Introduction to NMRbox Project and NMRbox Virtual Machine

Introduction to NMRbox Project and NMRbox Virtual Machine Mark Maciejewski UConn Health “Think inside the box” UMD 2017 @NMRbox

Outline Lecture • • Motivation for the project NMRbox platform Benefits to users and developers Usage Hands on (at the beginning of the tutorial session) - Adam • • • Account management Connect to NMRbox VM Set the display resolution Software inventory File transfers UMD 2017 @NMRbox

Motivation: Abundance of software Hundreds of packages cited in Bio. Mag. Res. Bank depositions, J. Bio-NMR, and other journals. UMD 2017 @NMRbox
Motivation: Fragmentation Operating systems Programming languages Libraries BLAS UMD 2017 @NMRbox
Motivation: Persistency Platforms become obsolete Developers graduate Software time bombs Grants end UMD 2017 @NMRbox
Motivation: Meta-software packages NMRPipe Sparky Python scripts Rosetta SHIFTX 2 MODELLER Experimental protein structure verification by scoring with a single, unassigned NMR spectrum. Courtney, Rienstra, et al. , Structure, 2015. UMD 2017 @NMRbox

Motivation: Computational reproducibility A computational study is reproducible when it provides the “complete software environment needed to reproduce the figures” - D. Donoho, Stanford Obstacles • • • Missing primary empirical data Missing meta-data Missing software (scripts, programs) Non-persistence of software Manual interventions UMD 2017 @NMRbox

Challenges • Abundance of software (discovery) • Fragmentation of OSes, programming languages, libraries • Persistence of resources • Complexity of design and installation • Reproducibility of results Question How do we address these challenges? Answer NMRbox VM UMD 2017 @NMRbox

Deliverables – primary tools Platform • NMRbox VM: A virtual machine pre-configured with a wide range of software used in biological NMR • Significant computational resources Data • BMRB integration & richer depositions • Metadata management and workflow annotation Analytics • Bayesian tools to enhance data analysis and interpretation • API for developers to incorporate Bayesian inference UMD 2017 @NMRbox

Deliverables – community services Training and Dissemination • Workshops, tutorials, and guides • User and developer support Driving Biological Projects (DBPs) • Test beds for NMRbox technology development • What limits your progress? Collaboration and Service (C&S) • Apply technologies to challenging biomedical research problems UMD 2017 @NMRbox

NMRbox VM. What’s included? Acquisition Agnostic – Install software available Access Persistent – Archive all versions Content – Software packages 100+ packages installed (see https: //nmrbox. org) • • • Spectral reconstruction Spectral visualization Automated assignment Structure determination Molecular visualization Validation UMD 2017 • • • Chemical shift prediction Dynamics Residual dipolar coupling Meta packages General purpose Instrument manufactures @NMRbox

NMRbox VM. What’s included? Content – Productivity Tools • • • over a dozen editors scientific python packages R and R tools office tools drawing tools Octave shells browsers Dropbox UMD 2017 OS xubuntu 16. 04 @NMRbox

NMRbox VM. What’s included? Release 3 features added • GPUs to support 3 D drawing • Py. MOL, VMD, Chimera, and others • GPUs to support CUDA processing • NAMD, others coming soon • Commercial software • data. Chord spectrum Analyst, data. Chord spectrum Miner, Mest. Re. Nova • Matlab compiled binaries • ALATIS, GUARDD, TITAN • virtual on-screen keyboard See Release notes at https: //nmrbox. org/files/release-notes-version-3 -0. pdf UMD 2017 @NMRbox

Virtual Machine Terminology VM = A software-based emulation of a guest computer backed by the physical resources of a host computer, managed by a hypervisor. Access • Local installation (standalone or downloadable) • Connect to server (Paa. S = Platform-as-a-Service) Advantages • • • Over-subscribe the host computer Snapshot the VM and restore to any point Run multiple OS’s on a single computer “spin-up” VMs in minutes Dynamically load balance VMs across multiple hosts No performance penalties on modern computers UMD 2017 @NMRbox
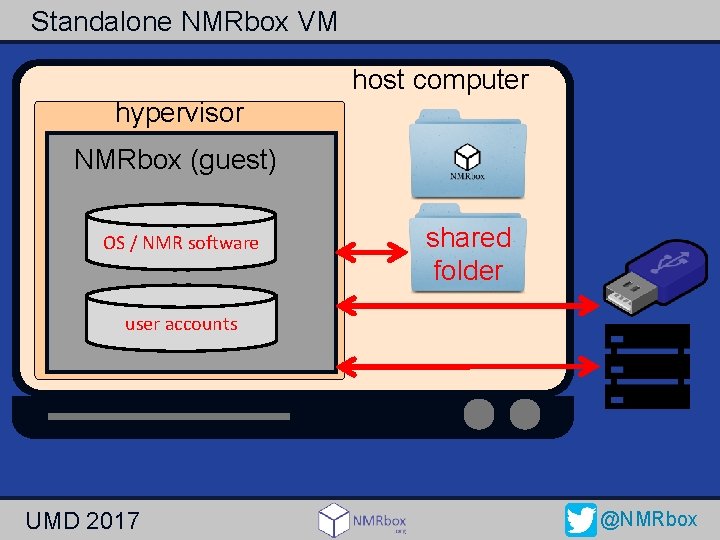
Standalone NMRbox VM host computer hypervisor NMRbox (guest) OS / NMR software shared folder user accounts UMD 2017 @NMRbox
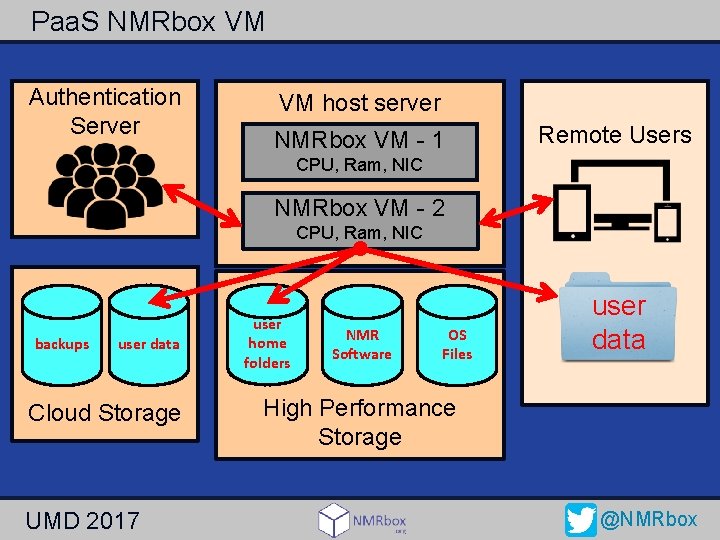
Paa. S NMRbox VM Authentication Server VM host server NMRbox VM - 1 Remote Users CPU, Ram, NIC NMRbox VM - 2 CPU, Ram, NIC backups user data Cloud Storage UMD 2017 user home folders NMR Software OS Files user data High Performance Storage @NMRbox
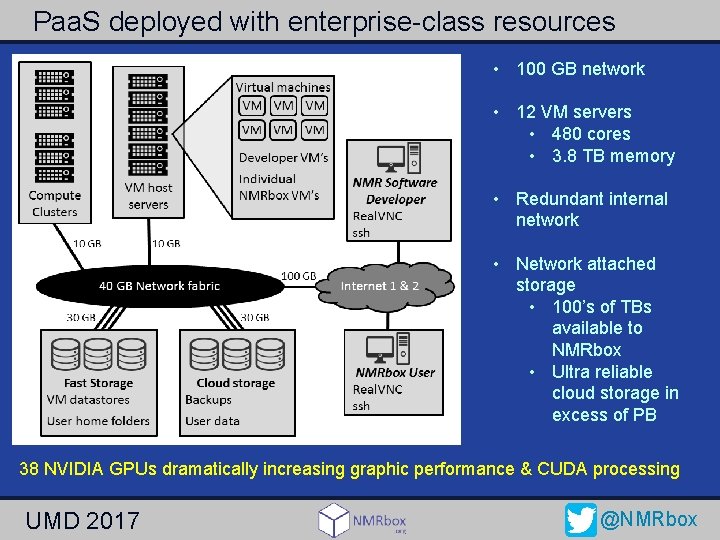
Paa. S deployed with enterprise-class resources • 100 GB network • 12 VM servers • 480 cores • 3. 8 TB memory • Redundant internal network • Network attached storage • 100’s of TBs available to NMRbox • Ultra reliable cloud storage in excess of PB 38 NVIDIA GPUs dramatically increasing graphic performance & CUDA processing UMD 2017 @NMRbox

VM Requirements for Users Standalone VM • 64 -bit hardware (Windows, OSX, Linux, …) • any modern laptop and desktop Oracle Virtual. Box VMware Workstation VMware Fusion VMware Player Server based Paa. S VM • ssh or VNC (Windows, OSX, Linux, tablet, phone, 32 -bit hardware, …) • Network connection UMD 2017 @NMRbox

Benefits Users Developers • • • • “Zero-configuration” Access Training Computational resources Discovery Persistence Reproducibility Cost Single platform Discovery Usage metrics Persistence Community Developer tools Computational resources Instructors • Access to NMRbox VMs for courses and workshops UMD 2017 @NMRbox

Practical aspects Large VM model • NMRbox VMs configured with many cores, high memory, and GPUs • Multiple users per VM, each user has two VMs (username. nmrbox. org and username 2. nmrbox. org) • CPU and memory utilization restricted to 50% of full VM • GPUs restrict VM management Updates • Additional software will be added to “live” NMRbox VMs • Version numbers updated • All states archived • Software versions updated on major releases • Older major VM releases continue to run with reduced resources at version. nmrbox. org Backups • User data backed up daily UMD 2017 @NMRbox

Practical aspects Large memory VM • A large memory VM can be “spun-up” for users if needed Home folder and archive folder • Each user has two home folders; /home/nmrbox/username and /nmr/archive/username Google Group • We have started a Google Group at https: //groups. google. com Search for NMRbox to join. Support • Email support@nmrbox. org Downloadable version • Downloadable version in final testing UMD 2017 @NMRbox

Practical aspects Host workshops with NMRbox VMs • The NMRbox team will “spin-up” custom VMs to support other workshops File permissions and access • Home and archive folders are not accessible to others by default. Will setup lab groups if desired. • /public folder for quick sharing Contact us • Suggestions for packages to include • Suggestions about the package • Issues with the NMRbox platform UMD 2017 @NMRbox
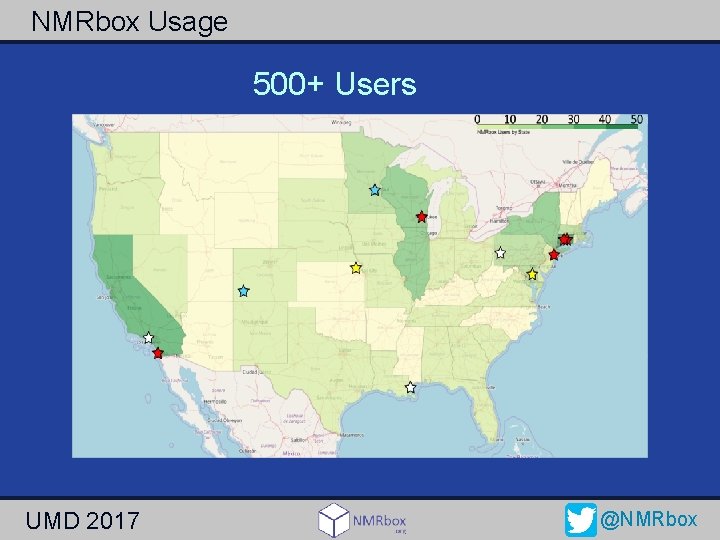
NMRbox Usage 500+ Users UMD 2017 @NMRbox
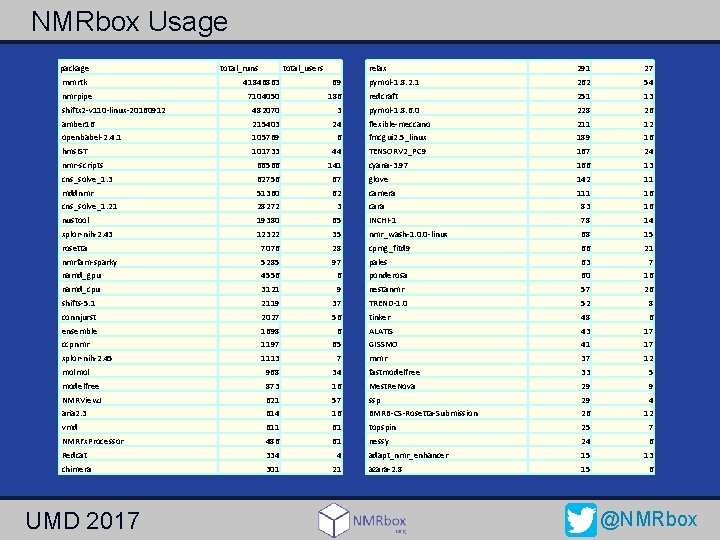
NMRbox Usage package rnmrtk nmrpipe total_runs total_users 41846863 69 7104050 186 relax 291 27 pymol-1. 8. 2. 1 262 54 redcraft 251 13 shiftx 2 -v 110 -linux-20160912 482070 3 pymol-1. 8. 6. 0 228 26 amber 16 215403 24 flexible-meccano 211 12 openbabel-2. 4. 1 105769 6 fmcgui 2. 5_linux 189 16 hms. IST 101733 44 TENSORV 2_PC 9 167 24 nmr-scripts 66566 141 cyana-3. 97 166 13 cns_solve_1. 3 62756 67 glove 142 11 mddnmr 51360 62 camera 111 16 cns_solve_1. 21 28272 3 cara 83 16 nustool 19380 65 INCHI-1 78 14 xplor-nih-2. 43 12322 35 nmr_wash-1. 0. 0 -linux 68 15 rosetta 7076 28 cpmg_fitd 9 66 21 nmrfam-sparky 5285 97 pales 63 7 namd_gpu 4556 6 ponderosa 60 16 namd_cpu 3121 9 nestanmr 57 26 shifts-5. 1 2119 37 TREND-1. 0 52 8 connjurst 2027 56 tinker 48 6 ensemble 1698 6 ALATIS 43 17 ccpnmr 1197 65 GISSMO 41 17 xplor-nih-2. 45 1113 7 rnmr 37 12 molmol 968 34 fastmodelfree 33 5 modelfree 873 16 Mest. Re. Nova 29 9 NMRView. J 621 57 ssp 29 4 aria 2. 3 614 16 BMRB-CS-Rosetta-Submission 26 12 vmd 611 61 topspin 25 7 NMRFx. Processor 486 61 nessy 24 6 Redcat 334 4 adapt_nmr_enhancer 15 13 chimera 301 21 azara-2. 8 15 6 UMD 2017 @NMRbox

Cite NMRbox Very Important!! If you utilize NMRbox in your research please cite and acknowledge us. Details at https: //nmrbox. org NMRbox: A Resource for Biomolecular NMR Computation. Maciejewski, M. W. , Schuyler, A. D. , Gryk, M. R. , Moraru, I. I. , Romero, P. R. , Ulrich, E. L. , Eghbalnia, H. R. , Livny, M. , Delaglio, F. , and Hoch, J. C. , Biophys J. , 112: 1529 -1534, 2017. [PMID: 28445744, DOI: 10. 1016/j. bpj. 2017. 03. 011] "This study made use of NMRbox: National Center for Biomolecular NMR Data Processing and Analysis, a Biomedical Technology Research Resource (BTRR), which is supported by NIH grant P 41 GM 111135 (NIGMS). " UMD 2017 @NMRbox
- Slides: 25