PeakCalling with MACS 2 Mahony Lab Department of

Peak-Calling with MACS 2 Mahony Lab Department of Biochemistry and Molecular Biology

Outline: • Basic molecular biology. • Review Ch. IP-seq (Single-End) • Overview of MACS 2 Mahony Lab
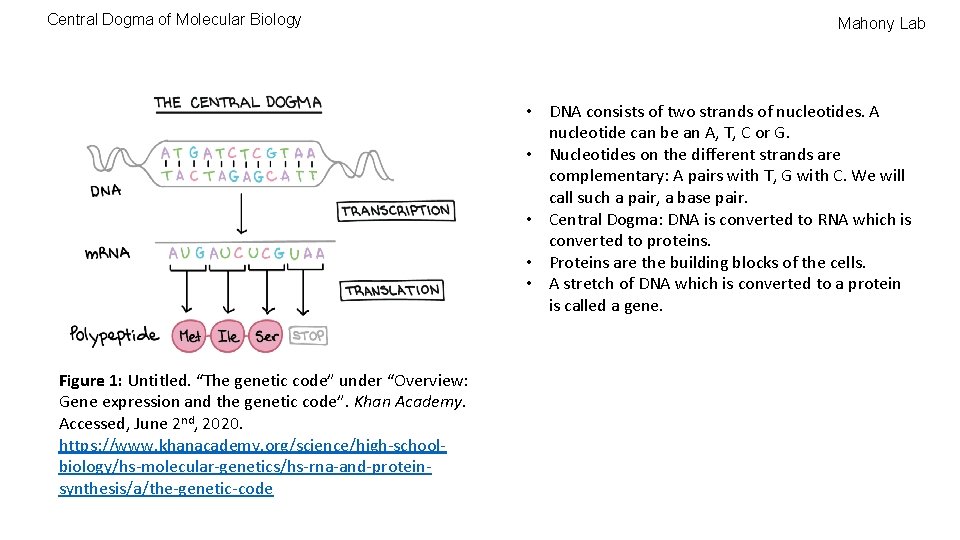
Central Dogma of Molecular Biology Mahony Lab • DNA consists of two strands of nucleotides. A nucleotide can be an A, T, C or G. • Nucleotides on the different strands are complementary: A pairs with T, G with C. We will call such a pair, a base pair. • Central Dogma: DNA is converted to RNA which is converted to proteins. • Proteins are the building blocks of the cells. • A stretch of DNA which is converted to a protein is called a gene. Figure 1: Untitled. “The genetic code” under “Overview: Gene expression and the genetic code”. Khan Academy. Accessed, June 2 nd, 2020. https: //www. khanacademy. org/science/high-schoolbiology/hs-molecular-genetics/hs-rna-and-proteinsynthesis/a/the-genetic-code
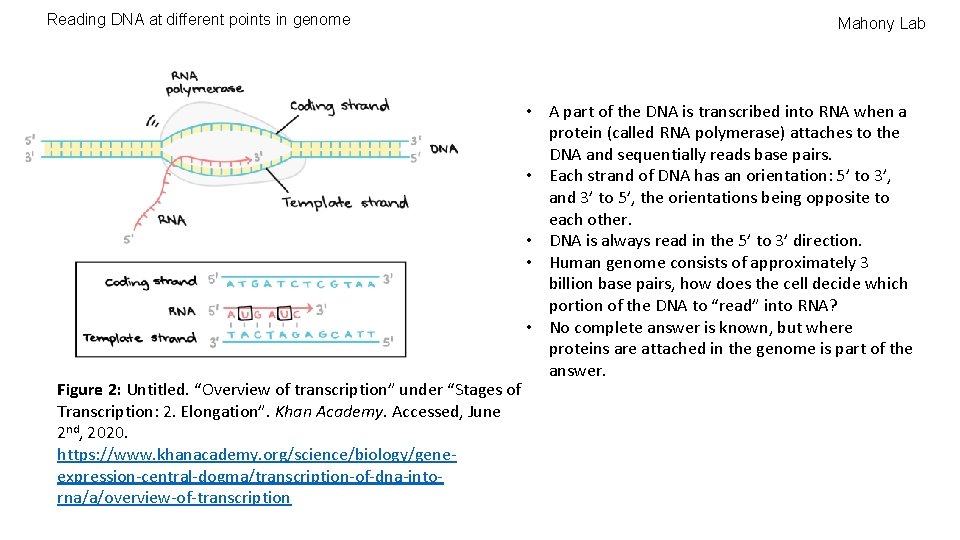
Reading DNA at different points in genome Figure 2: Untitled. “Overview of transcription” under “Stages of Transcription: 2. Elongation”. Khan Academy. Accessed, June 2 nd, 2020. https: //www. khanacademy. org/science/biology/geneexpression-central-dogma/transcription-of-dna-intorna/a/overview-of-transcription Mahony Lab • A part of the DNA is transcribed into RNA when a protein (called RNA polymerase) attaches to the DNA and sequentially reads base pairs. • Each strand of DNA has an orientation: 5’ to 3’, and 3’ to 5’, the orientations being opposite to each other. • DNA is always read in the 5’ to 3’ direction. • Human genome consists of approximately 3 billion base pairs, how does the cell decide which portion of the DNA to “read” into RNA? • No complete answer is known, but where proteins are attached in the genome is part of the answer.

Reading DNA at different points in genome Figure 3: Untitled. “Overview: Gene regulation in bacteria” under “Anatomy of an operon”. Khan Academy. Accessed, June 2 nd, 2020. https: //www. khanacademy. org/science/biology/geneexpression-central-dogma/transcription-of-dna-intorna/a/overview-of-transcription Mahony Lab • Proteins have an affinity to attach to DNA sequences. • Proteins attached at certain parts of the genome can form logical gates to either allow or disallow RNA polymerase to attach there. • Often what controls a particular attribute is more complex than a single stretch of DNA. • How to determine where proteins are attached in the genome? • Proteins that attach to DNA to regulate conversion to RNA are called Transcription Factors (TFs)
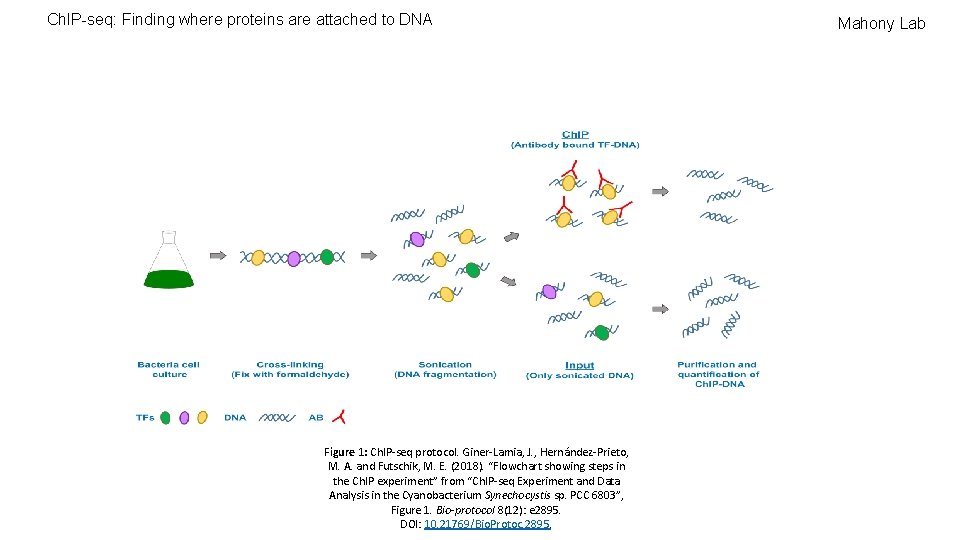
Ch. IP-seq: Finding where proteins are attached to DNA Figure 1: Ch. IP-seq protocol. Giner-Lamia, J. , Hernández-Prieto, M. A. and Futschik, M. E. (2018). “Flowchart showing steps in the Ch. IP experiment” from “Ch. IP-seq Experiment and Data Analysis in the Cyanobacterium Synechocystis sp. PCC 6803”, Figure 1. Bio-protocol 8(12): e 2895. DOI: 10. 21769/Bio. Protoc. 2895. Mahony Lab

Ch. IP-seq (contd. ) • DNA fragments are sequenced (or read) in the 5’ to 3’ direction. • Sequencers per base accuracy decreases as they read more base pairs. Read length varies but around 50 bp (base pairs) long are common. The typical size of a fragment is 150 -250 bp. • The Human Genome Project was a massive effort to determine the entire sequence of the genome from a few humans. • As the genome between humans doesn’t vary “a lot”, we assume that every human has the same genome and perform an approximate string match of our 50 bp across the whole genome to find its position. • Sequencing doesn’t tell us which strand (Plus or Minus) the read came from. • However, when a read is aligned to the genome, it will be aligned to either the plus or minus strand. Mahony Lab

Example: Ch. IP-seq read alignment 5’ ACTGATTGCATGGCCAAAATGTGAACTTTTGGGG 3’ TGACTAACGTACCGGTTTTACACTTGAAAACCCC 5’ 3’ Figure 4: Example Genome 5’ ACTGATTGCATGGCCAAAATGTGAACTTTTGGGG 3’ TGACTAACGTACCGGTTTTACACTTGAAAACCCC 5’ 3’ Figure 5: Aligning GATTGC 5’ ACTGATTGCATGGCCAAAATGTGAACTTTTGGGG 3’ TGACTAACGTACCGGTTTTACACTTGAAAACCCC 5’ 3’ Figure 6: Aligning GTTCAC Mahony Lab • Align the following reads to the example genome: • GATTGCXXXXX • GTTCACXXXXX • Given a set of reads (also called tags), want to find areas of “large tag pileup”. These are known as peaks. Based on how we set up our experiment, they should correspond to where the protein is bound on the DNA.

MACS 2 Step 1: Decide the Shift Size d/2 Figure 7: 5’ Ends of tags for a FOXA 1 Ch. IP. Adapted from “Model-based Analysis of Ch. IP-Seq (MACS). ”, Figure 1 a. Zhang, Y. , Liu, T. , Meyer, C. A. et al. Genome Biol 9, R 137 (2008). https: //doi. org/10. 1186/gb-20089 -9 -r 137 Mahony Lab • Sonication is less likely to fragment the DNA where the protein is bound, so for the actual location of protein there will be a dip in number of reads starting at that position. • The length between the two summits for any two peaks is assumed to be the binding location (or atleast where the protein is inhibiting sonication). • Shift windows of size 2 x sonication size across the genome. • Sample 1000 sites whose enrichment (# of tags) is some m-fold enriched compared to what you would expect from a random tag distribution. • From these 1000 sites, calculate an average value of d. • Perform a read extension of d towards 3’ end for each tag. (In the MACS paper, this is stated as shift by d/2 towards 3’ direction then extend by d/2 in each direction. This is equivalent to our (the MACS 2 software) statement)

MACS 2 Step 1: Decide the Shift Size d/2 (detailed, contd. ) Mahony Lab 1 bp (on +/- strand) Probability of tag at this position: 1/2 s Window of size 2 x sonication size Expected # of tags in window: (2 x f x t)/ (2 x s) • Assume that tags are uniformly randomly distributed across the genome in the treatment experiment. • Let: • t = total # of tags • f = Sonication size/bandwidth • s = Genome size • mlow = m-fold lower bound. (default 5) • mup = m-fold upper bound. (default 50) • Given a particular base pair (bp) in the genome and a tag, let X be a Bernoulli random variable which is 1 if the left end (5’ end) of the tag aligns there and 0 otherwise. • P(X = 1) = 1/2 s. • E(X) = 1/2 s. • For t trials, E(t. X) = t/2 s. • Let E be the expected # of tags in a random window of size 2 f across the genome. If the number of tags P in a window of size 2 f is: mlow x E < P < mup x E, we call this a peak. • 1000 such peaks are sampled to find the shift size d/2.

MACS 2 Step 2: Shift windows of length 2 d across genome 2 d 2 d λ Figure 8: Depicted in the left and right pictures is the same window of length 2 d of the genome. The left picture shows the # of tags in the window in the treatment. The right picture is the expected # of tags from the control Mahony Lab

MACS 2 Step 2: Shift windows of length 2 d across genome (more details) 2 d 2 d λ Figure 9: Depicted in the left and right pictures is the same window of length 2 d of the genome. The left picture shows the # of tags in the window in the treatment. The right picture is the expected # of tags from the control Mahony Lab
- Slides: 12