Optimisation of Multiple Sequence Alignments for Bacterial Identification

Optimisation of Multiple Sequence Alignments for Bacterial Identification By Manal Helal Centre for Infectious Diseases and Microbiology, Western Clinical School, University of Sydney and School of Computer Science, University of New South Wales
Why Multiple Sequence Alignment Compare a new sequence with the sequences in a protein family. Phylogenetic analysis: Gain insight into evolutionary relationships. Identify conserved domains/elements in sequences Compare regions of similarity among multiple organisms Identify probes for similar sequences in other organisms Develop PCR primers

Definition • A multiple alignment of strings S 1, … Sk is a series of strings with spaces S 1’, …, Sk’ such that • • |S 1’| = … = |Sk’| Sj’ is an extension of Sj by insertion of spaces • Goal: Find an optimal multiple alignment. MSCS 230: Bioinformatics I - Multiple Sequence Alignment 3
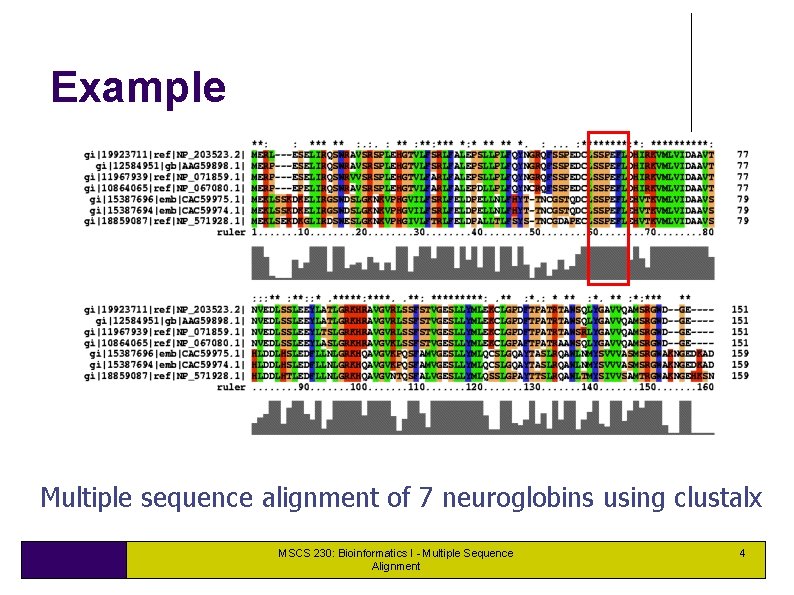
Example Multiple sequence alignment of 7 neuroglobins using clustalx MSCS 230: Bioinformatics I - Multiple Sequence Alignment 4

Problems with Existing Methods Assume Min. percent identity of ~40% for proteins and ~70% for DNA, otherwise, much higher likelihood of errors. Sensitive to sequences input order. Depend on pairwise alignments, which is less sensitive and cause bias in the positioning of gaps. Statistical Uncertainty. Assume the MSA outcome before doing it, using annotations and biological knowledge in the objective function, or adding homologs or profiles. Assume conserved order of aligned residues. ABA, Pro. DA, TBA, MAUVE don't assume this. Care must be made in choosing scoring matrices and penalties. MSCS 230: Bioinformatics I - Multiple Sequence Alignment 5

Clustal – A Common MSA Tool • Progressive alignment strategy: – Sequences used to make guide tree – Most similar two sequences aligned = consensus – Next closest sequence aligned to the consensus 13 MITTEN 1 2 3 4 MITTENS KITTIES SMITTEN KITTENS 1 -MITTENS 3 SMITTEN- 13 MITTEN 4 KITTENS 134 ITTEN Manual editing: ITT-E columns Fine adjustment 134 of particular 2 KITTIES Incorporate specific knowledge Removal of gappy bits Important for phylogenetic analysis Removal of parts of/whole sequences Non-homologous regions Sequences included by error Lab 4. 1 6

Other Progressive Approaches PILEUP Similar to CLUSTALW Uses UPGMA (Unweighted Pair Group Method with Arithmetic mean) to produce tree rather than Neighbor-joining method T-Coffee – Muscle, MAFT and others use different objective functions. MSCS 230: Bioinformatics I - Multiple Sequence Alignment 7

Dynamic Programming The dynamic programming approach can be adapted to MSA For simplicity, assume k sequences of length n The dynamic programming array F is kdimensional of length n+1 (including initial gaps) The entry F(i 1, …, ik) represents score of optimal alignment for s 1[1. . i 1], … sk[1. . ik] MSCS 230: Bioinformatics I - Multiple Sequence Alignment 8
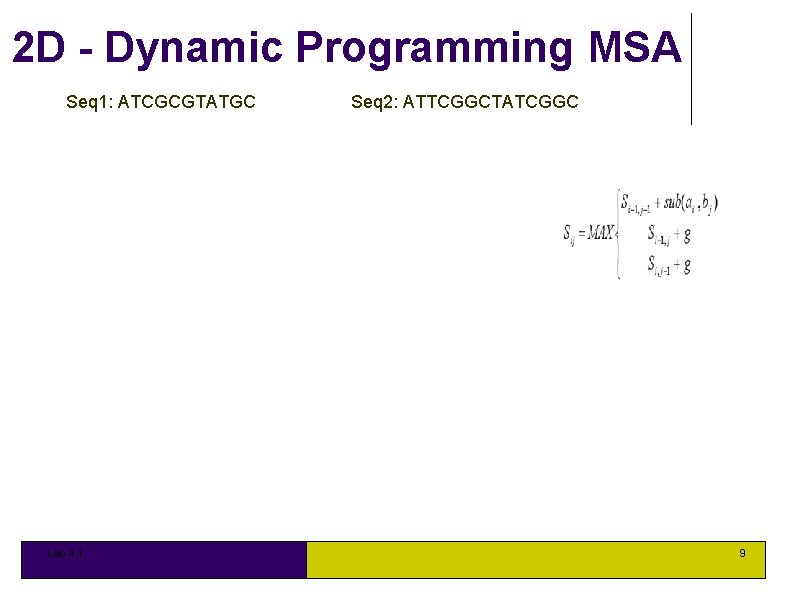
2 D - Dynamic Programming MSA Seq 1: ATCGCGTATGC Lab 4. 1 Seq 2: ATTCGGCTATCGGC 9
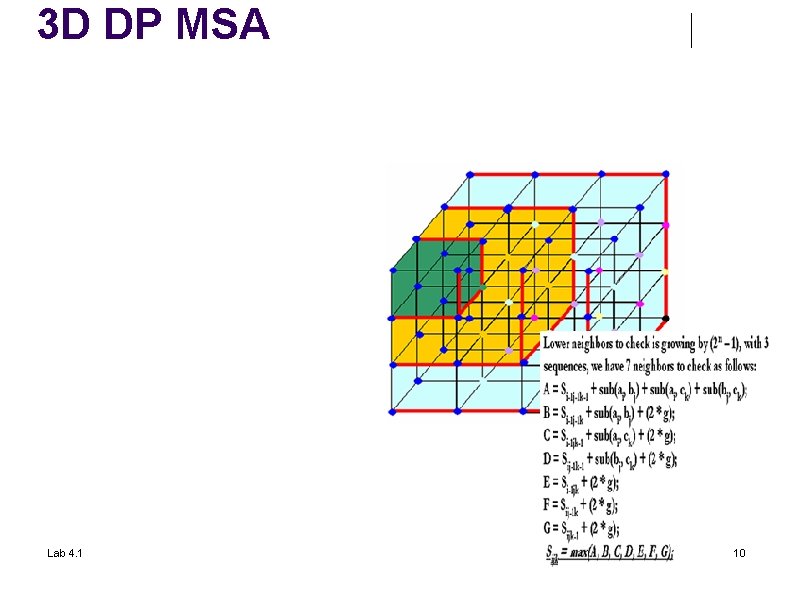
3 D DP MSA Lab 4. 1 10

Dynamic Multidimensional Array Indexing Mathematics of Arrays (Mo. A) represents arrays dynamically as a linear array in memory. The scoring tensor of any dimension is created where neighbouring and partitioning is done using algebraic transformations of the array index. Lab 4. 1 11

Performance Space Problem As sequences grow in number and lengths, scoring tensor won’t fit in cache This is solved by MOA partitioning to allow running smaller partitions of the scoring tensor in parallel processors. Execution Time Problem Again is addressed by parallelization and using bigger HPC or computer clusters.

Parallelization Technique Distributed MSA based on MOA is designed by retrieving diagonals of partitions that can be scored simultaneously in one wave of computation. Their dependencies are computed in an earlier wave of computation, and sent to the waiting processors.
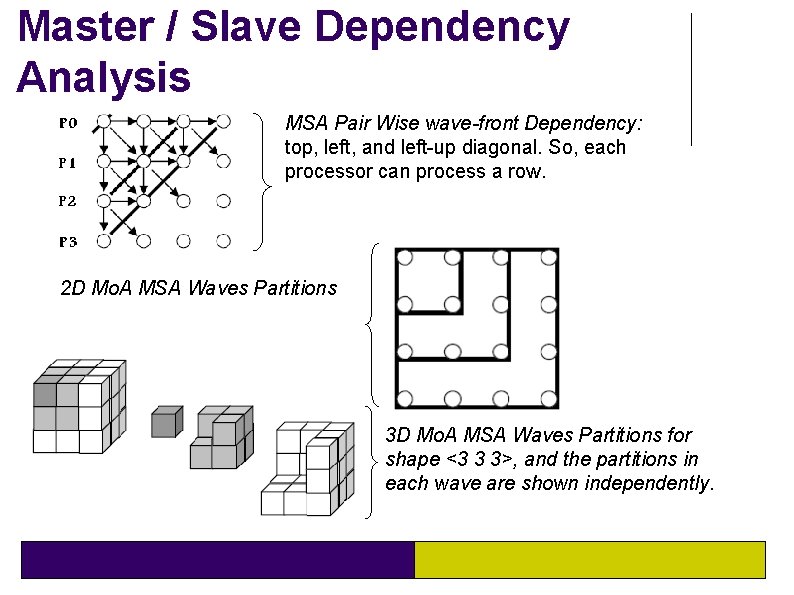
Master / Slave Dependency Analysis MSA Pair Wise wave-front Dependency: top, left, and left-up diagonal. So, each processor can process a row. 2 D Mo. A MSA Waves Partitions 3 D Mo. A MSA Waves Partitions for shape <3 3 3>, and the partitions in each wave are shown independently.

Distributed Scoring & Dependency S(i 0 i 1 i 2 i 3. . . ik) = max G 1 + TS (G 1) G 2 + TS (G 2) : G 2 k-1 + TS (G 2 k-1) Where: TS (Gi) = (sub(dj, dk) for each pair j, k in G) +( g. S * (K-D)) Gi: Neighbor i of current cell, up to 2 k-1 neighbors D: No of decremented indices to get this particular neighbor TS: Temporary Score function assigned to each neighbor based on how many multidimensional indices were decremented to get to this neighbor g. S: gap Score Value * (K-D): multiply the gap Score Value with number of indices that remained the same (were not decremented to get this neighbor), retrieved by Total Dimensions K (Sequences) – D.
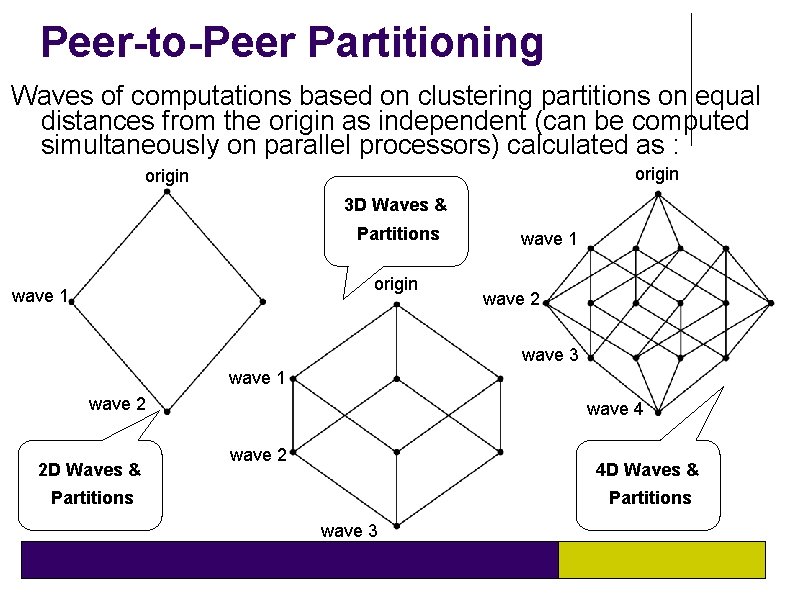
Peer-to-Peer Partitioning Waves of computations based on clustering partitions on equal distances from the origin as independent (can be computed simultaneously on parallel processors) calculated as : origin 3 D Waves & Partitions origin wave 1 wave 2 wave 3 wave 1 wave 2 2 D Waves & wave 4 wave 2 4 D Waves & Partitions wave 3
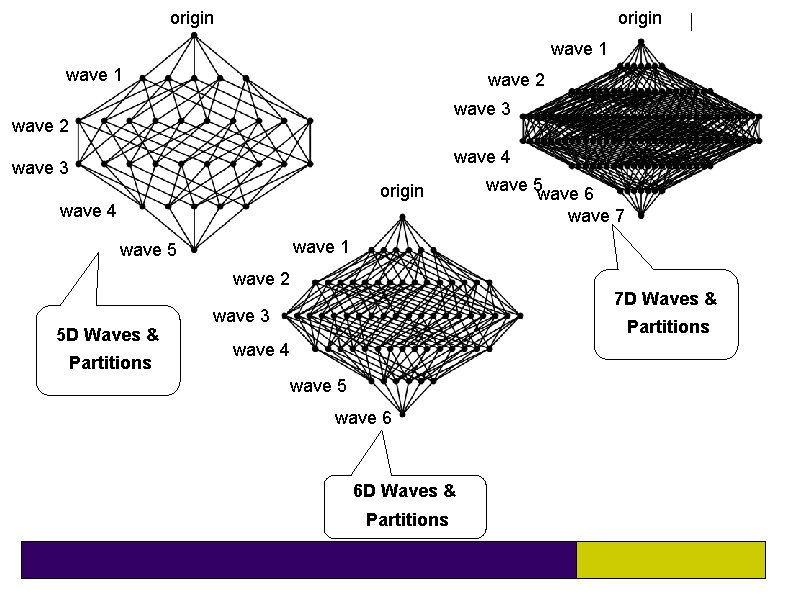
origin wave 1 wave 2 wave 3 wave 2 wave 4 wave 3 origin wave 4 wave 5 wave 6 wave 7 wave 1 wave 5 wave 2 5 D Waves & Partitions 7 D Waves & wave 3 Partitions wave 4 wave 5 wave 6 6 D Waves & Partitions
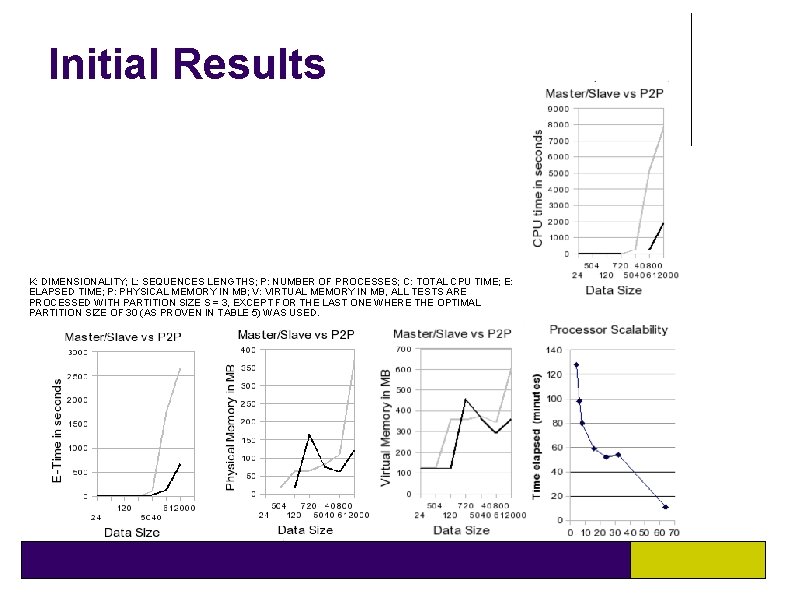
Initial Results K: DIMENSIONALITY; L: SEQUENCES LENGTHS; P: NUMBER OF PROCESSES; C: TOTAL CPU TIME; E: ELAPSED TIME; P: PHYSICAL MEMORY IN MB; V: VIRTUAL MEMORY IN MB, ALL TESTS ARE PROCESSED WITH PARTITION SIZE S = 3, EXCEPT FOR THE LAST ONE WHERE THE OPTIMAL PARTITION SIZE OF 30 (AS PROVEN IN TABLE 5) WAS USED.
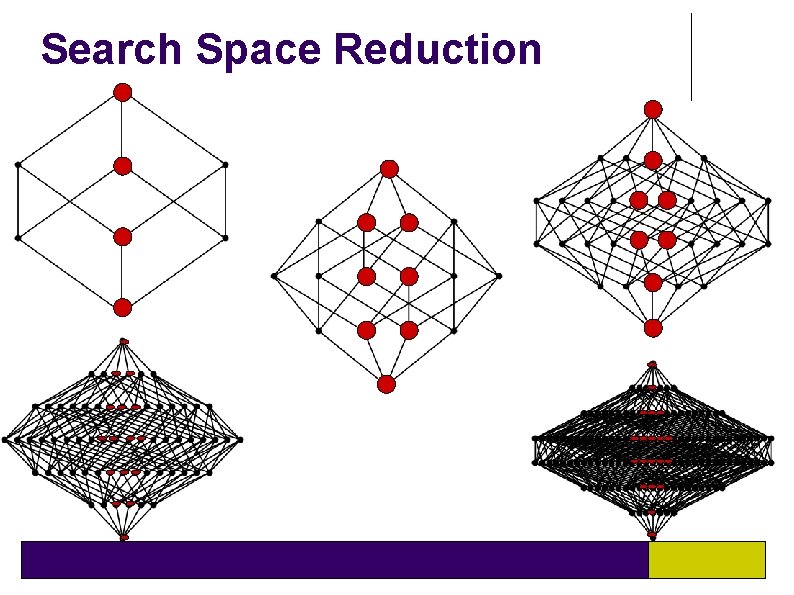
Search Space Reduction

Thanks
- Slides: 20