Rule Bender A Tutorial Outline n Rulebased Modeling

+ Rule. Bender A Tutorial

+ Outline n Rule-based Modeling n Bio. Net. Gen Language (BNGL) n Rule. Bender

+ Rule-based Modeling n n Molecules n Types n Names n Initial Concentrations Molecular Interactions n Reactants and Products n Reaction Directions n Reaction Rates

+ Rule-based Modeling A Simple Model: Toy-Jim n Molecules – Data Objects: n n n Ligand L Receptor R Adaptor A Kinase K Molecular Interactions - Rules: n n n n L can bind to R Two R can dimerize if they are bound to L A can bind R , regardless of whether it is bonded to L/dimerized or not A can bind K , regardless of its phosphorylation state K can be phosphorylated When bound to A, one K can transphorylate the other …
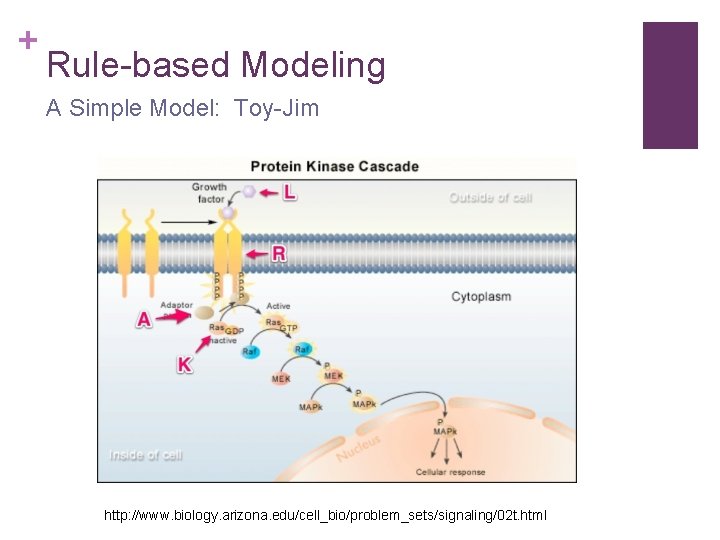
+ Rule-based Modeling A Simple Model: Toy-Jim http: //www. biology. arizona. edu/cell_bio/problem_sets/signaling/02 t. html

+ Rule-based Modeling A Simple Model: Toy-Jim n Rules: n n n Problems n n L + R <-> LR LR + LR <-> LRLR A + R <-> AR A + K <-> AK … How to express a bond? How to express a molecule’s binding state? How to express phosphorylation state? Solution: Bio. Net. Gen Language
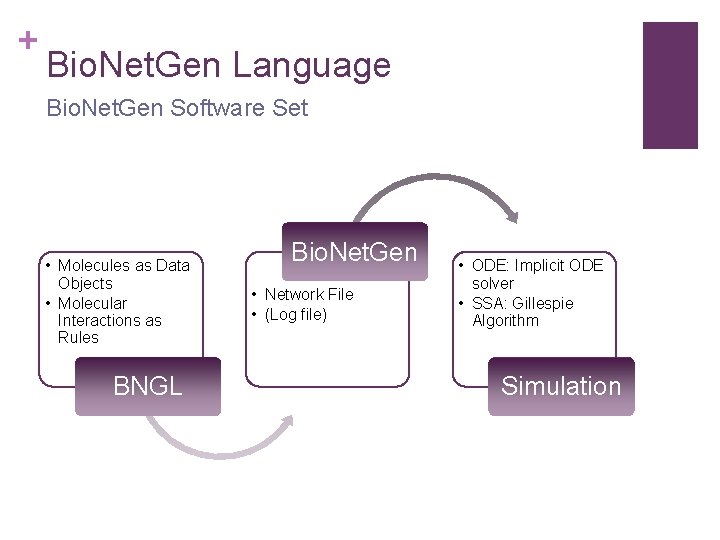
+ Bio. Net. Gen Language Bio. Net. Gen Software Set • Molecules as Data Objects • Molecular Interactions as Rules BNGL Bio. Net. Gen • Network File • (Log file) • ODE: Implicit ODE solver • SSA: Gillespie Algorithm Simulation

+ Bio. Net. Gen Language Elements of a BNGL program n A BNGL program consists of following blocks: n Parameters n Molecule Types n Seed Species n Reaction Rules n Observables n Actions

+ Bio. Net. Gen Language Molecule Types n n Surrounded by n begin molecule types n end molecule types Declare a Molecule: In toy-jim. bngl: begin molecule types 1 L(r) n Molecule name 2 R(l, r, a) n List of Components in Parentheses 3 A(r, k) n Tilde character (‘~’) after the component to declare the state of the component n ALL possible components and states should be declared 4 K(a, Y~U~P) begin molecule types

+ Bio. Net. Gen Language Seed Species n n Molecules / Complexes and concentrations present at the initial time Surround by n n n begin seed species end seed species Bindings to form a Complex n n n Use ‘. ’ to concatenate two molecules Use ‘!’ followed by a bond name (Integer) to declare the binding position Each bond name must occur TWO times within a species. begin seed species L(r) 0 R(l, r, a) R 0 A(r, k) A 0 K(a, Y~U) K 0 end seed species
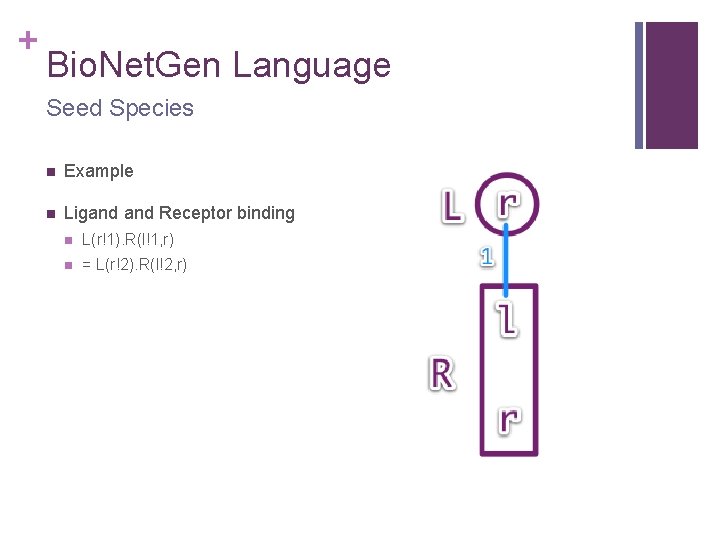
+ Bio. Net. Gen Language Seed Species n Example n Ligand Receptor binding n L(r!1). R(l!1, r) n = L(r!2). R(l!2, r)
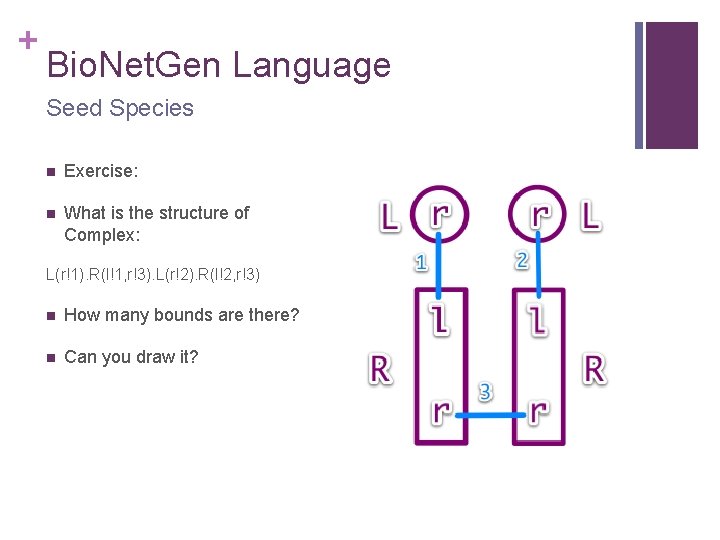
+ Bio. Net. Gen Language Seed Species n Exercise: n What is the structure of Complex: L(r!1). R(l!1, r!3). L(r!2). R(l!2, r!3) n How many bounds are there? n Can you draw it?

+ Bio. Net. Gen Language Reaction Rules n n Surrounded by n begin reaction rules n end reaction rules n Edge names may be given with wildcards to select various connectivity n “? ”: a bond may or may not be present n “+”: a bond must be present Molecules - Pattern Matching Molecules / Complexes do NOT have to be fully specified n Components of defined molecules may be missing n n K(Y~P) matches K(a, Y~P) State labels may be absent n K(a, Y) matches K(a, Y~U) and K(a, Y~P) n Other molecules in a complex may be absent n R(r!+) matches all dimerized R

+ Bio. Net. Gen Language Reaction Rules n Exercise: n Which of the following does pattern A(r!+, k!? ) match? 1. A(r!1). R(a!1) 2. A(r, k!2). K(a!2, Y~P) 3. R(a!1). A(r!1, k!2). K(a!2) 4. A(r!+, k!+) 5. A(r!? , k!? ) 6. L(r!8). R(l!8, a!4, r!7). A(r!4). R(r!7, l!11). L(r!11) 7. A. A

+ Bio. Net. Gen Language Reaction Rules n Declare a Reaction n Direction n One direction: -> n Both directions: <-> n Reactants: Left hand side n Products: Right hand side n Reaction rates

+ Bio. Net. Gen Language Reaction Rules begin reaction rules # Ligand Receptor Binding L(r) + R(l, r) <-> L(r!1). R(l!1, r) kp. L, km. L # Receptors can dermize if bounded to Ligand L(r!1). R(l!1, r) + L(r!1). R(l!1, r) <-> L(r!1). R(l!1, r!3). L(r!2). R(l!2, r!3) kp. D, km. D # Adaptor and Receptor binding A(r) + R(a) <-> A(r!1). R(a!1) kp. A, km. A # Adaptor and Kinase binding, regardless of phosphorylation state A(k) + K(a) <-> A(k!1). K(a!1) kp. K, km. K

+ Bio. Net. Gen Language Reaction Rules # Kinase transphorylation K(Y~U) -> K(Y~U). K(Y~P) p. K # Kinase transphorylation K(Y~P). K(Y~U) -> K(Y~P) p. Ks # Dephosphylation in membrane complex R(a!1). A(r!1, k!2). K(a!2, Y~P) -> R(a!1). A(r!1, k!2). K(a!2, Y~U) d. M # Dephosphylation in cytosol K(a, Y~P) -> K(a, Y~U) end reaction rules d. C
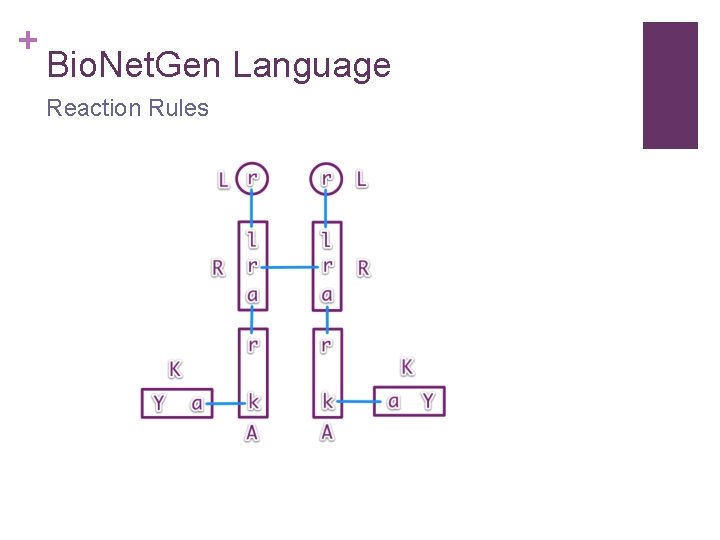
+ Bio. Net. Gen Language Reaction Rules

+ Bio. Net. Gen Language Reaction Rules n Exercise n What is the reaction describing n n An Adapter with a bond to a dermized Receptor n Binds n A Kinase One direction, reaction rate kp R(r!+, a!1). A(r!1, k)+K(a) -> R(r!+, a!1). A(r!1, k!2). K(a!2) kp

+ Bio. Net. Gen Language Parameters n n Surrounded by n begin parameters n end parameters Defines parameters in n Initial concentrations n Reaction Rates begin parameters # initial concentrations L 0 1 R 0 1 … # reaction rates kp. L 0. 1 km. L 0. 1 … end parameters

+ Bio. Net. Gen Language Observables n Surrounded by n n n begin observables end observables Type: n Species n Fully defined molecules/ Complexies n Molecules n Weighted sum over Species matched by the pattern begin parameters Molecules Rec. Dim R(r!+) Molecules Rec_A R(a!1). A(r!1) … Molecules L_tot L … end parameters n Name n Pattern

+ Bio. Net. Gen Language Actions n Generate Network n n Simulation with an ODE solver n n Simulation Parameters: n n simulate_ode Simulation with Stochastic Simulation Algorithm n n generate_network n simulate_ssa n n Set up a simulation n simulate_xxx({param 1=>val 1, par am 2=>val 2, …}) n t_end n Simulation end time n_steps n Number of intervals at which to report concentrations atol, rtol n Absolute error tolerance n Relative error tolerance sample_times n Times at which to report concentrations suffix/prefix n The suffix/prefix of the result file

+ Rule. Bender The Graphical Interface of BNGL
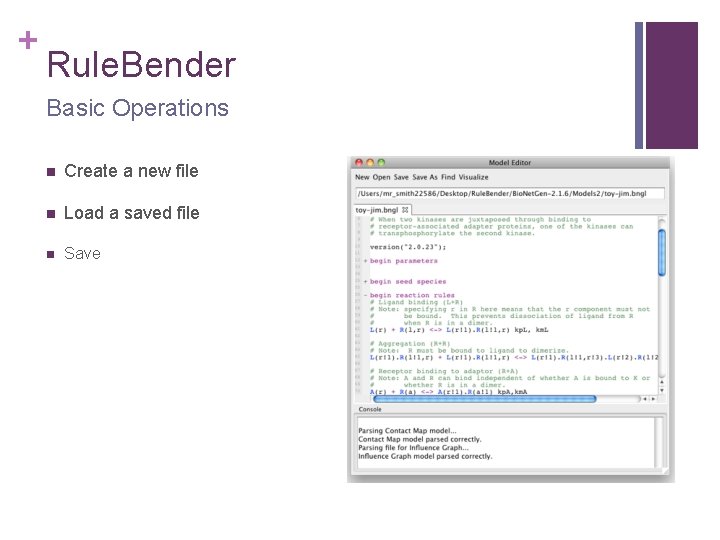
+ Rule. Bender Basic Operations n Create a new file n Load a saved file n Save

+ Rule. Bender Visualizing a Model n Contact Map n Connections between Molecules n Click a Component can get its States information n Click a Bond between two molecules can get the associated Reaction information (Reactants, Products, Reaction Rates)

+ Rule. Bender Visualizing a Model n Influence Graph n Influences between two reactions n Reactions with one direction n Reactions with two directions
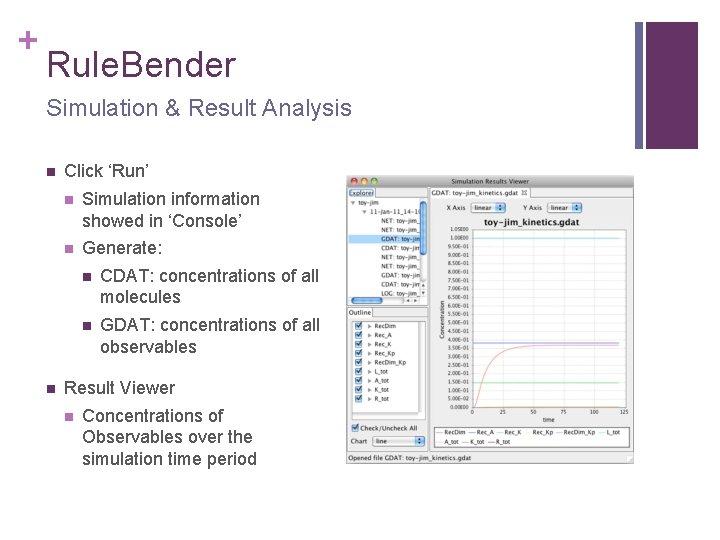
+ Rule. Bender Simulation & Result Analysis n n Click ‘Run’ n Simulation information showed in ‘Console’ n Generate: n CDAT: concentrations of all molecules n GDAT: concentrations of all observables Result Viewer n Concentrations of Observables over the simulation time period
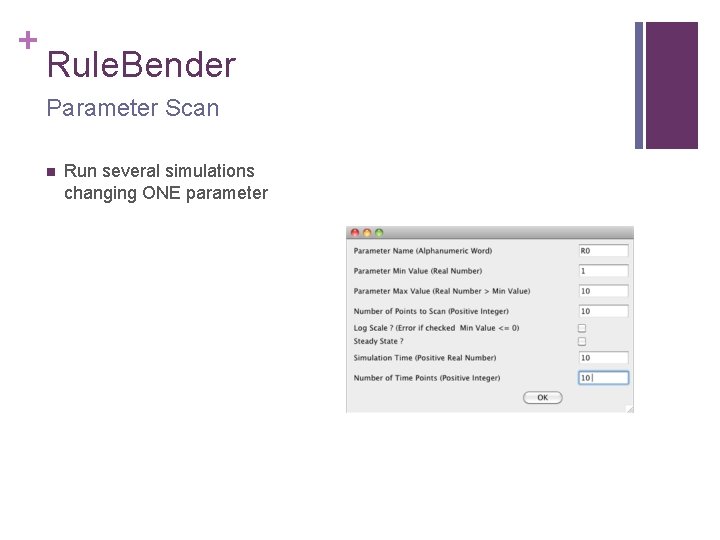
+ Rule. Bender Parameter Scan n Run several simulations changing ONE parameter

+ Rule. Bender Parameter Scan n Report how the changing of the parameter affects the other reactions n Plot a graph n Initial Value of the selected parameter vs. Concentrations of Observables

+ The END Thank You! 30
- Slides: 30