Goals of tutorial Introduce NMRbox platform Showcase NMRbox

Goals of tutorial • Introduce NMRbox platform • Showcase NMRbox with NUS tools • A dozen different NUS processing tools installed and configured – more coming. • Demonstrate potential of NMRbox • Now that the platform is maturing we can focus attention to enhancing tools, providing training materials, etc. • For this tutorial – Created a prototype, autonus, for wrapping NUS tools into a single package. • Learn some information about NUS • NUS can improve: • Time – if there is sensitivity to spare • Resolution – Collect to longer time increments without increase experiment time • Sensitivity per unit time – Focus more sampling to increments with smaller time increments without sacrificing resolution • Learn about features / characteristics of different techniques • Example scripts for various NUS techniques

Logistics of tutorial • In the tutorial all commands are shown in quotes • “autonus dft-nmrpipe” • Tutorial assumes a basic knowledge of terminal commands, NMRPipe, and NMRDraw • All data in the tutorial is 3 D with two indirect NUS dimensions. • Processed data is saved as 3 D NMRPipe data and 2 D projections. It is helpful to examine both and to view horizontal and vertical slices through peaks. • For the HNCACB planes 79 and 167 will be used to illustrate characteristics of the different NUS programs. • You can work alone or in pairs • GET DATA
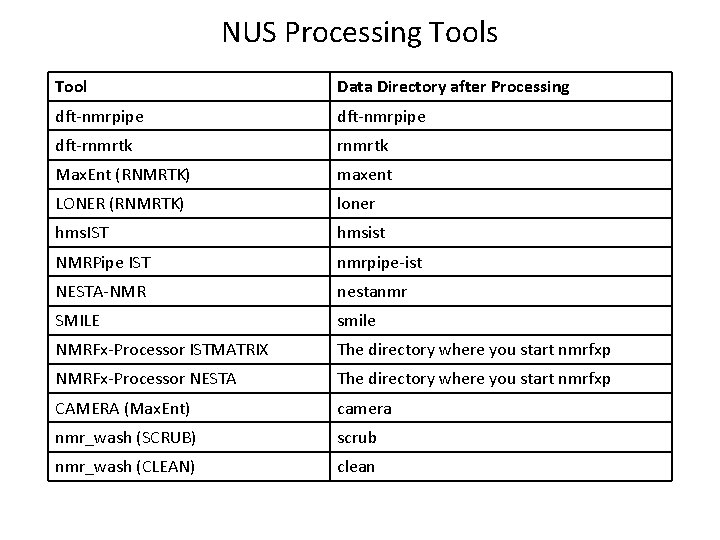
NUS Processing Tools Tool Data Directory after Processing dft-nmrpipe dft-rnmrtk Max. Ent (RNMRTK) maxent LONER (RNMRTK) loner hms. IST hmsist NMRPipe IST nmrpipe-ist NESTA-NMR nestanmr SMILE smile NMRFx-Processor ISTMATRIX The directory where you start nmrfxp NMRFx-Processor NESTA The directory where you start nmrfxp CAMERA (Max. Ent) camera nmr_wash (SCRUB) scrub nmr_wash (CLEAN) clean

Differences in NUS techniques • • • Acquisition dimension (DFT) maxent & loner: Output frequency – no DFT of indirect dimensions scrub & clean: Start with nu. DFT spectrum and scrub sampling artifacts hms. IST, NMRPipe-IST, CAMERA, NESTA-NMR, SMILE, NMRFX-Processor IST & NESTA, MDD & MDD-CS, : “fill-in” missing time domain data – data processed with DFT afterwards Keep or replace experimental data? • Keeping experimental data is akin to overfitting the data Phases: Some tools are independent of phase, some need to know the phase during reconstruction, and some will not work properly with a first order phase correction. Extending data beyond last collected point Deconvolution: Max. Ent can deconvolve linewidth and J-couplings Non-linearity

HNCACB 12 k. Da four helix bundle Experimental setup • grpdly = 67. 9862060546875 t 1 t 2 t 3 Nucleus N CACB HN Echo-Antiecho no no no Reference 119. 087 47. 742 4. 773 Phases 0, 0 90, 0 ZF size 256 1024 FT options alt, neg alt Extract Region 9. 52 – 5. 9 ppm Sampling Max increment 800 points (hyper-complex) (21%) 50 76 1024 (complex) 1024
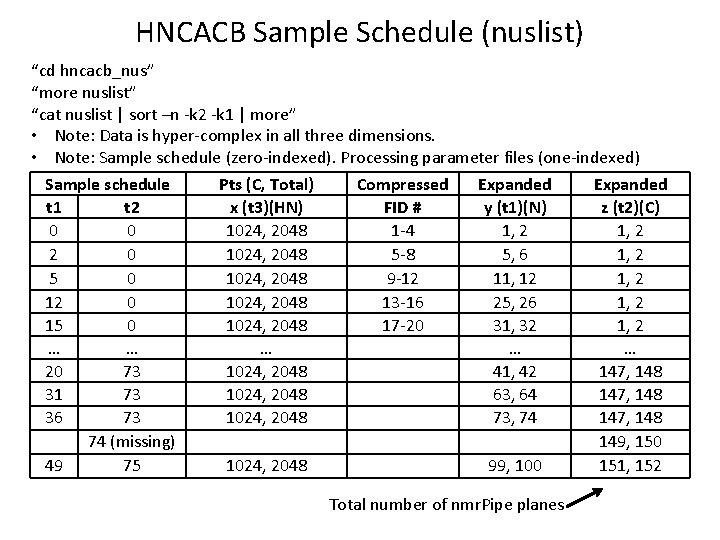
HNCACB Sample Schedule (nuslist) “cd hncacb_nus” “more nuslist” “cat nuslist | sort –n -k 2 -k 1 | more” • Note: Data is hyper-complex in all three dimensions. • Note: Sample schedule (zero-indexed). Processing parameter files (one-indexed) Sample schedule t 1 t 2 0 0 2 0 5 0 12 0 15 0 … … 20 73 31 73 36 73 74 (missing) 49 75 Pts (C, Total) x (t 3)(HN) 1024, 2048 1024, 2048 … 1024, 2048 Compressed FID # 1 -4 5 -8 9 -12 13 -16 17 -20 Expanded y (t 1)(N) 1, 2 5, 6 11, 12 25, 26 31, 32 … 41, 42 63, 64 73, 74 99, 100 Total number of nmr. Pipe planes Expanded z (t 2)(C) 1, 2 1, 2 … 147, 148 149, 150 151, 152

HNCACB Sample Schedule (nuslist) Sample schedule with x axis t 1 and y axis t 2. Red dots represent data collected. The max value in t 1 is 49 (50) and the max value in t 2 is 75 (76). Total coverage is 21%

Expanding NUS data We will see if a bit how we expand NUS data, but lets take a look at some NUS data that is already expanded. “cd HNCACB_nus/data” “nmr. Draw -in test%03 d. ft 3” For z (t 2) planes 1 and 2 we see identical FIDs at y (t 1) positions of 1, 2, 5, 6, 11, 12, etc. “Goto z plane 147 and FIDs for y (t 1 ) positions of 41, 42, 63, 64, 73, and 74 Thus, The NUS data has been fully expanded with zeros for any FIDs not in the sample schedule and the collected time domain data for points in the sample schedule. Benefits: • Perform fixes for sensitivity enhancement during conversion – leads to consistency in processing parameters • Allows a nu. DFT for a quick view of the data. Expanded y (t 1)(N) 1, 2 5, 6 11, 12 25, 26 31, 32 … 41, 42 63, 64 73, 74 99, 100 Expanded z (t 2)(C) 1, 2 1, 2 … 147, 148 149, 150 151, 152
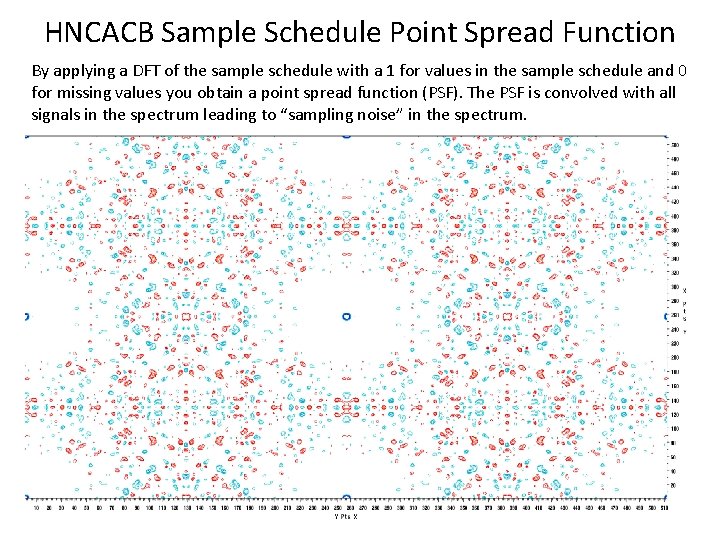
HNCACB Sample Schedule Point Spread Function By applying a DFT of the sample schedule with a 1 for values in the sample schedule and 0 for missing values you obtain a point spread function (PSF). The PSF is convolved with all signals in the spectrum leading to “sampling noise” in the spectrum.
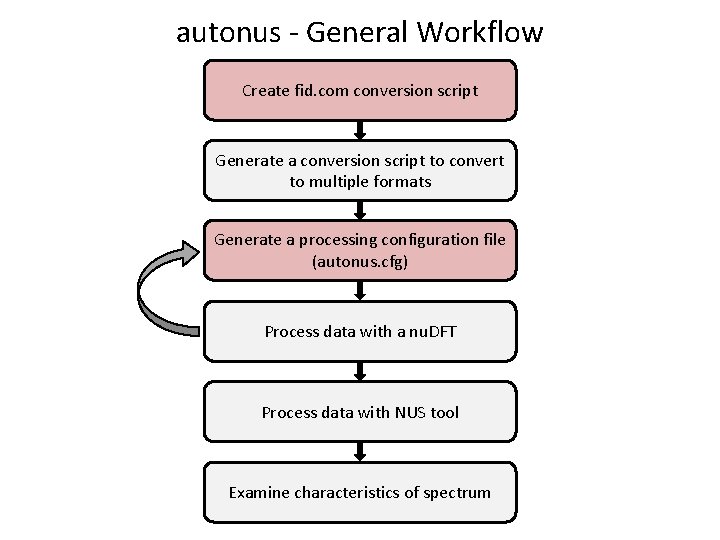
autonus - General Workflow Create fid. com conversion script Generate a conversion script to convert to multiple formats Generate a processing configuration file (autonus. cfg) Process data with a nu. DFT Process data with NUS tool Examine characteristics of spectrum

nu. DFT File conversion (fid. com) “cd HNCACB_nus” “bruker” • Press “Read Parameters”; Change output template from ”. / fid/” to “. /data/” • Press “Save Script” • Press “Quit” “more fid. com”

Generate processing configuration file (autonus. cfg) For the workshop the autonus. cfg files have been pre-created for speed. The file format is ugly and cumbersome • Need to decide if autonus will be a separate GUI based program or whether to wrap it into NMRFx-Processor “more autonus. cfg” • The text file contains some basic input questions and selections for processing along the acquisition dimension, t 1, t 2, as well as sections for the NUS tools. • Initially one can guess as to some of the values, but the workflow is to process data with a nu. DFT to determine values such as phases, sign alterations, apodization functions, etc. This will likely need to be done in an iterative process. • For the workshop the values should be set appropriately for the HNCACB spectrum without any editing necessary • Restore fid. com and autonus. cfg to defaults for Scripps workshop if necessary. • “cp fid. com. scripps fid. com” • “cp autonus. cfg. scripps autonus. cfg”

Create conversion scripts Before any data processing can be performed the data must be converted to the correct format. • All the NUS tools utilize the nmr. Pipe format except rnmrtk, maxent, and loner. • The autonus tool will attempt to create a single conversion script that will read in Varian or Bruker data and convert to all the formats. • Note that while a bit slow, the step only needs to be performed once. “autonus convert” “more convert. com” • All the conversion scripts concatenated. Individual scripts are also created. “. /convert. com” • This will run all the conversion scripts • All data saved in current directory under data nmrpipe & rnmrtk formats are fully expanded with zeros in place of any missing FIDs. • Allows a nu. DFT of the spectrum

nu. DFT with nmrpipe “cd HNCACB_nus” “autonus dft-nmrpipe” “ls -ltr” “more dft-nmrpipe. com” Script broken into 6 sections 1. Acquisition dimension for expanded data (x saved back to x [xyz]) • Intermediate files will be used by: • dft-nmrpipe, nmrpipe-ist, clean, & scrub 2. Acquisition dimension for expanded data for nestanmr (x saved back to x [xyz]) • Intermediate files will be used by: • nestanmr 3. Acquisition dimension for expanded data (x saved to z [yzx]) • Intermediate files will be used by: • smile 4. Acquisition dimension for compressed data (x saved to z [yzx]) • Intermediate files will be used by: • Hmsist & camera 5. Process t 1 dimension for nu. DFT (y saved back to y) 6. Process t 2 dimension for nu. DFT (z saved to y)

nu. DFT (nmrpipe) “. /dft-nmrpipe. com” “. /ls -ltr” • nu. DFT data saved in dft-nmrpipe directory. • Intermediate t 3 processed data saved in dft-nmrpipe, xyz and yzx folders for later use “nmr. Draw -in dft-nmrpipe/dft%03 d. ft 3 “ Move through various planes and examine the large amount of “noise”. The noise is both the empirical noise and “sampling noise” due to the convolution of the point-spreadfunction with every signal in the spectrum The NUS processing tools will attempt to remove the “sampling noise” from the final spectrum.

Rowland NMR Toolkit (RNMRTK) Overview • Originally developed as a platform for developing new NMR data processing methods • Now widely used as a general processing platform • Provides • Rich set of apodization (window) functions • DFT processing • General processing tools (phasing, reversing, truncating, etc. ) • Robust, efficient Linear Prediction (LP) extrapolation • Max. Ent reconstruction (deconvolution, uniform or nonuniform sampling) • universal data reader • exports to nmr. Pipe, XEASY, Felix • Generates synthetic data, noise (testing/error analysis) • Based on a program “VNMR” developed by Jeff Hoch in the late 1970’s • Redesigned in 1985 and then in the early 1990 s • Free for Academic use and included in NMRbox • l 1 -norm real (LONER) has recently been added

RNMRTK – Suite of Programs section – Manages the shared memory sections rnmrtk – The main processing program in the Toolkit Loading, apodization, DFT, phasing, saving, etc. flip – Forward linear prediction msa / msa 2 d / msa 3 d – Maximum entropy reconstruction in one, two, and three dimensions inject – Add synthetic peaks to time domain data select / zsample – Expands or compresses NUS data seepln / contour – graphical display for 1 D and 2 D data Many others …

RNMRTK – Shared memory With a suite of programs for performing tasks – How do you get data output from one command to the next command without needing to write out intermediate files? • Pipes (nmr. Pipe strategy) • Shared memory (RNMRTK strategy) • A shared memory section is a chunk of persistent memory where NMR data can be stored and used by different programs. section • RNMRTK program to manage (create / delete) shared memory sections • Let’s create a shared memory section “section -c 1024 128” • The size of the shared memory section is the product of the arguments multiplied by 4 (32 bit data) plus 512 extra bytes for a header. • Shared memory sections must be large enough to store all the data. “section -d” • Deletes the shared memory section The maximum size of shared memory sections is defined by kernel values shmmax and shmall. NMRbox takes care of these advanced settings.

RNMRTK – Shared memory Share memory example Assume a data set size of t 3=1024 complex points, t 1=80 complex points, and t 2=48 complex points. A shared memory section, just large enough, to load the dataset would be: section -c 1024 2 80 2 48 2 (note the 2’s to handle the complex data) However, lets assume we zerofill to the next Fourier number in all dimensions during processing, then we would need “section -c 2048 2 128 2 64 2”

RNMRTK – rnmrtk • Command for performing many of the traditional processing techniques • Can be run: • one command at a time • in interactive mode • scripted • Row (column) oriented. • Many commands need to have the dimension set.

RNMRTK – Issuing commands • Arguments are entered as different classes: • floats (defined as a number with a decimal) • setpar SF 1 599. 8763 • integers (defined as a number without a decimal) • zerofill 2048 • string (defined as a character string without a period) • sinebell square 70. 0 • filename (defined as a character string with a decimal point) • loadvnmr. /fid • load hsqc. sec • The order of arguments entered is important, but only within a given class of argument • sstdc 16 20 156. 25 COS • sstdc 156. 25 16 COS 20 • rnmrtk is case insensitive (except for filenames) DIM command sets the dimension which is necessary for many commands rnmrtk << EOF LOAD test. sec LOAD command does not need dimension set DIM t 1 FFT command must have dimension set FFT EOF

RNMRTK – Issuing commands From the command line: bash% rnmrtk loadvnmr. /fid bash% rnmrtk setpar SF 1 599. 8763 bash% * DIM command not persistent In interactive mode bash% rnmrtk% loadvnmr. /fid rnmrtk% DIM t 2 rnmrtk% FFT rnmrtk% exit bash% As a script bash% processing_script. com bash% Typical script #! bin/bash section –c 1024 256 rnmrtk << EOF loadvnmr. /fid setpar PPM 2 4. 772 dim t 2 zerofill 1024 fft 0. 5 phase -152. 0 0. 0 realpart EOF rnmrtk << EOF dim t 1 zerofill 128 fft 0. 5 realpart save test. sec EOF Here doc Cannot have spaces at the beginning of lines

RNMRTK – Load data • Varian • rnmrtk loadvnmr. /fid • Bruker • Need to create a parameter file called ser. par • Example: Dom T 1 T 2 T 3 Order of dimensions in memory (after loading) N 64 C 140 C 512 C Note spacing between value and C sw 2583. 0 4498. 0 8992. 8 sf 75. 972 749. 666 ppm 118. 5 7. 706 4. 706 Quad STATES # of padding bytes at beginning of fid Format Little-endian Int-32 0 0 0 # of padding bytes at end of fid # of header bytes Layout t 2: 280 t 1: 128 t 3: 1024 Endian is defined by the computer architecture Layout defines order of dimensions in input file NOTE: spectrum-translator being developed to handled data conversions so little need to manually create load script anymore

RNMRTK – msa 2 d Maximum Entropy Reconstruction in 2 dimensions Reads parameters from a file and dimensions are specified on invocation line • Ex: msa 2 d t 1 t 2 msa 2 d. param Constant Aim parameter file Max loops, convergence may be quicker DEBUG 1 NLOOPS 400 DEF 0. 1 AIM 2. 5 NUS SCALEFIRST 0. 5 SCHED. /sample_schedule NUSE 256 128 NOUT 512 256 Uniform PHASE 0. 0 90. 0 Data will be LW 0. 0 extrapolated JVALUE 0. 0 Constant LAMBDA parameter file DEBUG 1 NLOOPS 400 DEF 0. 1 LAMBDA 1. 0 SCALEFIRST 0. 5 SCHED. /sample_schedule NUSE 256 128 NOUT 512 256 PHASE 0. 0 90. 0 LW 0. 0 JVALUE 0. 0 For 3 D NUS datasets the acquisition dimension is processed with conventional FT and then the indirect dimensions are processed with msa 2 d

RNMRTK – msa 2 d Monitoring convergence Very important! • Ensure that the value of test is less than 0. 1 unless fewer than NLOOPS required • Without deconvolution convergence typically occurs is around 40 iterations if proper def, aim, lambda values were use. • High numbers of loops suggest msa 2 d is modeling noise • With deconvolution the number of loops may be significantly higher.

nu. DFT (rnmrtk) “cd HNCACB_nus” “autonus dft-rnmrtk” “more dft-rnmrtk. com” • Script processes data in all three dimensions with nu. DFT • Two intermediate files are generated • noisecalc. sec • Used by noisecalc to estimate values for DEF and AIM for Max. Ent reconstruction • f 3_proc. sec • Used by Max. Ent and LONER as starting point for reconstruction in the indirect dimensions “nmr. Draw -in rnmrtk/dft-rnmrtk_f 3 f 2. ft 3” • If everything worked the data should look virtually identical to the nmr. Pipe nu. DFT earlier • Intermediate files ready for automatic Max. Ent reconstruction

Auto Max. Ent Reconstruction The maximum entropy algorithm in RNMRTK has three adjustable parameters; def, aim, and lambda. Data can be processed with constant aim mode or constant lambda mode. Workflow for 3 D processing with msa 2 d Choose Def and Aim (Based on empirical noise) Process spectrum in Constant Aim mode Look up the converged Lambda value Process the whole dataset in Constant Lambda mode Choosing def and aim • aim should be a small integer multiple of the rms noise in the data • compute the rms for a blank region of a 1 D (typically use fid with weakest signal) • def should be a value smaller than that of the smallest expected peak • too small and the baseline noise distribution becomes “spikey” • too large and the noise level will be too high • noisecalc – A tool developed by Mehdi Mobli to compute reasonable values of def and aim given a blank region of a 1 D spectrum • autonus – A tool that will perform all these steps in an automated fashion

Auto Max. Ent Reconstruction “cd HNCACB_nus” “autonus maxent” • Script performs the Workflow steps of the previous slide automatically • Reads in t 3 processed data • Analyzes spectrum for noise and estimates DEF and AIM • Processes whole spectrum in constant AIM mode • Examines the log file and averages converged Lambda values • Processes whole spectrum in constant Lambda mode • Saves data and outputs some information • Maximum number of loops • Values between 20 and 80 are typical • Reports DEF, AIM, and LAMBDA “ls -ltr” • maxent. com (Processing script) • msa 2 d_param (Max. Ent paramter file) • msa 2 d. txt (Max. Ent reconstruction log)

Auto Max. Ent Reconstruction “more msa 2 d_test_param” “more msa 2 d_param” DEF determined from noisecalc multiplied by “def multiplication factor” LAMBDA determined from trial run in constant AIM mode Final data size extrapolated to 256 x 256 To re-run with different parameters, if desired, simply edit the msa 2 d_param file and execute “. /maxent. com” “nmr. Draw -in maxent/maxent_f 3 f 2. ft 3”

Automatic LONER Reconstruction The l 1 -norm real algorithm in RNMRTK has one adjustable parameters; aim Processing Workflow • Read in t 3 processed data • Analyzes spectrum for noise and estimates AIM • Reconstruct indirect dimensions with LONER • Saves data and outputs some information • Maximum number of loops • Reports AIM “more loner_param” “nmr. Draw –in loner/loner_f 3 f 2. ft 3”
hms. IST, nmrpipe-ist, SMILE, NESTA-NMR, CAMERA, NMRFx-IST, & NMRFx-NESTA Workflows Workflow Convert time domain data Process the acquisition dimension with NMRPipe Reconstruct missing time domain data Process two indirect dimensions with NMRPipe

NMRFx-Processor (nmrfxp) NMRFx-Processor is a relatively new NMR Processing package from Bruce Johnson at CUNY. Fully GUI based program Lots of intelligence built-in Handles Bruker, Varian, and other data Processing scripts are built in the GUI, but the executed script (process. py by default) is a python script which can be saved and run in a stand-alone manner (not yet implemented in NMRbox) Has IST and NESTA algorithms implemented Has the framework to allow calls to external programs from within the python scripts Under active development “cd HNCACB_nus” “nmrfxp”

NMRFx-Processor (nmrfxp) Instructions File --> Open and Draw. Navigate to Scripps-2017/HNCACB_nus and select ser and open From the Dialog box without the spectrum choose Scripts and Auto Generate For ref select ppm at center and H 20 Select the Operations Tab and make sure D 1 dimension is selected Add TD-Solvent TDSS Select ZF and add 1024 for size Select phase and change ph 0 to 90 and select dimag Add Regions --> EXTRACT and set start=160 and end=415 Select D 2, 3 then select ISTMATRIX and check the disabled checkbox Select D 2 Change ZF size to 256 Change FT to turn off negate. Imag Select the dimag checkbox for phase Select D 3 Change ZF size to 256 Change phase to -23. 0 50. 0 and select the dimag checkbox Click the process button at the bottom – performs a nu. DFT After processing select Y and choose 13 C Select Z and Center – then move about the spectrum Go back to D 2, 3 Turn off the disable button & click Process Disable ISTMATRIX Add Sampling --> NESTA & Re. Process

hms. IST “autonus hmsist” “ls -ltr” • hmsist. com (IST script) • phf 2 pipe. com (Script to rearrange data) • run-hmsist. com (Script to run hmsist. com and parallalize calculations) • hmsist-ft 23. com (Script to process t 1/t 2 dimensions after hms. IST_ • hmsist-all. com (Script to run them all) “more hmsist. com” “hms. IST –help” “. /hmisist-all. com” “nmr. Draw –in hmsist/hmsist%03 d. ft 3”

NMRPipe IST “autonus nmrpipe-ist” “more nmrpipe-ist. com” Notes • ist 3 D. com processes the acquisition dimension with FT, performs an IST calculation to fill in missing points in t 1/t 2, and then processes the t 1/t 2 planes with FT all internally. • Arguments can be passed to change default values for phases, FT arguments, apodization, etc. • In the latest version -ist. Max. Res can be set to Auto • Also in latest version data can be extended beyond the last point collected – slows calculation • Likely the slowest of all the NUS techniques – especially when the highest increment values are large. “. /nmrpipe-ist. com” “nmr. Draw -in nmrpipe-ist/ist%03 d. ft 3”

NESTA-NMR “cd hncacb_nus” “autonus nestanmr” “ls -ltr” “more nestanmr. com” “. /nestanmr. com” “nmr. Draw -in nestanmr/nestanmr%03 d. ft 3”

SMILE, CAMERA, SCRUB “cd HNCACB_nus” “autonus smile” “ls -ltr” “more smile. com” “. /smile. com” “nmr. Draw –in smile/smile%03 d. ft 3” Repeat for camera and scrub

Additional Data Sets • • • 15 N-NOESYHSQC. fid – A Varian data set collected uniformly and sub-sampled into a NUS with 26% coverage. w Process with autonus to see how the NUS tools work with a high dynamic range spectrum • Note: Don’t use NMRPipe-IST

Uniform HNCO (ni=128, ni 2=128) with Max. Ent “cd hnco-large. fid” • The HNCO was run with t 1 and t 2 collected out to 128 points. • The process. com script processes the data with a sample schedule entered as an argument with RNMRTK’s maximum entropy reconstruction and performs a nu. DFT. Any point not in the sample schedule will be ignored during processing. • The only criteria is that the sample schedule not exceed 128 in either dimension and that the schedule is 1 indexed. • The output files are: • • Base. Output. Name_nudft. ft 3 (3 D) & Base. Output. Name_nudft_proj. ft 2 (Projection) Base. Output. Name_msa 2 d. ft 3 (3 D) & Base. Output. Name_msa 2 d_proj. ft 2 (Projection) “more rnmrtk. com” • Script to process HNCO with FT in all dimensions with RNMRTK “more nmrpipe. com” • Script to process HNCO with FT in all dimensions with NMRPipe “. /rnmrtk. com” • Create a sample schedule making sure that increment values do not exceed 128 in either of the two dimensions “. /msa 2 d. com Sample. Schedule. scd Base. Output. Name” • After running the data will be saved with the filename Base_output_name. ft 3
- Slides: 39