Genes Expression or Genes and How They Work


































- Slides: 64

Genes Expression or Genes and How They Work: Transcription, Translation, & More Chapter 15

Overview: Central Dogma • • Central Dogma – DNA RNA Protein During polypeptide synthesis, ribosomal RNA (r. RNA) is the site of polypeptide assembly. – Transfer RNA (t. RNA) transports and positions amino acids. – Messenger RNA (m. RNA) directs which amino acids are assembled into polypeptides. 2 2

The Central Dogma DNA Central Dogma Proposed by Francis Crick in 1958 to describe the flow of information in a cell. Information stored in DNA is transferred residue-by-residue to RNA which in turn transfers the information residue-by-residue to protein. RNA Protein The Central Dogma was proposed by Crick to help scientists think about molecular biology. It has undergone numerous revisions in the past 45 years. 3 3

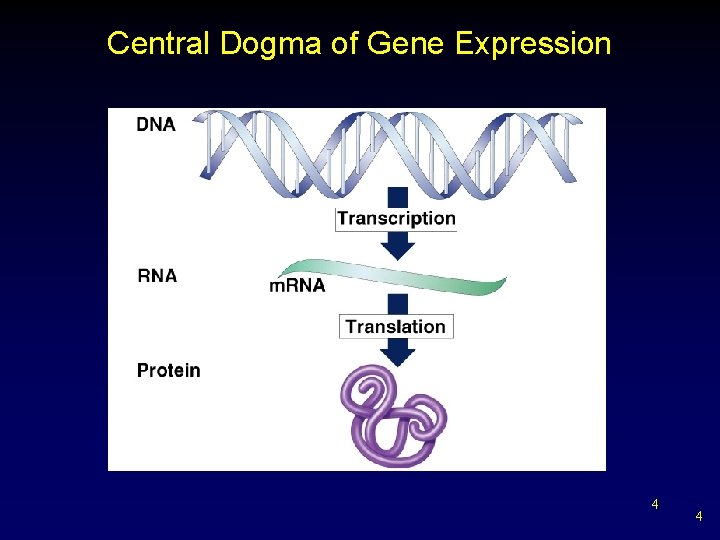
Central Dogma of Gene Expression 4 4

Transcription Overview Def: DNA sequence is transcribed into RNA sequence – initiated when RNA polymerase binds to promoter binding site § moves along DNA strand adds corresponding complementary RNA nucleotide v disengages at stop signal 5 5

Translation Overview • • Def: nucleotide sequence of m. RNA transcript is translated into amino acid sequence in the polypeptide § r. RNA recognizes and binds to start sequence v moves three nucleotides at a time Ø disengages at stop signal Gene expression - collective of transcription and translation 6 6

Genetic Code • • How does the order of nucleotides in a DNA molecule encode the information that specifies the order of amino acids in a polypeptide? The answer came in 1961 through an experiment lead by Crick. 7 7

Genetic Code • • Crick and colleagues reasoned that there must be codons or block of info that coded for an amino acid They hypothesized that it was most likely 3 nucleotides – Why 3? – 2 nucleotides did not have enough combinations (42 is only 16 possible amino acids) 3 – 3 nucleotides (4 is 64 which is enough to cover the roughly 20 known amino acids) 8 8

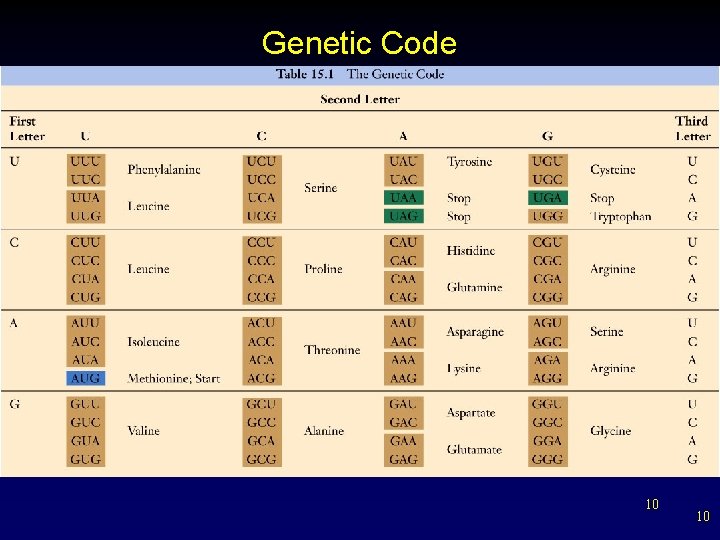
Genetic Code • • Now known Genetic code consists of a series of information blocks called codons. – reading frame (triplet) § each codes for one amino acid § highly redundant 9 9

Genetic Code 10 10

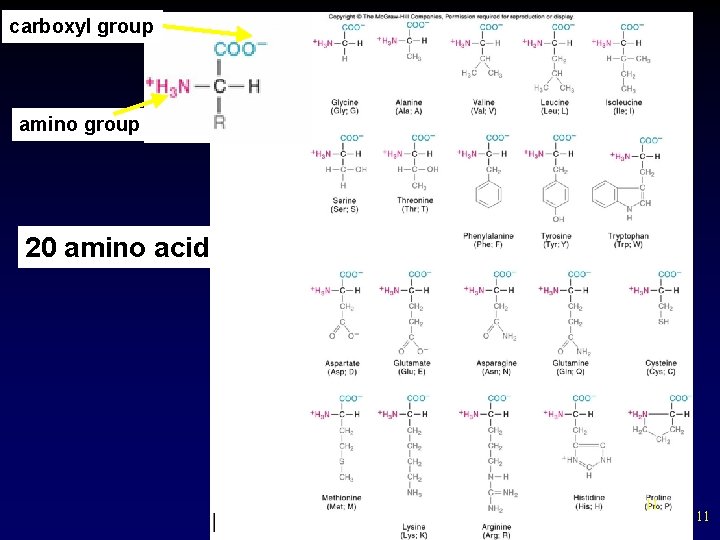
carboxyl group amino group 20 amino acids 11 11

Genetic Code • • Could be punctuated or not – Punctuated code would have a something in the code that separates codons – Non-punctuated code would not In the following example, O is not a base pair but the “punctuation. ” 12 12

Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Delete 1 base 15_06. jpg WHYDIDTHEREDBATEATTHEFATRAT? Hypothesis A : Delete T unpunctuated WHY DID HER EDB ATE ATT HEF ATR AT? (Nonsense) Hypothesis B : punctuated WHYODIDOTHEOREDOBATOEATOTHEOFATORAT? Delete T O O R B E T F R WHY DID HEO EDO ATO HEO AT? (Nonsense) Delete 3 bases Hypothesis A : WHYDIDTHEREDBATEATTHEFATRAT? unpunctuated Delete T, R, and A WHY DID HEE DBT EAT THE FAT RAT? (Sense) (Nonsense) WHYODIDOTHEOREDOBATOEATOTHEOFATORAT? Hypothesis B : Delete T, R, and A punctuated O O E T T WHY DID HEO DOB OEA OTH OFA ORA? (Nonsense) 13 13

Genetic Code • • • Could be punctuated or not – Punctuated code would have a something in the code that separates codons – Non-punctuated code would not In the following example, O is not a base pair but the “punctuation. ” Crick concluded that it is not punctuated as mutations lead to a different sequence of amino acids, not nonsense. 14 14

Genetic Code • Code is practically universal – ex: AGA codes for arginine in bacteria, humans and all other organisms studied – great evidence that all life has a common ancestor – Genes coded in one organism can be transcribed in another § SWEET biotechnology 15 15

Genetic Code • • • Code is practically universal…but not quite In 1979 mammalian mitochondria found to have different “universal code” – In mitochondrial DNA, UGA is not a stop codon as it is in “universal code” – Other codons are different – Chloroplasts and ciliates (protists) have minor differences as well It is thought that the changes to mitochondria and chloroplasts happen after their endosymbiotic existence 16 16

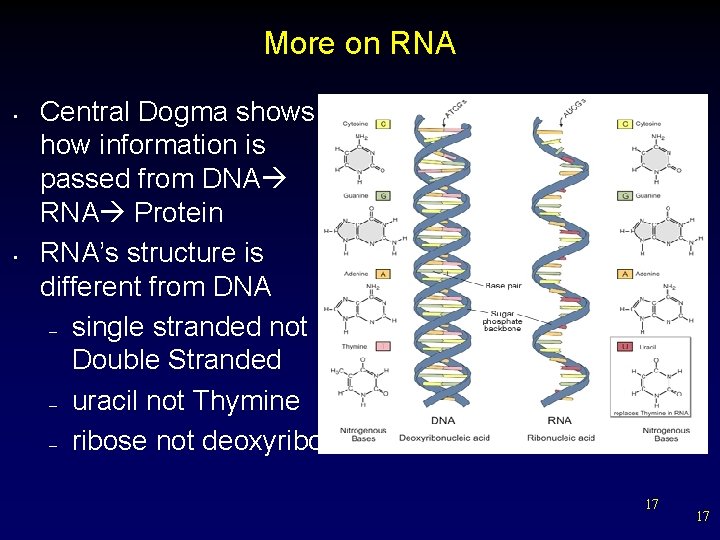
More on RNA • • Central Dogma shows how information is passed from DNA RNA Protein RNA’s structure is different from DNA – single stranded not Double Stranded – uracil not Thymine – ribose not deoxyribose 17 17

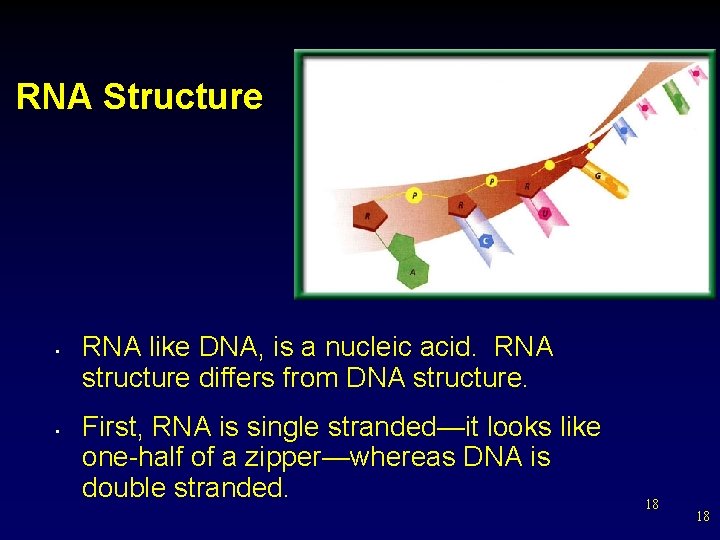
RNA Structure • • RNA like DNA, is a nucleic acid. RNA structure differs from DNA structure. First, RNA is single stranded—it looks like one-half of a zipper—whereas DNA is double stranded. 18 18

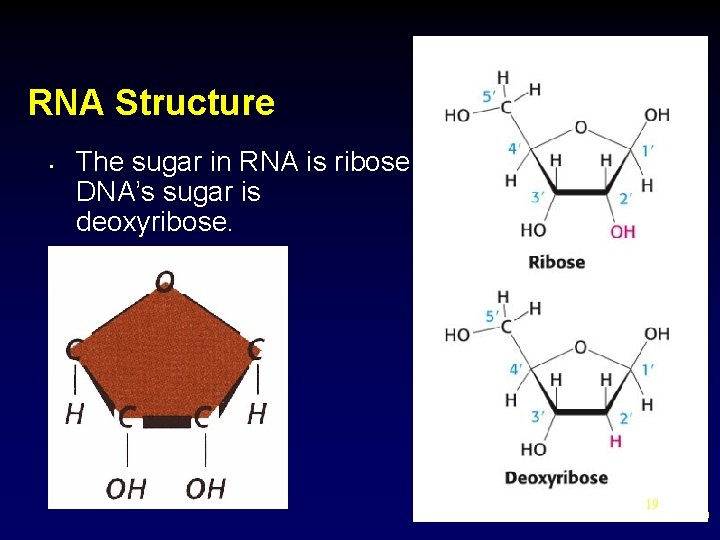
RNA Structure • The sugar in RNA is ribose; Ribose DNA’s sugar is deoxyribose. 19 19

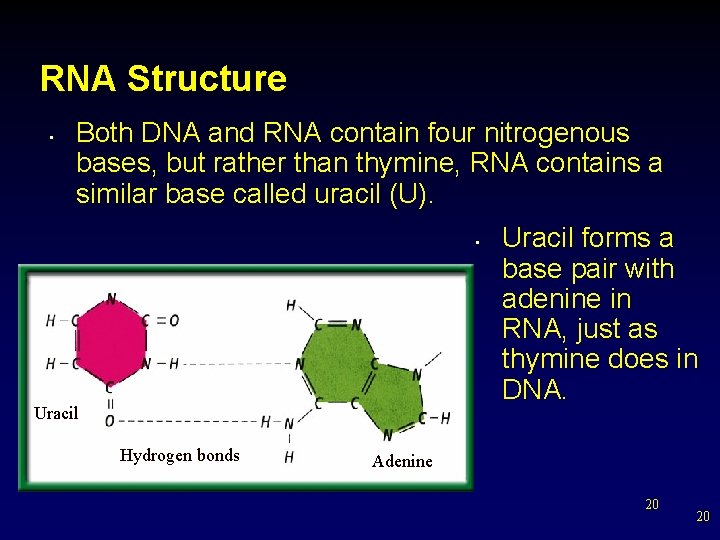
RNA Structure • Both DNA and RNA contain four nitrogenous bases, but rather than thymine, RNA contains a similar base called uracil (U). • Uracil Hydrogen bonds Uracil forms a base pair with adenine in RNA, just as thymine does in DNA. Adenine 20 20

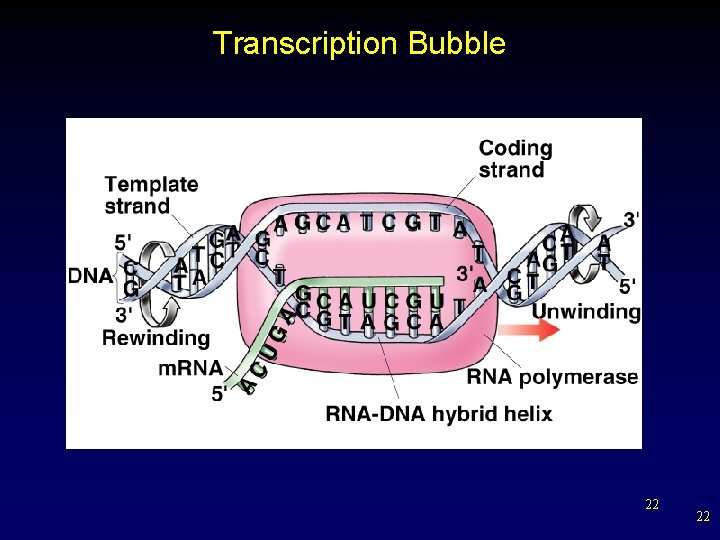
Transcription • RNA polymerase – only one of two DNA strands (template or antisense strand) is transcribed – non-transcribed strand is termed coding strand or sense strand – In both bacteria and eukaryotes, the polymerase adds ribonucleotides to the growing 3’ end of an RNA chain. § synthesis proceeds in 5’ 3’ direction 21 21

Transcription Bubble 22 22

Transcription • • Promoter – Transcription starts at RNA polymerase binding sites called promoters on DNA template strand. Initiation – Other eukaryotic factors bind, assembling a transcription complex. § RNA polymerase begins to unwind DNA helix. 23 23

Transcription • • Elongation – Transcription bubble moves down DNA at constant rate leaving growing RNA strands protruding from the bubble. Termination – Stop sequences at the end of the gene cause phosphodiester bond formation to cease, transcription bubble to dissociate, and RNA polymerase to release DNA. 24 24

25 25

Eukaryotic Transcription • Eukaryotic transcription differs from prokaryotic transcription: – three RNA polymerase enzymes – initiation complex forms at promoter – RNAs are modified after transcription 26 26

Translation: From m. RNA to Protein • • The process of converting the information in a sequence of nitrogenous bases in m. RNA into a sequence of amino acids in protein is known as translation. Translation takes place at the ribosomes in the cytoplasm. 27 27

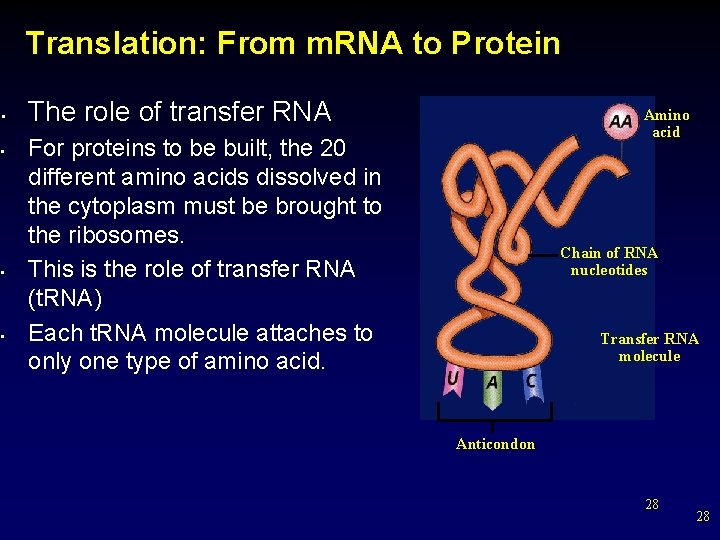
Translation: From m. RNA to Protein • • The role of transfer RNA Amino acid For proteins to be built, the 20 different amino acids dissolved in the cytoplasm must be brought to the ribosomes. This is the role of transfer RNA (t. RNA) Each t. RNA molecule attaches to only one type of amino acid. Chain of RNA nucleotides Transfer RNA molecule Anticondon 28 28

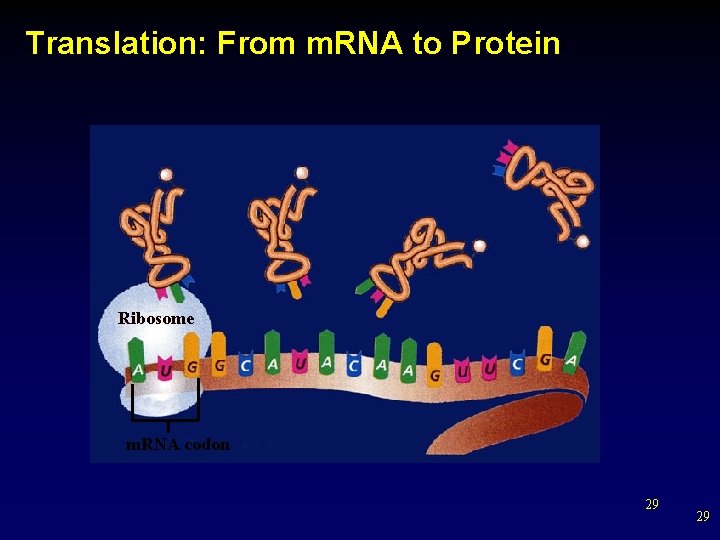
Translation: From m. RNA to Protein Ribosome m. RNA codon 29 29

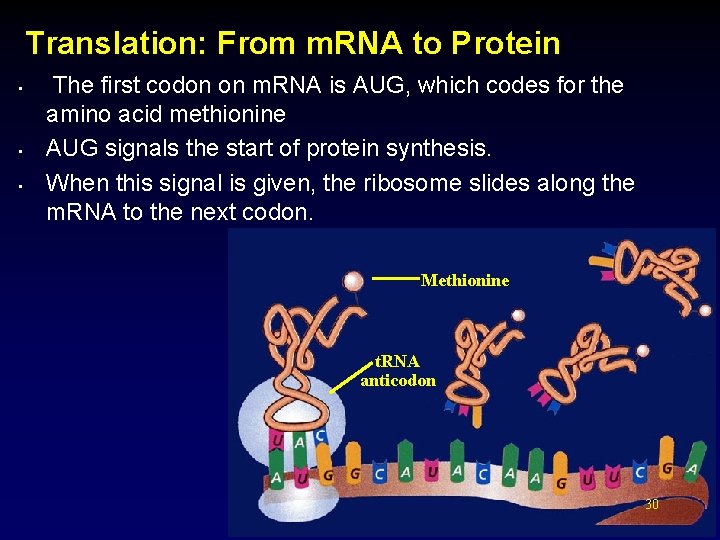
Translation: From m. RNA to Protein • • • The first codon on m. RNA is AUG, which codes for the amino acid methionine AUG signals the start of protein synthesis. When this signal is given, the ribosome slides along the m. RNA to the next codon. Methionine t. RNA anticodon 30 30

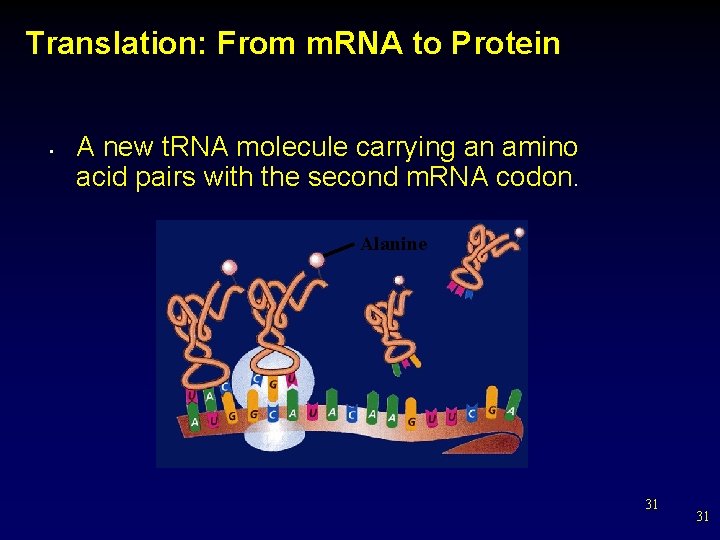
Translation: From m. RNA to Protein • A new t. RNA molecule carrying an amino acid pairs with the second m. RNA codon. Alanine 31 31

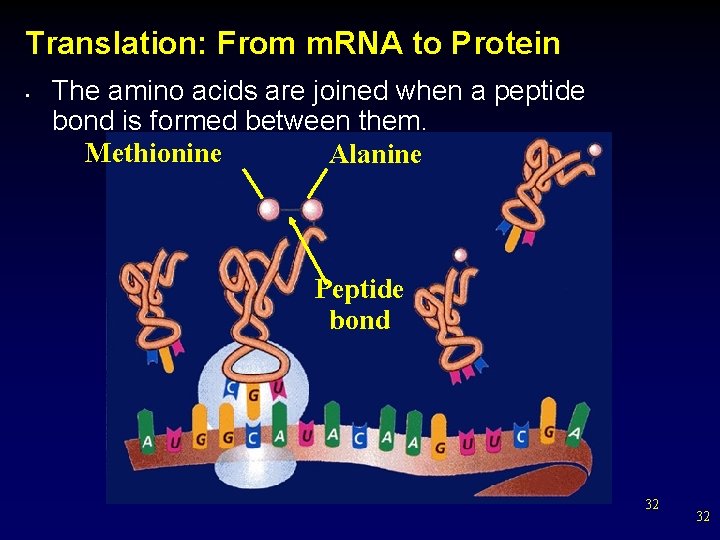
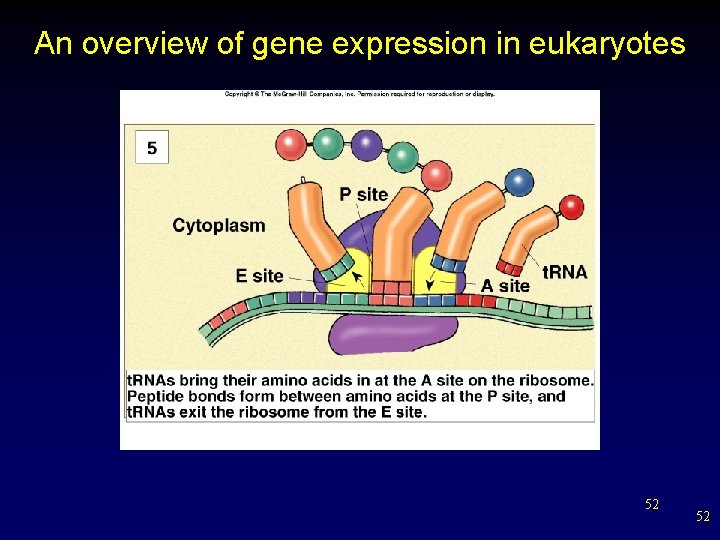
Translation: From m. RNA to Protein • The amino acids are joined when a peptide bond is formed between them. Methionine Alanine Peptide bond 32 32

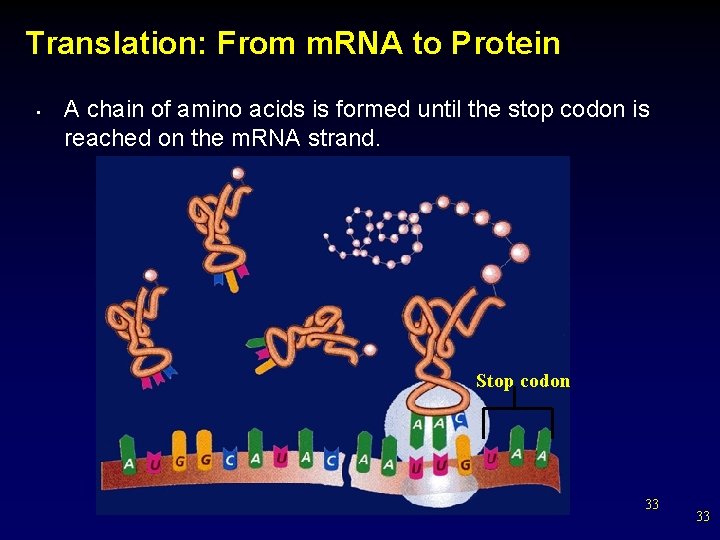
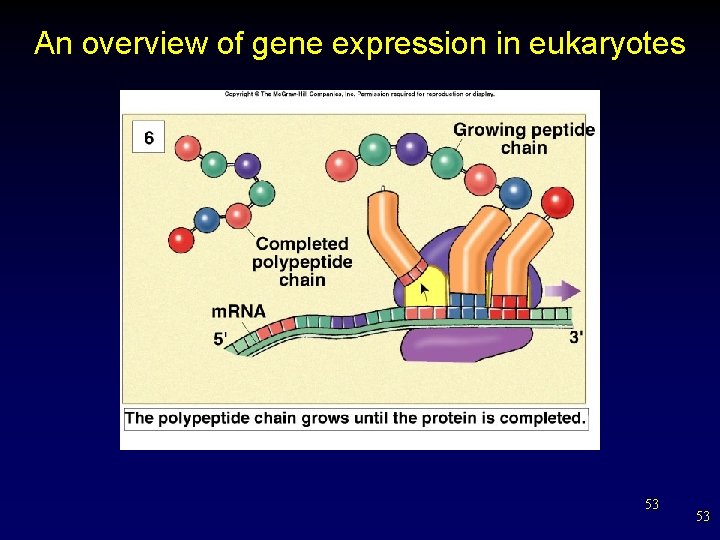
Translation: From m. RNA to Protein • A chain of amino acids is formed until the stop codon is reached on the m. RNA strand. Stop codon 33 33

Translation (in more detail) • Begins when initial portion of m. RNA molecule binds to r. RNA in a ribosome – t. RNA molecule with complimentary anticodon binds to exposed codon on m. RNA § some t. RNA molecules recognize more than one codon 34 34

Translation (in more detail) • Activating enzymes – t. RNA molecules attach to specific amino acids through the action of activating enzymes (aminoacyl-t. RNA syntheases). § must correspond to specific anticodon sequences on a t. RNA molecule as well as particular amino acids 35 35

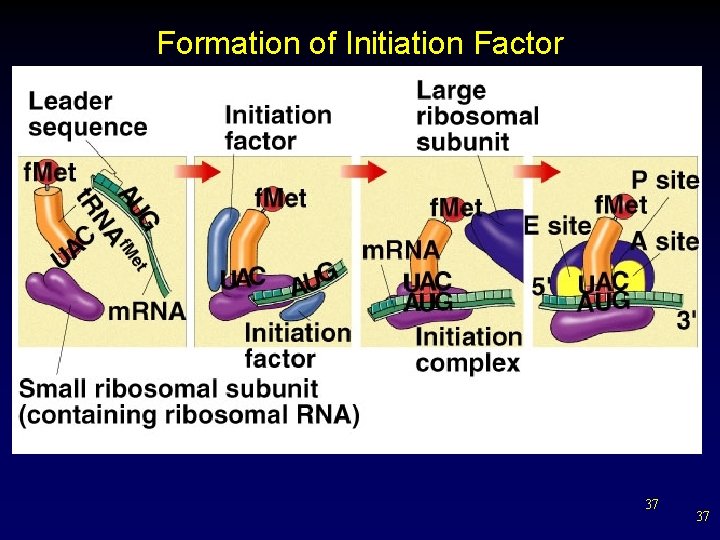
Translation (in more detail) • • Start and stop signals – start signal coded by AUG codon – stop signal coded by one of three nonsense codons: UAA - UAG – UGA – What do you think “nonsense codons” means here? Initiation – Polypeptide synthesis begins with the formation of an initiation complex. § initiation factors 36 36

Formation of Initiation Factor 37 37

Translation (in more detail) • Elongation – After initiation complex forms, large ribosome subunit binds, exposing m. RNA codon adjacent to the initiating codon, positioning it for interaction with another amino acid-bearing t. RNA molecule. 38 38

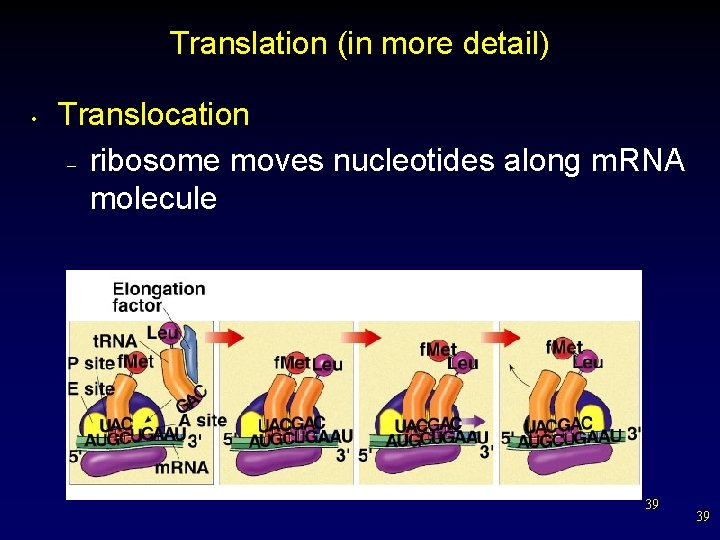
Translation (in more detail) • Translocation – ribosome moves nucleotides along m. RNA molecule 39 39

A bit about the peptide bond formation • • A peptide bond (or amide bond) is a covalent chemical bond formed between two molecules when the carboxyl group of one molecule reacts with the amine group of the other molecule, thereby releasing a molecule of water. This is a dehydration synthesis reaction (also known as a condensation reaction), and usually occurs between amino acids. The resulting C(O)NH bond is called a peptide bond, and the resulting molecule is an amide. The four-atom functional group -C(=O)NH- is called a peptide link 40 40

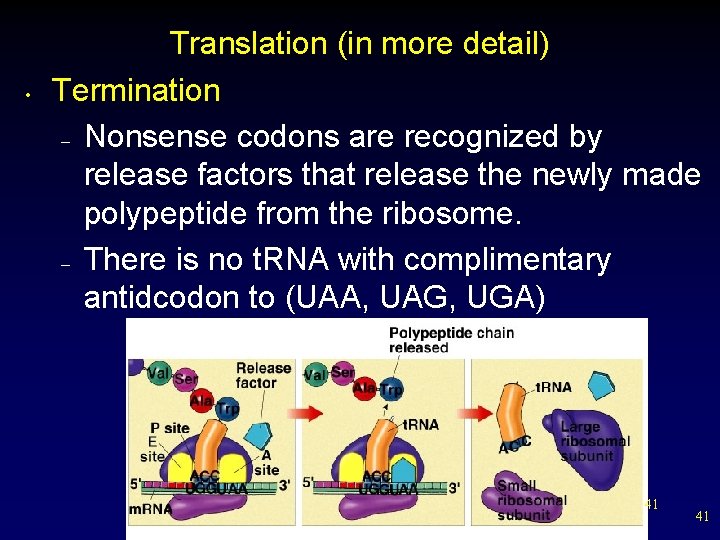
• Translation (in more detail) Termination – Nonsense codons are recognized by release factors that release the newly made polypeptide from the ribosome. – There is no t. RNA with complimentary antidcodon to (UAA, UAG, UGA) 41 41

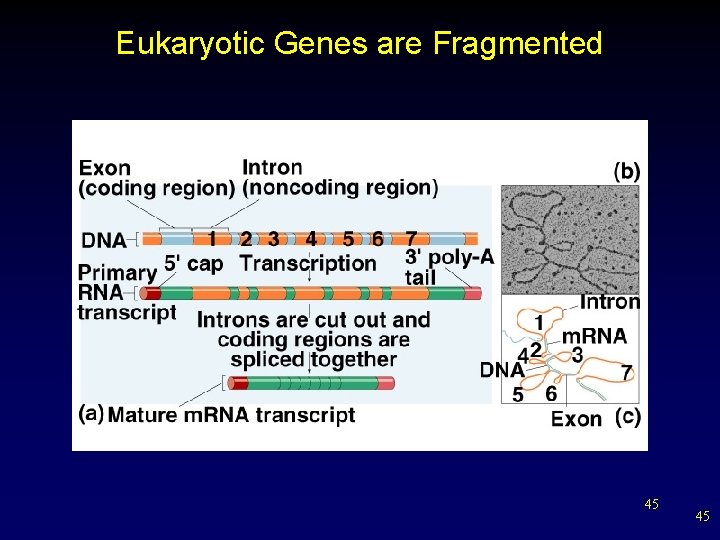
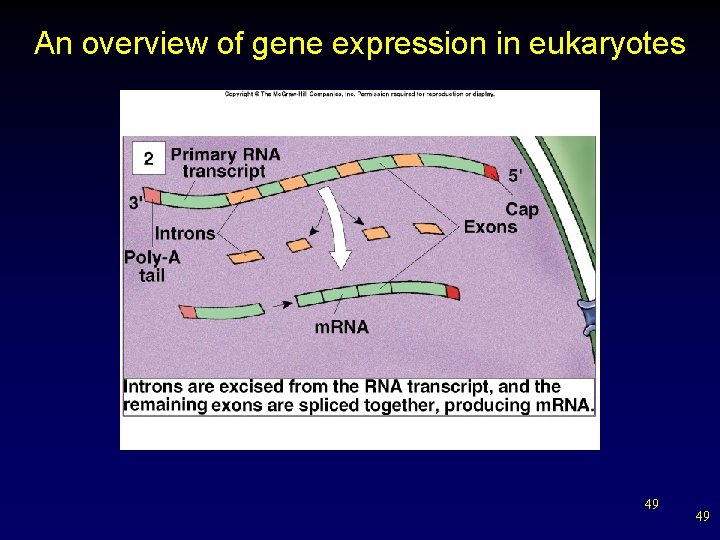
Spliced Gene Transcripts • • DNA sequence specifying a protein is broken into segments (exons) scattered among longer noncoding segments (introns). Initially, primary RNA transcript is produced for the entire gene. – Small nuclear ribonuclearproteins (sn. RNPs) associate with proteins to form spliceosomes. § Lariat forms, excising introns and splicing exons to form mature m. RNA. v alternative splicing 42 42

43 43

RNA Splicing • During RNA processing, intron sequences are cut of primary transcript before it is used in polypeptide synthesis. – remaining sequences are not translated § remaining exon sequences are spliced together to form final processed m. RNA 44 44

Eukaryotic Genes are Fragmented 45 45

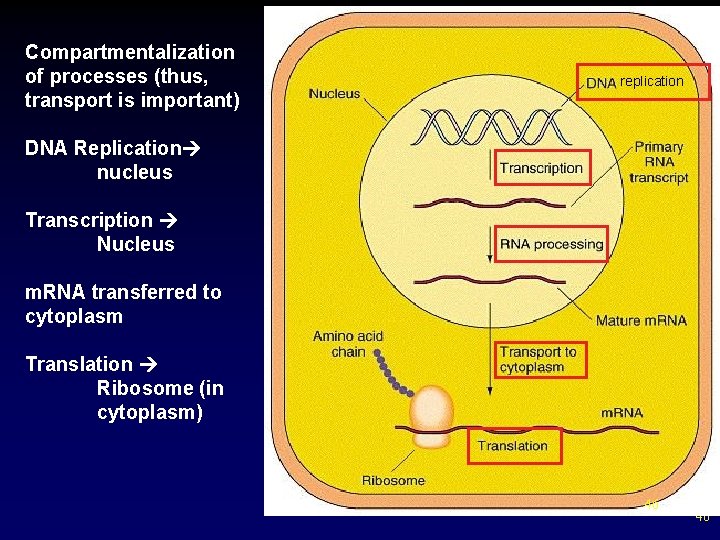
Compartmentalization of processes (thus, transport is important) replication DNA Replication nucleus Transcription Nucleus m. RNA transferred to cytoplasm Translation Ribosome (in cytoplasm) 46 46

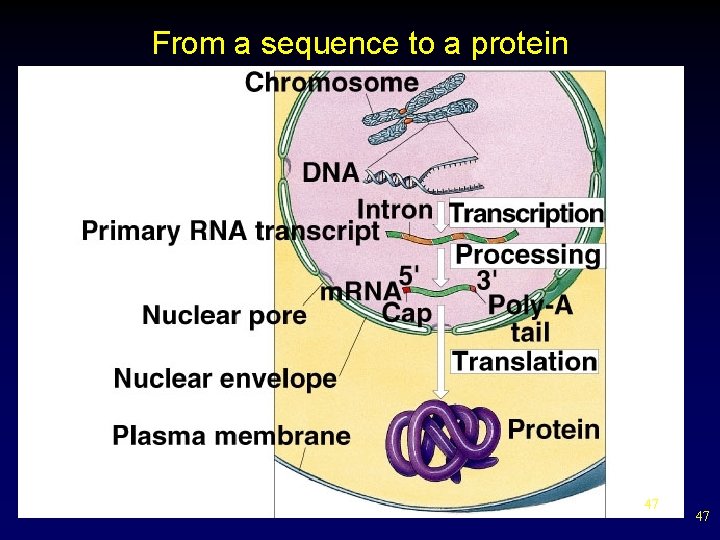
From a sequence to a protein 47 47

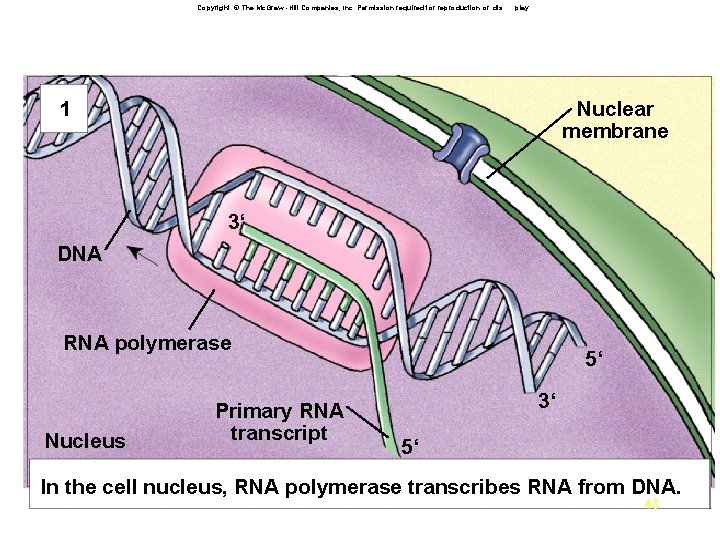
Copyright © The Mc. Graw -Hill Companies, Inc. Permission required for reproduction or dis play. An overview of gene expression in eukaryotes 1 Nuclear membrane 3‘ DNA RNA polymerase Nucleus Primary RNA transcript 5‘ 3‘ 5‘ In the cell nucleus, RNA polymerase transcribes RNA from DNA. 48 48

An overview of gene expression in eukaryotes 49 49

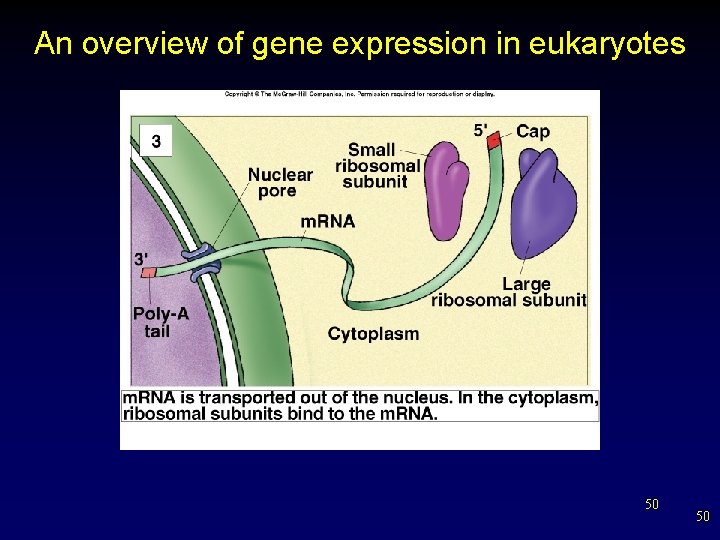
An overview of gene expression in eukaryotes 50 50

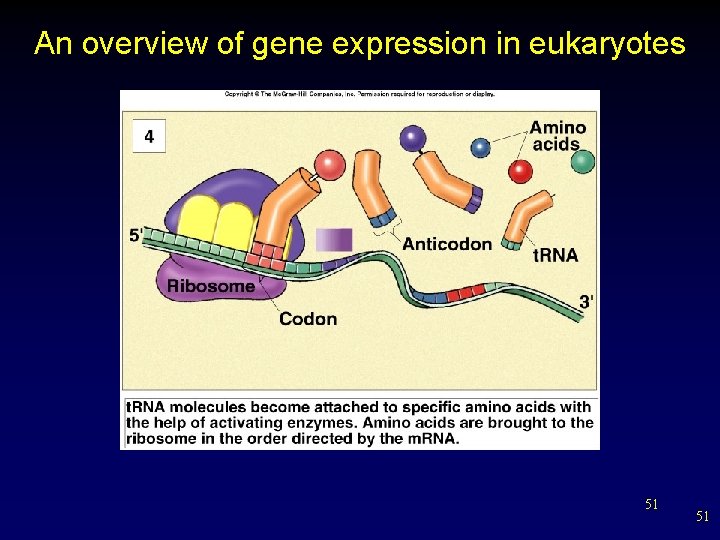
An overview of gene expression in eukaryotes 51 51

An overview of gene expression in eukaryotes 52 52

An overview of gene expression in eukaryotes 53 53

Differences Between Prokaryotic and Eukaryotic Gene Expression • • Most eukaryotic genes possess introns (prokaryotic genes do not. ) Individual bacterial m. RNA molecules often contain transcripts of several genes. Eukaryotic m. RNA molecules must be completely formed and must pass across the nuclear membrane before translation. In prokaryotes, translation begins at the AUG codon preceded by a special nucleotide sequence. 54 54

Differences Between Prokaryotic and Eukaryotic Gene Expression • • Eukaryotic m. RNA molecules have introns cut out and exons joined together before translation. Eukaryotic ribosomes are larger than prokaryotic ribosomes. 55 55

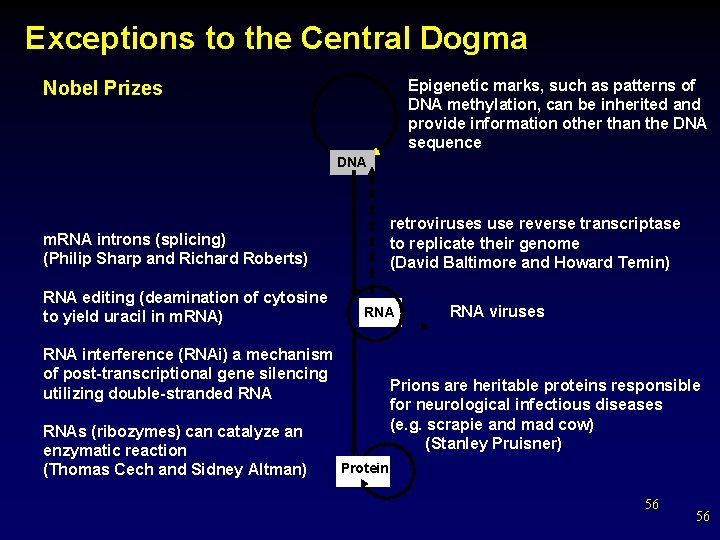
Exceptions to the Central Dogma Epigenetic marks, such as patterns of DNA methylation, can be inherited and provide information other than the DNA sequence Nobel Prizes DNA retroviruses use reverse transcriptase to replicate their genome (David Baltimore and Howard Temin) m. RNA introns (splicing) (Philip Sharp and Richard Roberts) RNA editing (deamination of cytosine to yield uracil in m. RNA) RNA interference (RNAi) a mechanism of post-transcriptional gene silencing utilizing double-stranded RNAs (ribozymes) can catalyze an enzymatic reaction (Thomas Cech and Sidney Altman) RNA viruses Prions are heritable proteins responsible for neurological infectious diseases (e. g. scrapie and mad cow) (Stanley Pruisner) Protein 56 56

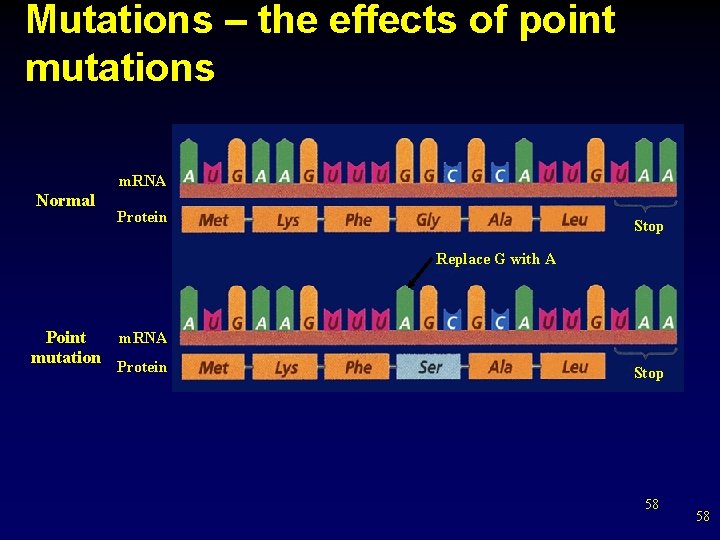
How do mutation effect proteins • • Any change in DNA sequence is called a mutation. Mutations can be caused by errors in replication, transcription, cell division, or by external agents The effects of point mutations • A point mutation is a change in a single base pair in DNA • A change in a single nitrogenous base can change the entire structure of a protein because a change in a single amino acid can affect the shape of the protein 57 57

Mutations – the effects of point mutations m. RNA Normal Protein Stop Replace G with A m. RNA Point mutation Protein Stop 58 58

Frameshift mutations • A mutation in which a single base is added or deleted from DNA is called a frameshift mutation because it shifts the reading of codons by one base. • Structural changes in chromosomes are called chromosomal mutations. 59 59

Causes of Mutations • • • Any agent that can cause a change in DNA is called a mutagen. Mutagens include radiation, chemicals, and even high temperatures. Forms of radiation, such as X rays, cosmic rays, ultraviolet light, and nuclear radiation, are dangerous mutagens because the energy they contain can damage or break apart DNA. 60 60

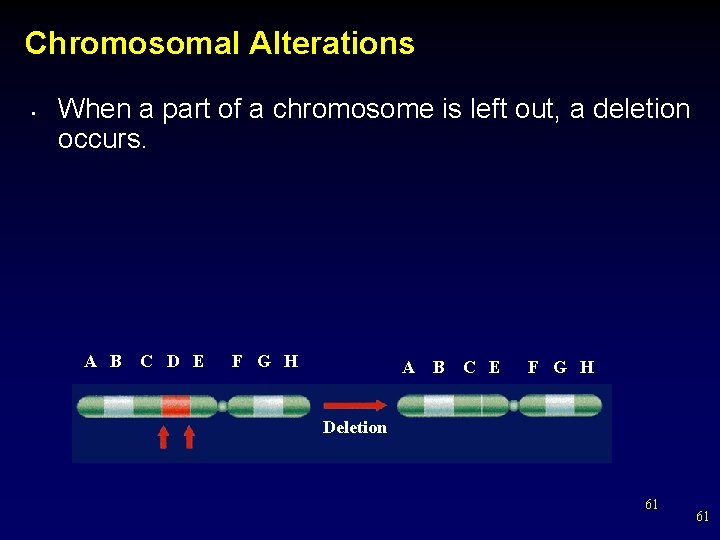
Chromosomal Alterations • When a part of a chromosome is left out, a deletion occurs. A B C D E F G H A B C E F G H Deletion 61 61

Chromosomal Alterations • • When part of a chromatid breaks off and attaches to its sister chromatid, an insertion occurs. The result is a duplication of genes on the same chromosome. A B C D E F G H Insertion 62 62

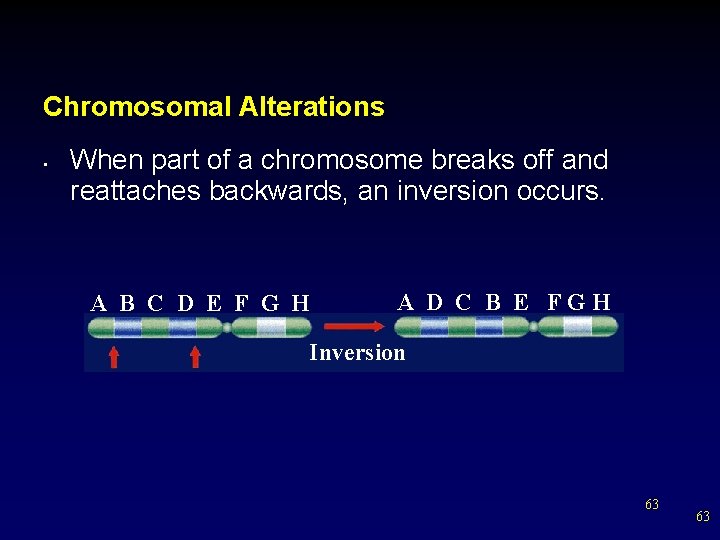
Chromosomal Alterations • When part of a chromosome breaks off and reattaches backwards, an inversion occurs. A B C D E F G H A D C B E FGH Inversion 63 63

Chromosomal Alterations • When part of one chromosome breaks off and is added to a different chromosome, a translocation occurs. AB C D E F GH WX Y Z W X AB C DE F GH Translocation Y Z 64 64