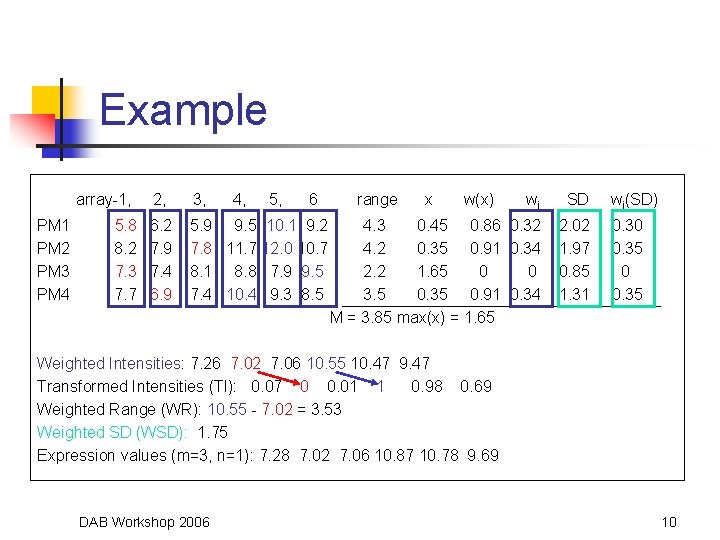
Example array-1, PM 1 PM 2 PM 3 PM 4 5. 8 8. 2 7. 3 7. 7 2, 3, 4, 5, 6 6. 2 7. 9 7. 4 6. 9 5. 9 9. 5 10. 1 9. 2 7. 8 11. 7 12. 0 10. 7 8. 1 8. 8 7. 9 9. 5 7. 4 10. 4 9. 3 8. 5 range x w(x) wi SD 4. 3 0. 45 0. 86 0. 32 4. 2 0. 35 0. 91 0. 34 2. 2 1. 65 0 0 3. 5 0. 35 0. 91 0. 34 M = 3. 85 max(x) = 1. 65 2. 02 1. 97 0. 85 1. 31 wi(SD) 0. 30 0. 35 Weighted Intensities: 7. 26 7. 02 7. 06 10. 55 10. 47 9. 47 Transformed Intensities (TI): 0. 07 0 0. 01 1 0. 98 0. 69 Weighted Range (WR): 10. 55 - 7. 02 = 3. 53 Weighted SD (WSD): 1. 75 Expression values (m=3, n=1): 7. 28 7. 02 7. 06 10. 87 10. 78 9. 69 DAB Workshop 2006 10

Why Weight? • Some PMs may have poor behavior • • Use information across arrays • • Give small or 0 weight to “poor” PM Assess quality of PM based on overall behavior SD of range provides information for detecting differentially expressed genes DAB Workshop 2006 11
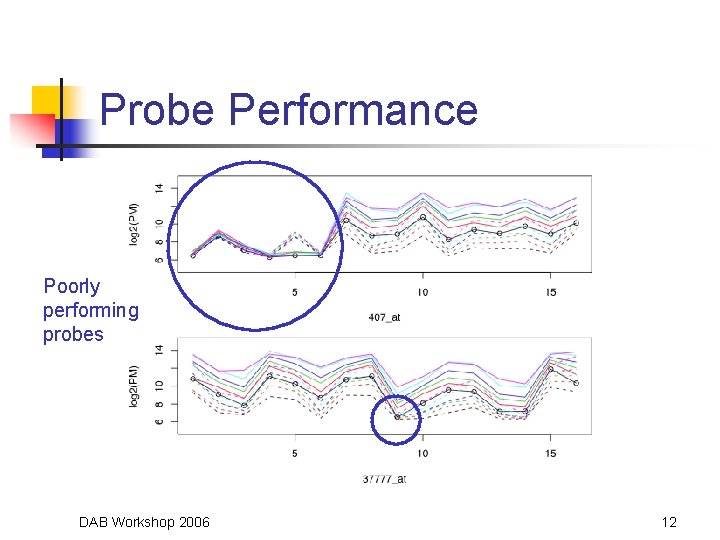
Probe Performance Poorly performing probes DAB Workshop 2006 12

Comparison Data Sets • Affymetrix Latin Square Spike-In Experiments • • • Two experiments: on HGU-95 Av 2 platform and HGU-133 A platform HGU-95 experiments has 14 transcripts spiked-in at concentrations from 0 to 1024 p. M (59 arrays) HGU-133 experiment has 42 transcripts spiked-in in triplicate at concentrations from 0 to 512 (42 arrays) Mc. Gee and Chen (2006) report 22 more spike-ins “Golden. Spike” Experiment (Choe et al. , 2005) • • • Six arrays (3 experiment, 3 control) on Dros. Genome 1 Chip 1309 transcripts recognizing known fold differences (from 1. 2 to 4) 2551 recognizing transcripts included at the same concentration DAB Workshop 2006 13

Comparison Methods • • ROC curves, AUC values and CPU time Competitors: • Robust Multichip Average (RMA) • • Gene Chip RMA (GCRMA) • • Affymetrix 2001, 2004 Model-Based Expression Index (MBEI) • • Wu et al. , 2004 MAS 5. 0, PLIER • • Bolstad, 2004; Irizarry et al. , 2003 Li & Wong, 2001 a, b Factor Analysis for Robust Array Summarization (FARMS) • Hochreiter et al. , 2006 DAB Workshop 2006 14
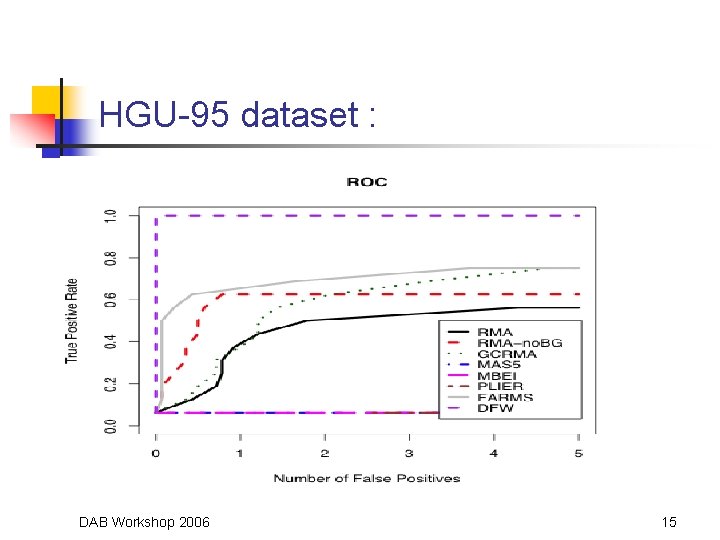
HGU-95 dataset : DAB Workshop 2006 15
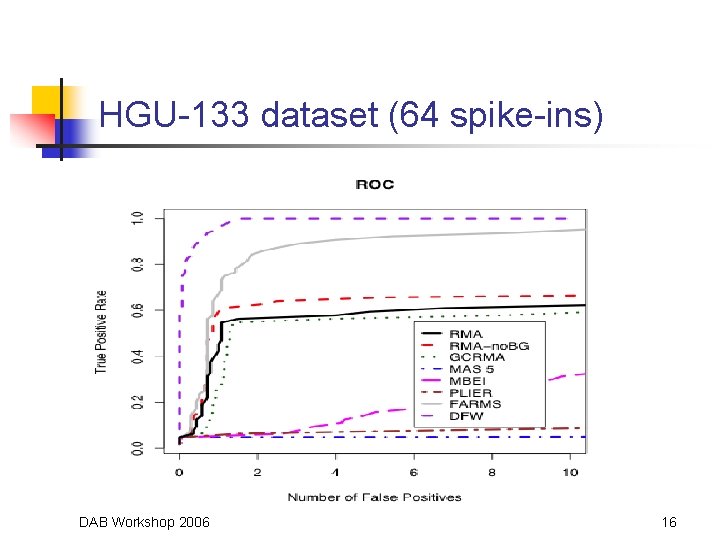
HGU-133 dataset (64 spike-ins) DAB Workshop 2006 16

“Preferred” Method • • • Choe et al. tested dozens of combinations of background correction, normalization, and summarization methods Preferred = the “best performing” method (according to DEGs obtained by Cyber. T - Baldi & Long, 2001) MAS 5. 0 background correction Quantile normalization median polish summarization second expression level normalization using LOESS procedure DAB Workshop 2006 17

Golden. Spike Data (FC = 1. 2) DAB Workshop 2006 18
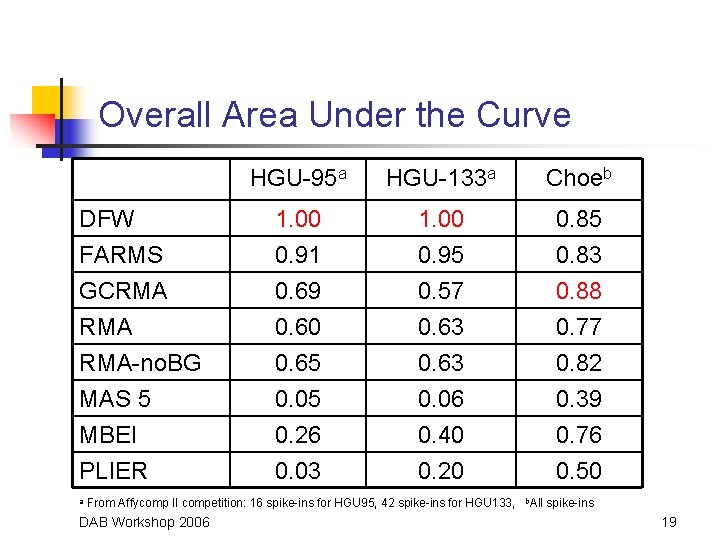
Overall Area Under the Curve DFW FARMS GCRMA RMA-no. BG MAS 5 MBEI PLIER a HGU-95 a HGU-133 a Choeb 1. 00 0. 91 0. 69 0. 60 0. 65 0. 05 0. 26 0. 03 1. 00 0. 95 0. 57 0. 63 0. 06 0. 40 0. 20 0. 85 0. 83 0. 88 0. 77 0. 82 0. 39 0. 76 0. 50 From Affycomp II competition: 16 spike-ins for HGU 95, 42 spike-ins for HGU 133, DAB Workshop 2006 b. All spike-ins 19
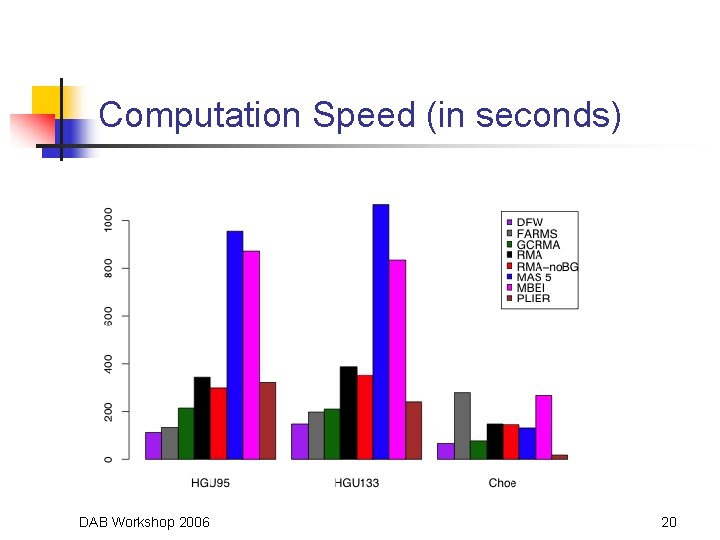
Computation Speed (in seconds) DAB Workshop 2006 20
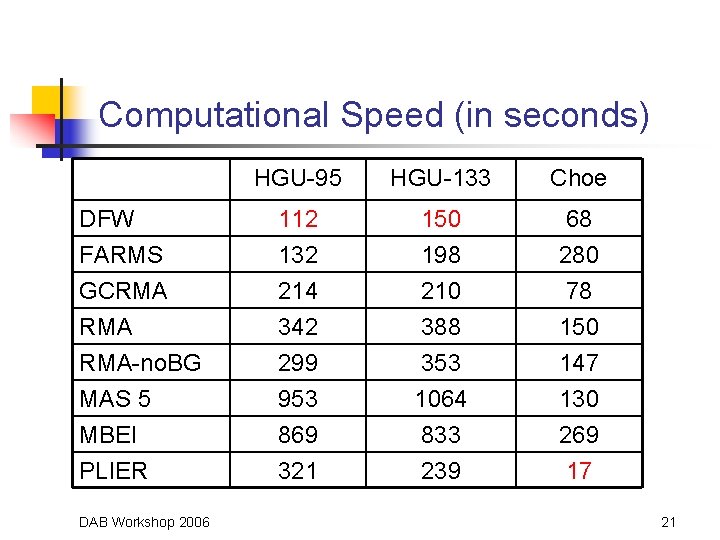
Computational Speed (in seconds) DFW FARMS GCRMA RMA-no. BG MAS 5 MBEI PLIER DAB Workshop 2006 HGU-95 HGU-133 Choe 112 132 214 342 299 953 869 321 150 198 210 388 353 1064 833 239 68 280 78 150 147 130 269 17 21

Further Comparisons • Affycomp II Competition Cope, et al. , 2004; Irizarry et al. , 2006 • For Hgu 95 spikein data, uses 16 spike-ins • For Hgu 133 spikein data, uses 42 spike-ins http: //affycomp. biostat. jhsph. edu/AFFY 2/TABLES. hgu/0. html • • SMU Technical Report http: //www. smu. edu/statistics/Tech. Reports/TR 344. pdf • Monnie Mc. Gee’s website http: //faculty. smu. edu/mmcgee DAB Workshop 2006 22

References Affymetrix, Inc. . (2002) Statistical algorithms description document. Affymetrix, Inc. (2005) Technical note: guide to probe logarithmic intensity error (PLIER) estimation. Baldi, P. and Long, A. D. (2001) A Bayesian framework for the analysis of microarray expression data: regularized t-test and statistical inferences of gene changes. Bioinformatics, 17, 509 -519. Bolstad, BM. (2004) Low Level Analysis of High-density oligonucleotide array data: Background, normalization and summarization [dissertation]. Department of Statistics, University of California at Berkeley. Choe, S. E. et al. (2005) Preferred analysis methods for Affymetrix Gene. Chips revealed by a wholly defined control datasets. Genome Biol. , 6, R 16. 1 -R 16. 6. Cope, L. M. et. al. (2004) A benchmark for Affymetrix Gene. Chip expression measures. Bioinformatics, 20, 323331. Hochreiter, S. et al. (2006) A new summarization method for Affymetrix probe level data. Bioinformatics, 22, 943949 Irizarry, R. A. et al. (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics, 4, 249 -264. Irizarry, R. A. et al. (2006) Comparison of Affymetrix Gene. Chip expression measures. Bioinformatics, 22, 789 -794. Li, C. and Wong, H. W. (2001 a) Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc. Nat. Acad. Sci. , 98, 31 -36. Li, C and Wong, H. W. (2001 b) Model-based analysis of oligonucleotide arrays: model validation, design issues and standard error application. Genome Biol. , 2, research 0032. 1 -0032. 11. Mc. Gee, M. and Chen, Z. (2006) New spiked-in probe sets for the Affymetrix HG-U 133 A Latin Square experiment. COBRA Preprint Series, Article 5 Wu, Z. et. al. (2004) A model-based background adjustment for oligonucleotide expression arrays. J. Am. Stat. Assoc. , 99, 909 -917. DAB Workshop 2006 23
- Slides: 23