PUCaller Sensitive somatic variant calling using positiveunlabeled learning

PU-Caller: Sensitive somatic variant calling using positive-unlabeled learning Elham Sherafat and Ion Mandoiu Computer Science & Engineering Department University of Connecticut

Outline • • Motivation Positive-unlabeled learning Results Ongoing work
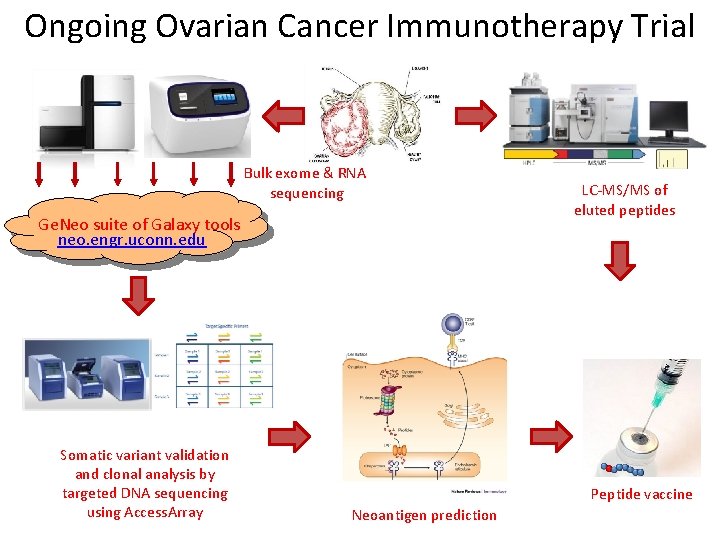
Ongoing Ovarian Cancer Immunotherapy Trial Bulk exome & RNA sequencing Ge. Neo suite of Galaxy tools neo. engr. uconn. edu Somatic variant validation and clonal analysis by targeted DNA sequencing using Access. Array LC-MS/MS of eluted peptides Peptide vaccine Neoantigen prediction
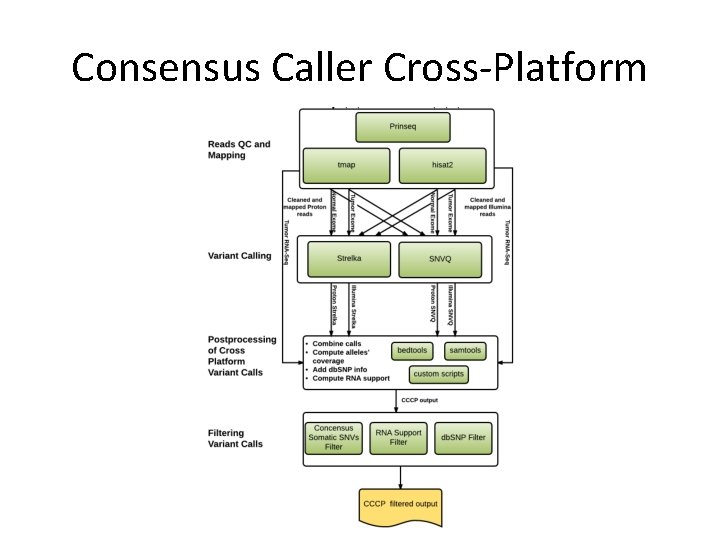
Consensus Caller Cross-Platform
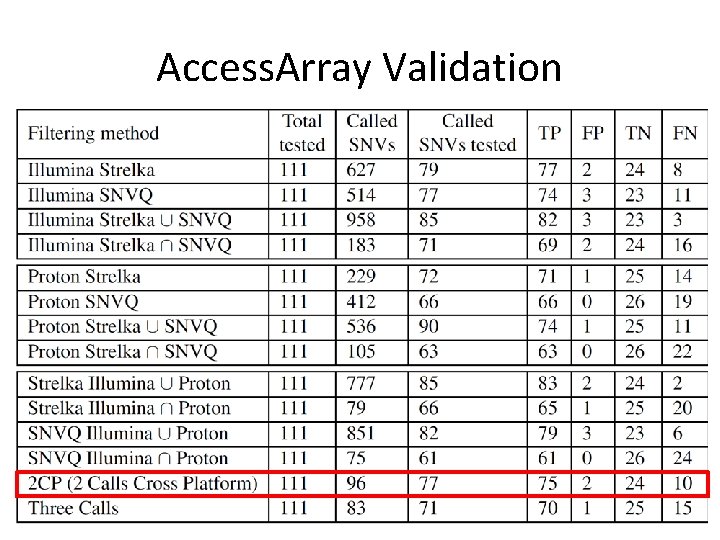
Access. Array Validation
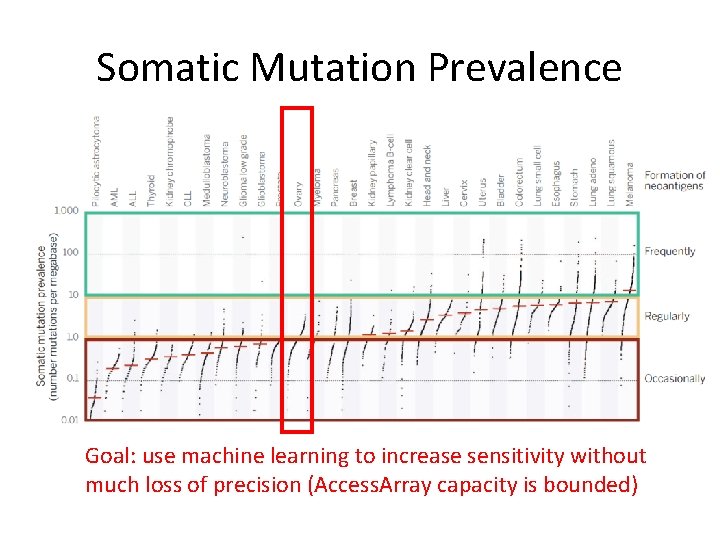
Somatic Mutation Prevalence Goal: use machine learning to increase sensitivity without much loss of precision (Access. Array capacity is bounded)

Previous Work • Supervised ML approaches • Need large amounts of training data • Assume matched distributions between training and test data https: //roywright. me/2017/11/16/positive-unlabeled-learning/

Outline • • Motivation Positive-unlabeled learning Results Ongoing work

PU Learning • Input: • 10 s-100 s of high confidence SNVs from CCCP/2 CP (“positives”) • 104 -105 of SNV candidates that fail 2 CP filter (“unlabeled”) • Two-step approach: • Infer “reliable negatives’’ from unlabeled data • Train classifier using positives and reliable negatives, then classify all points • Robust to patient-to-patient variability https: //roywright. me/2017/11/16/positive-unlabeled-learning/
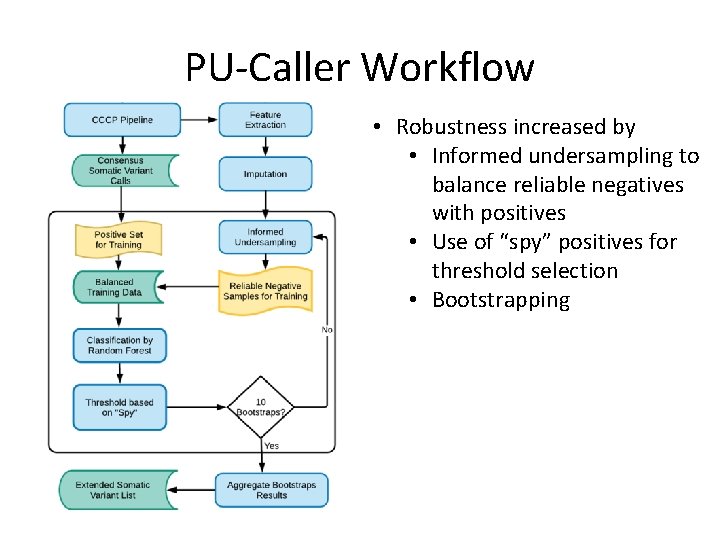
PU-Caller Workflow • Robustness increased by • Informed undersampling to balance reliable negatives with positives • Use of “spy” positives for threshold selection • Bootstrapping

Outline • • Motivation Positive-unlabeled learning Results Ongoing work
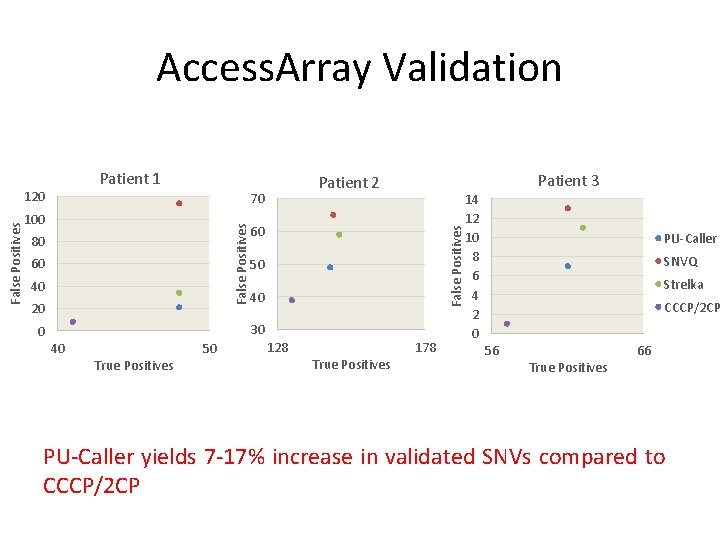
Access. Array Validation Patient 1 70 80 60 40 20 60 False Positives 100 False Positives 120 Patient 3 Patient 2 50 40 30 0 40 True Positives 50 128 True Positives 178 14 12 10 8 6 4 2 0 PU-Caller SNVQ Strelka CCCP/2 CP 56 True Positives 66 PU-Caller yields 7 -17% increase in validated SNVs compared to CCCP/2 CP
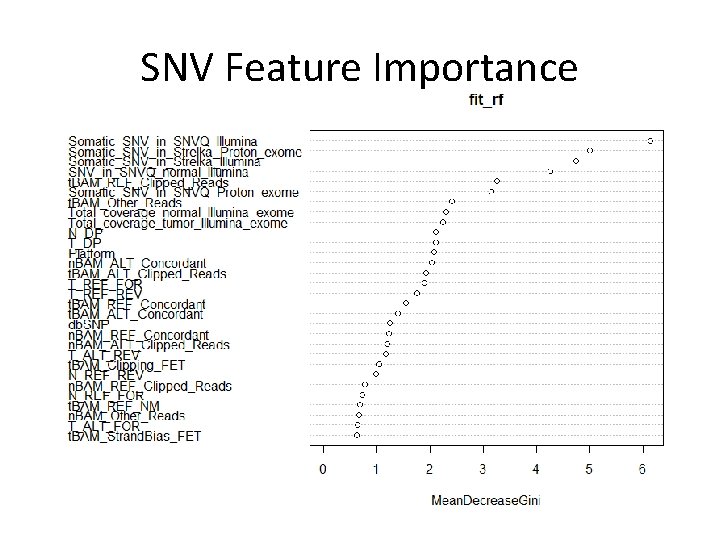
SNV Feature Importance

Outline • • Motivation Positive-unlabeled learning Results Ongoing work

Ongoing Work • PU learning technique is broadly applicable • • Currently using PU learning for improving sensitivity of peptide identification from LC-MS/MS data MS-GF+ database search engine generates 1000 s of confidentifications, but leaves 10, 000 s of spectra unmatched

Acknowledgments Elham Sherafat Jordan Force
- Slides: 16