ModelBased Information Integration in a Neuroscience Mediator System

Model-Based Information Integration in a Neuroscience Mediator System Bertram Ludaescher Amarnath Gupta Maryann E. Martone University of California San Diego 1
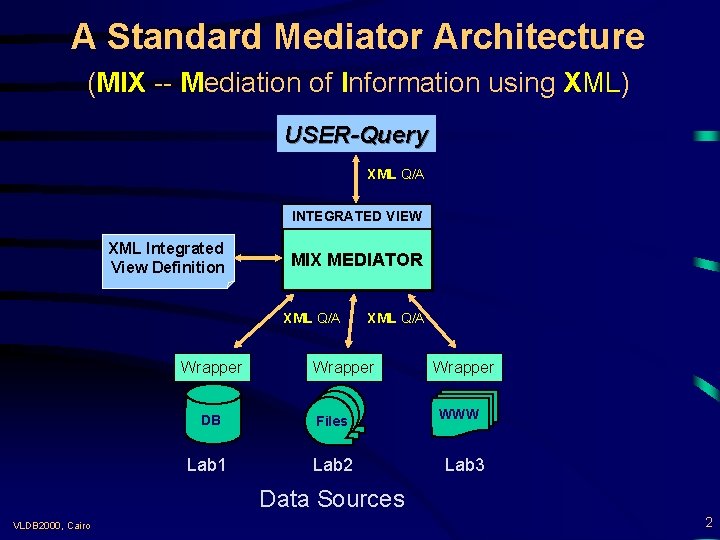
A Standard Mediator Architecture (MIX -- Mediation of Information using XML) USER-Query XML Q/A INTEGRATED VIEW XML Integrated View Definition MIX MEDIATOR XML Q/A Wrapper DB Files Lab 1 Lab 2 Wrapper WWW Lab 3 Data Sources VLDB 2000, Cairo 2

Integration Issues SYNTACTIC/STRUCTURAL Integration • Integrated Views (Src-XML => Intgr-XML) • Schema Integration (DTD =>DTD) • Wrapping, Data Extraction (Text => XML) TCP/IP HTTP CORBA VLDB 2000, Cairo MIX SYSTEM Integration Mediation of Information using XML SRB/MCAT Distributed Query Processing SEMANTIC Integration ? ? ? storage, query capabilities protocols & services 3

Integration Issues: Mediating across Multiple-Worlds • Structural Integration => common semistructured data model (XML) => XML queries & transformations to resolve schema conflicts • Limited Query Capabilities => mediator is aware of QCs exported by wrappers • . . . • Semantic Integration – most work deals with issues for “one-world” scenarios (e. g. , amazon. com vs. bn. com) – what if data comes from a “multiple-world” scenario (like Neuroscience), where data objects from different sources are not even similar, and only the hidden semantics (known to the domain expert) provides the “semantic link”? VLDB 2000, Cairo 4
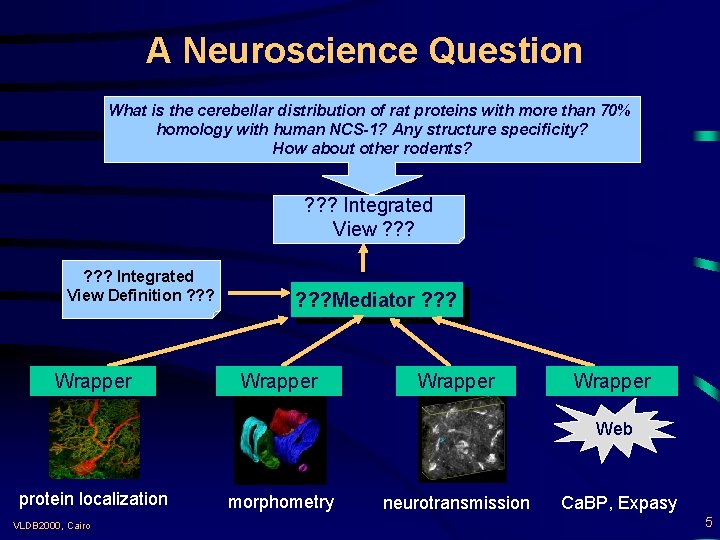
A Neuroscience Question What is the cerebellar distribution of rat proteins with more than 70% homology with human NCS-1? Any structure specificity? How about other rodents? ? ? ? Integrated View Definition ? ? ? Wrapper ? ? ? Mediator ? ? ? Wrapper Web protein localization VLDB 2000, Cairo morphometry neurotransmission Ca. BP, Expasy 5

Hidden Semantics: Protein Localization Purkinje Cell layer of Cerebellar Cortex <protein_localization> <neuron type=“purkinje cell” /> <protein channel=“red”> <name>Ry. R</> …. </protein> <region h_grid_pos=“ 1” v_grid_pos=“A”> <density> <structure fraction=“ 0. 8”> <name>spine</> <amount name=“Ry. R”>0</> Molecular layer of </> Cerebellar Cortex <structure fraction=“ 0. 2”> <name>branchlet</> Fragment of dendrite <amount name=“Ry. R”>30</> VLDB 2000, Cairo </> 6

Hidden Semantics: Morphometry <neuron name=“purkinje cell”> <branch level=“ 10”> Branch level beyond 4 <shaft> is a branchlet … </shaft> <spine number=“ 1”> <attachment x=“ 5. 3” y=“-3. 2” z=“ 8. 7” /> <length>12. 348</> <min_section>1. 93</> <max_section>4. 47</> <surface_area>9. 884</> <volume>7. 930</> <head> <width>4. 47</> Must be dendritic <length>1. 79</> because Purkinje cells </head> don’t have somatic spines </spine> 7 VLDB 2000, Cairo …

The Problem • Multiple Worlds Integration – compatible terms not directly joinable – complex, indirect associations among schema elements – unstated integrity constraints • Why not just use Ontologies? – typical ontologies associate terms along limited number of dimensions • What’s needed? – a “theory” under which non-identical terms can be “semantically joined” => lift mediation to the level of conceptual models (CMs) => domain knowledge, ICs become rules over CMs => Model-Based Mediation VLDB 2000, Cairo 8
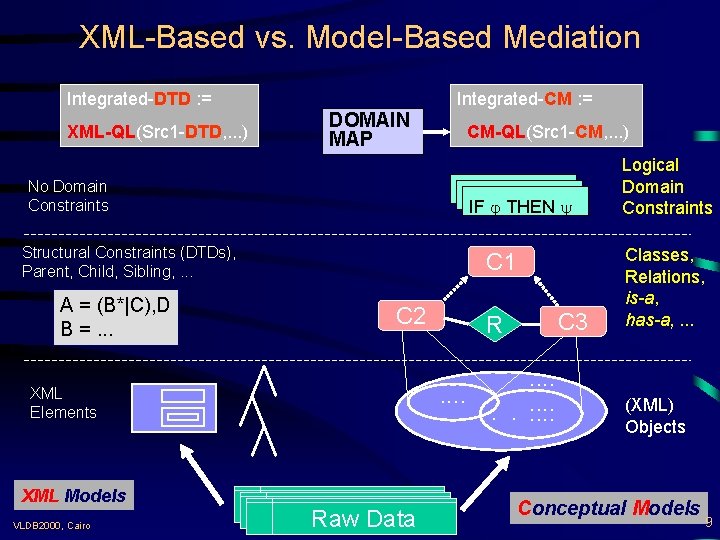
XML-Based vs. Model-Based Mediation Integrated-DTD : = XML-QL(Src 1 -DTD, . . . ) Integrated-CM : = DOMAIN MAP No Domain Constraints CM-QL(Src 1 -CM, . . . ) IF THEN IF IF THEN Structural Constraints (DTDs), Parent, Child, Sibling, . . . A = (B*|C), D B =. . . C 1 C 2. . XML Elements XML Models VLDB 2000, Cairo Raw Data Raw. Data C 3 R. . . Logical Domain Constraints Classes, Relations, is-a, has-a, . . . (XML) Objects Conceptual Models 9

Extended Mediator Architecture => Wrappers export Conceptual Models (CMs), i. e. , facts+rules for classes, relationships, ICs, . . . ) => Mediator imports CMs (from sources, auxiliary knowledge bases, and domain maps (DMs) => a generic conceptual model (GCM, a subset of F-logic), extensible via rules = common target CM language => new CMs can be plugged-in by specifying them in GCM + Flogic rules => prototype implementation in FLORA: • global-as-view approach • compiler: F-logic => XSB-Prolog • top-down evaluation => virtual (demand-driven) views • external interfaces (XML, RDBs, DM visualization, . . . ) VLDB 2000, Cairo 10
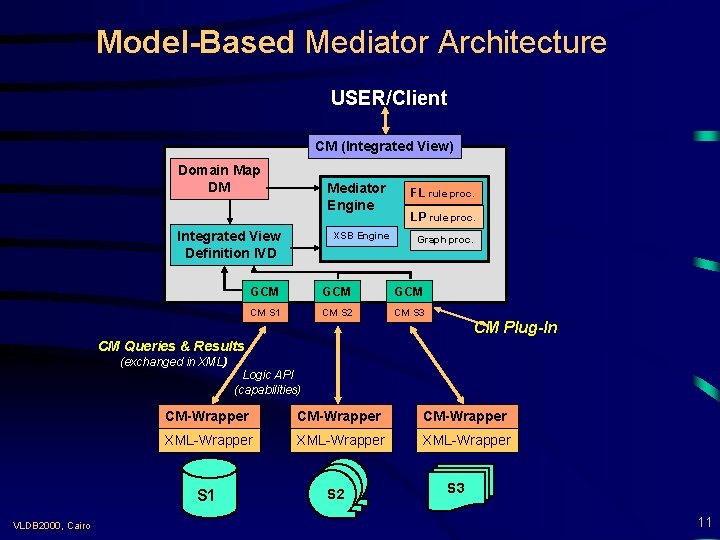
Model-Based Mediator Architecture USER/Client CM (Integrated View) Domain Map DM Mediator Engine Integrated View Definition IVD XSB Engine FL rule proc. LP rule proc. Graph proc. GCM GCM CM S 1 CM S 2 CM S 3 CM Plug-In CM Queries & Results (exchanged in XML) CM-Wrapper XML-Wrapper S 1 VLDB 2000, Cairo Logic API (capabilities) S 2 S 3 11

Definition of Integrated Views. . . • XML-2 -FL and CM-2 -FL Translators <!ELEMENT Studies (Study)*> <!ELEMENT Study (study_id, … animal, experiments, experimenters> <!ELEMENT experiments (experiment)*> <!ELEMENT experiment (description, instrument, parameters)> study. DB[studies =>> study]. study[study_id => string; … animal => animal; experiments =>> experiment; experimenters =>> string]. … • Specification of Domain Knowledge • Subclasses mushroom_spine : : spine S: mushroom_spine IF S: spine[head _; neck _]. • Rules ic 1(S): alert[type “invalid spine”; object S] IF S: spine[undef ->> {head, neck}]. • Integrity Constraints • Integrated View Definition protein_distribution(Protein, Organism, Brain_region, Feature_name, Anatom, Value) IF I: protein_label_image[ proteins ->> {Protein}; organism -> Organism; anatomical_structures ->> {AS: anatomical_structure[name->Anatom]}], NAE: neuro_anatomic_entity[name->Anatom; loccated_in->>{Brain_region}], AS. . segments. . features[name->Feature_name; value->Value]. VLDB 2000, Cairo 12
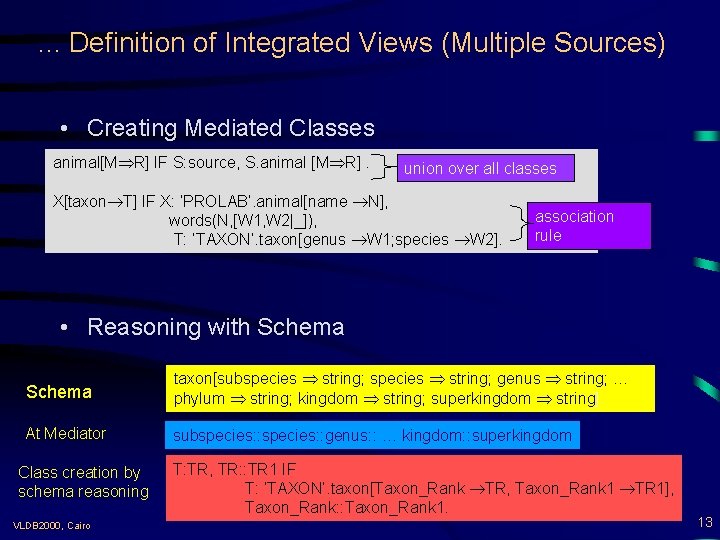
. . . Definition of Integrated Views (Multiple Sources) • Creating Mediated Classes animal[M R] IF S: source, S. animal [M R]. union over all classes X[taxon T] IF X: ‘PROLAB’. animal[name N], words(N, [W 1, W 2|_]), T: ‘TAXON’. taxon[genus W 1; species W 2]. association rule • Reasoning with Schema taxon[subspecies string; genus string; … phylum string; kingdom string; superkingdom string]. At Mediator subspecies: : genus: : … kingdom: : superkingdom Class creation by schema reasoning VLDB 2000, Cairo T: TR, TR: : TR 1 IF T: ‘TAXON’. taxon[Taxon_Rank TR, Taxon_Rank 1 TR 1], Taxon_Rank: : Taxon_Rank 1. 13

Model-Based Mediation with DOMAIN MAPS (DMs) • “Semantic Road Maps” for situating source data => navigational aid (browsing source classes at the conceptual level) => basis for integrated views across multiple worlds => link points (concepts) and labeled arcs (roles) => formal semantics (in FL and/or DLs) Example: ANATOM DM = antatomical entities (concepts) + is_a, has_a, overlaps, . . . (roles) => from syntactic equality to semantic joins LINK(X, Y): X. zip = Y. zip X. addr in Y. zip X. zip overlaps Y. county. . . VLDB 2000, Cairo Integrated-CM(Z 1, . . . ) : = get X 1, . . . from Src 1; get X 2, . . . from Src 2; LINK (Xi, Yj); Zj = CM-QL(X 1, . . . , Y 1, . . . ) 14
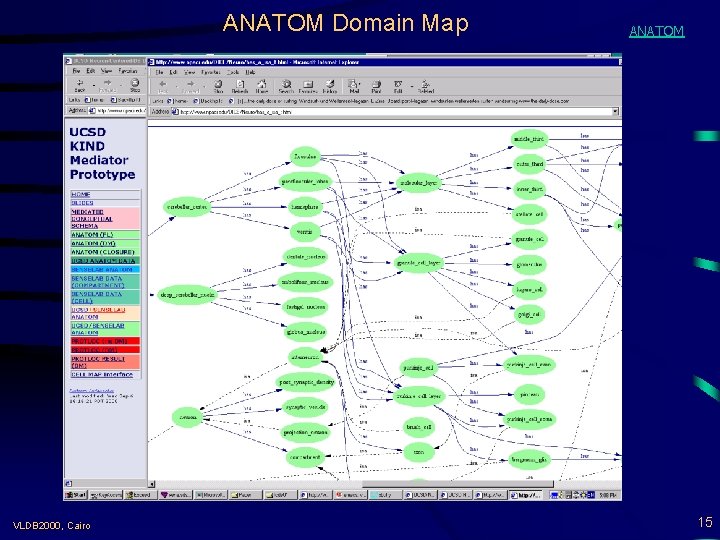
ANATOM Domain Map VLDB 2000, Cairo ANATOM 15
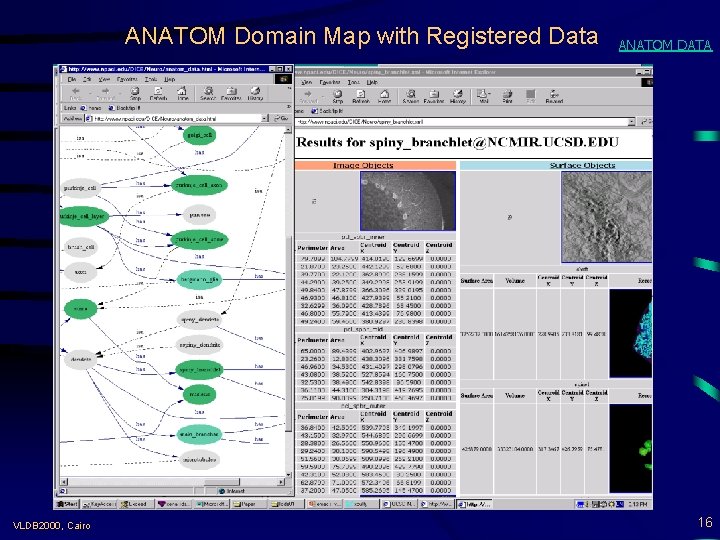
ANATOM Domain Map with Registered Data VLDB 2000, Cairo ANATOM DATA 16
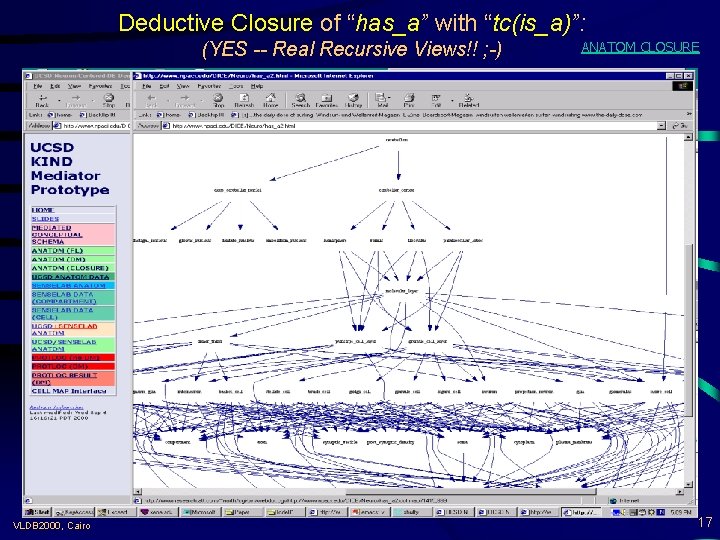
Deductive Closure of “has_a” with “tc(is_a)”: (YES -- Real Recursive Views!! ; -) VLDB 2000, Cairo ANATOM CLOSURE 17

Example Query Evaluation (I) • Example: protein_distribution – given: organism, protein, brain_region – ANATOM DM: • recursively traverse the has_a_star paths under brain_region collect all anatomical_entities – Source PROLAB: • join with anatomical structures and collect the value of attribute “image. segments. feature. protein_amount” where “image. segments. feature. protein_name” = protein and “study_db. study. animal. name” = organism – Mediator: • aggregate over all parents up to brain_region • report distribution VLDB 2000, Cairo 18
Interactive Queries (I) VLDB 2000, Cairo KIND 19

Example Query Evaluation (II) "How does the parallel fiber output (Yale/SENSELAB) relate to the distribution of Ryanodine Receptors (UCSD/NCMIR)? " @SENSELAB: X 1 : = select output from parallel fiber ; @MEDIATOR: X 2 : = “hang off” X 1 from Domain Map; @MEDIATOR: X 3 : = subregion-closure(X 2); @NCMIR: X 4 : = select PROT-data(X 3, Ryanodine Receptors); @MEDIATOR: X 5 : = compute aggregate(X 4); VLDB 2000, Cairo 20
Interactive Queries (II) VLDB 2000, Cairo KIND 01 21
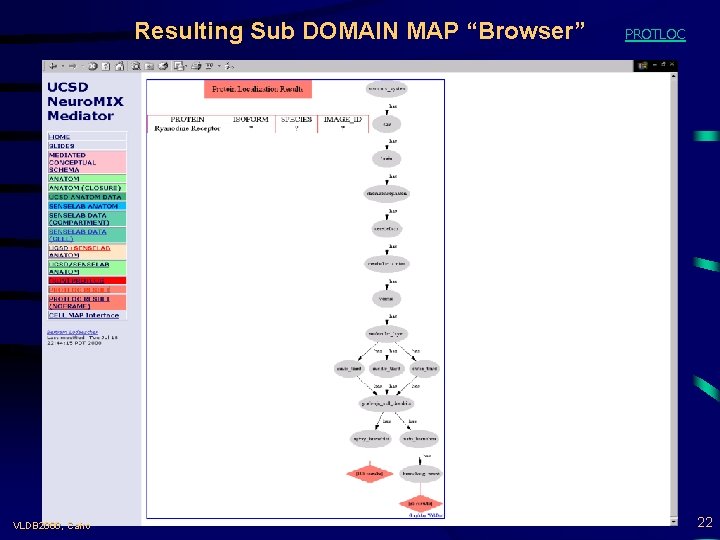
Resulting Sub DOMAIN MAP “Browser” VLDB 2000, Cairo PROTLOC 22
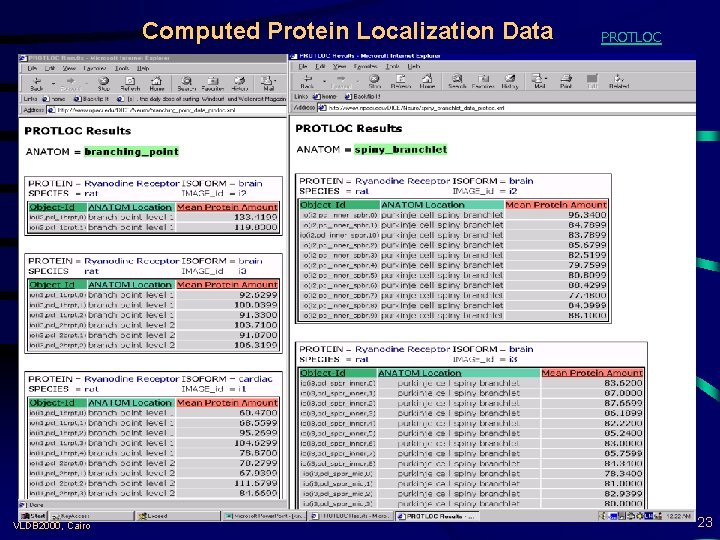
Computed Protein Localization Data VLDB 2000, Cairo PROTLOC 23
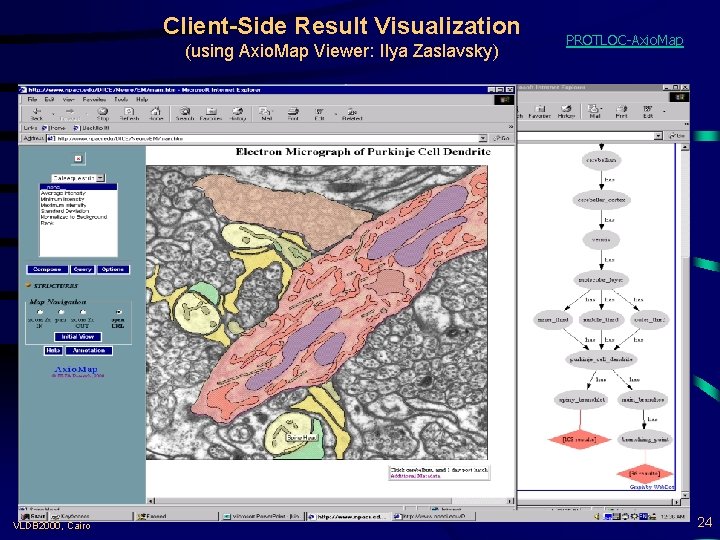
Client-Side Result Visualization (using Axio. Map Viewer: Ilya Zaslavsky) VLDB 2000, Cairo PROTLOC-Axio. Map 24
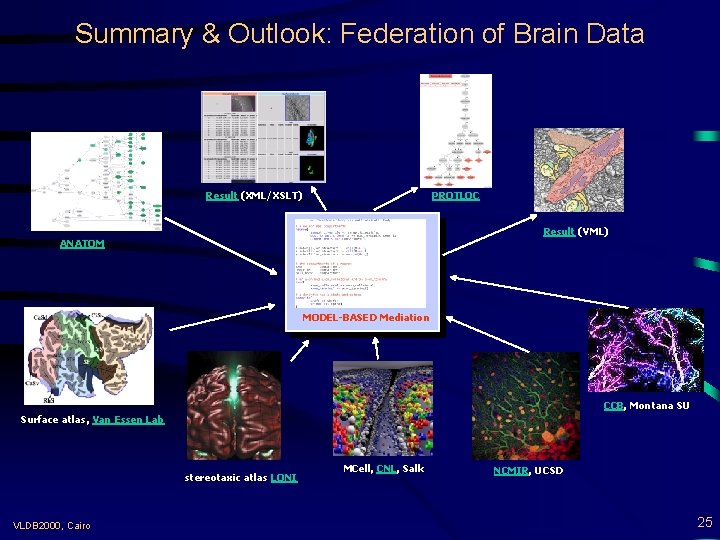
Summary & Outlook: Federation of Brain Data Result (XML/XSLT) PROTLOC Result (VML) ANATOM MODEL-BASED Mediation CCB, Montana SU Surface atlas, Van Essen Lab stereotaxic atlas LONI VLDB 2000, Cairo MCell, CNL, Salk NCMIR, UCSD 25
- Slides: 25