Bio Make Specifying biocompute pipelines with Makefiles and

Bio. Make Specifying biocompute pipelines with Makefiles and beyond Chris Mungall BDGP
There is no escaping pipelines in bioinformatics ● ● A pipeline is any specification of a set of tasks which are interdependent Compute pipelines – ● Data transformation pipelines – ● genomic, transcriptomic, proteomic and others db exports, generating xml/html, data warehouses Laboratory Information Management Systems (LIMS). . .
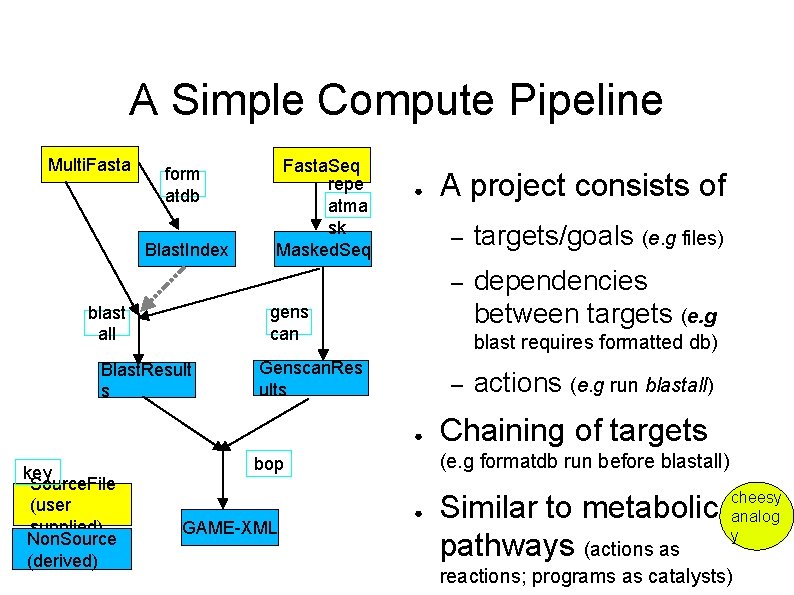
A Simple Compute Pipeline Multi. Fasta form atdb Blast. Index Fasta. Seq repe atma sk Masked. Seq ● – targets/goals (e. g files) – dependencies between targets (e. g gens can blast all Blast. Result s blast requires formatted db) Genscan. Res ults – ● key Source. File (user supplied) Non. Source (derived) A project consists of Chaining of targets (e. g formatdb run before blastall) bop GAME-XML actions (e. g run blastall) ● Similar to metabolic pathways (actions as cheesy analog y reactions; programs as catalysts)

First Attempt – Shell Pipes action. A input-file | action. B - | action. C - > final-target generalised form rmask myseq. fst | blastx mydb - | bop - > myseq. genscan rmask myseq. fst | genscan - | bop - > myseq. genscan cat glucose | hexo-kinase - | fructose-bis-kinase - > glucose-2 -6 -p – Unix shell pipe '|' – feeds the output of one program in as the input to another; hides intermediate files (e. g glucose-6 -p) ● OK for manual use, less good for full automation ● Linear dependency flows only; pipelines are graphs ● Not all programs accept '-' as an input; need intermediate tmp files ● no re-use of previous results (full path recompute!) ● no knowledge of when to re-compute final target

Second Attempt – write a script – Code pipeline in an imperative language (java/perl/C/C++/python/etc) ● explicitly code rules such as – ● ● don't recompute if we have output file already Repetitive, redundant code; unaesthetic Underlying logic becomes obscured once we start extending things (asynchronous execution via PBS, comparing timestamps to determine if a recompute is required) sub blastpipe ($mydb, $myseq): formatdb $mydb unless -f $mydb rmask $myseq > $myseq. m unless -f $myseq. m $blastout = blast-$mydb-$myseq. out blastx $mydb $myseq > $blastout unless -f $blastout

Third Attempt – Objects class Blast. Job extends Job: method run { $seq = $self->seq $mydb = $self->mydb $j = new Format. DBJob($seq) $self->check($j) $self->shell(blastx $mydb $seq) ● Attempts to abstract and reuse code – Similar problems to scripting – Different approaches, but end results are very complex (eg gadfly-pipe, biopipe, ensembl) and (IMHO) somewhat unsatisfactory – Crux of problem: mixing code/rules with data/config

Back to basics: Makefiles ● Makefiles are used in a variety of contexts – ● Mostly for building and compiling applications ● . class file depends on. java file; ● . jar file depends on list of. class files ● avoids unnecessary re-compiling; uses timestamps A Makefile is a set of rules – Each rule has a target, dependencies (subtargets) and action(s) – Targets are files, actions are unix shell commands
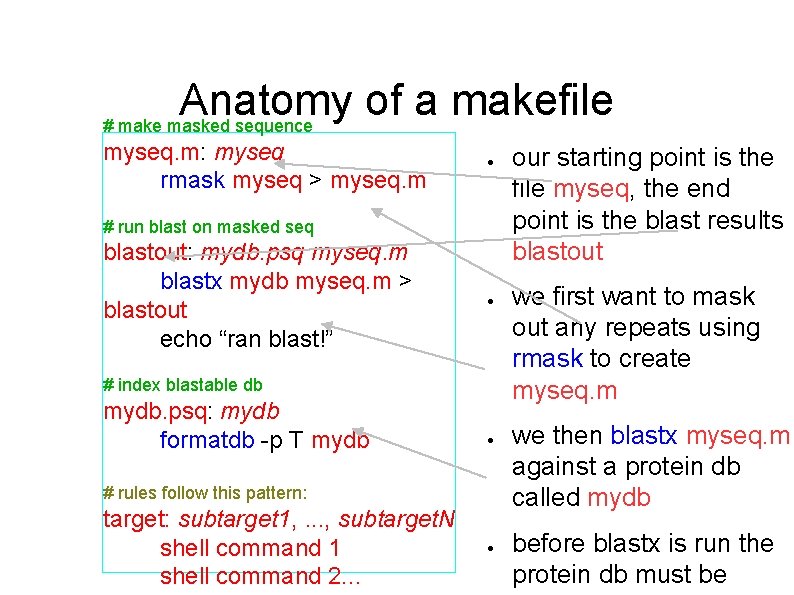
Anatomy of a makefile # make masked sequence myseq. m: myseq rmask myseq > myseq. m ● # run blast on masked seq blastout: mydb. psq myseq. m blastx mydb myseq. m > blastout echo “ran blast!” ● # index blastable db mydb. psq: mydb formatdb -p T mydb ● # rules follow this pattern: target: subtarget 1, . . . , subtarget. N shell command 1 shell command 2. . . ● our starting point is the file myseq, the end point is the blast results blastout we first want to mask out any repeats using rmask to create myseq. m we then blastx myseq. m against a protein db called mydb before blastx is run the protein db must be

The “make” command # run blast on masked seq blastout: mydb. psq myseq. m blastx mydb myseq. m > blastout echo “ran blast!” # index blastable db mydb. psq: mydb formatdb -p T mydb # make masked sequence myseq. m: myseq rmask myseq > myseq. m % make blastout formatdb -p T mydb rmask myseq. fst > myseq. m blastx mydb myseq. m > blastout % make blastout make: 'blastout' is up to date % cat newseqs >> mydb % make blastout formatdb -p T mydb blastx mydb myseq. m > blastout ● make uses unix file modification timestamps when checking dependencies – if a subtarget is more recent than the goal target, then re-execute

If compounds were files and cells were makefiles things might look a bit like this # glucose-2 -6 -p synthesis from glucose via glucose-6 -p # using fructose-bis-kinase and hexo-kinase-p enymes as catalysts # (all reactions carried out by unix command react) cheesy analog y # final step glucose-2 -6 -p: glucose-6 -p fructose-bis-kinase react -catalyst fructose-bis-kinase glucose-6 -p > glucose-2 -6 -p # initial step glucose-6 -p: glucose hexo-kinase-p react -catalyst hexo-kinase-p glucose > glucose-6 -p

Get wild with makefiles ● The preceding examples used hardcoded targets – ● ● ● what if we want to use the same logic over multiple targets (eg multiple input sequence fasta files and/or multiple blast databases)? Makefiles allow this via * wildcards Wildcards allow basic 'parameterization' of make rules Syntax is a little strange – even stranger than perl

Pattern matching examples ● blastout_%. m: mydb. psq %. m blastx mydb $*. m > $@ %. m: %. fst rmask %. fst > $@ ● ● % make blastout_seq 1. m rmask seq 1. fst > seq 1. m blastx mydb seq 1. m > blastout_seq 1. m ● ● A % is a wildcard match on the target/dependency file A $* in the action line(s) is replaced with the match A $@ in the action line is replaced by the target/goal Concise but ugly! We can iterate through lots of input

Defensive makefile logic blastout_%. m: mydb. psq %. m blastx mydb $*. m > $@. tmp && mv $@. tmp $@ %. m: %. fst rmask %. fst > $@. tmp && mv $@. tmp $@ ● Be on guard – make sure no target is created if the action fails, otherwise you get rubbish all the way downstream – ● With the unix shell construct foo 1 && foo 2, foo 2 will not be run if foo 1 exits with non-zero status (error) Things are looking even uglier! (but safer)

(some) Makefile limitations ● Wildcard problems: difficult to have more than one wildcard parameter for a rule – blast needs at least 2: fastadb and input sequence ● Dependencies inherently tied to filesystem ● Logic is fundamentally synchronous (always waits for actions to finish) – not good for managing remotely executed tasks ● Nasty syntax takes a while to master (and is easily forgotten) ● No programming language constructs ● Too dumb/low-level; leads to repetitive patterns One side effect of these limitations is the large number of makefile makers; these typically take a makefile templates and automatically generate one or more makefiles. This takes us back to evil scripting logic! Despite this, makefiles are an invaluable tool. But can they be improved?

Alternatives to make ● nmake – minor improvements (aimed at software maintenance) ● scons ● build ● apache ant – ● XML/java based - focused mainly on java code maintenance (does this very well) Nothing that would suffice for a full-on

Some things that would be nice ● ● Abstract away from filesystem Specify Goals and requirements as parameterized functions blastout_%. m: mydb. psq %. m blastx mydb $*. m > $@ NO! maskblastout(DB, Seq): formatdb(DB) rmask(Seq) blastx DB rmask(Seq) > target() YES!

Bio. Make using skam ● Skam – Skolem Asynchronous Make – Functional-Logical Makefile replacement ● Lightweight (~400 lines of prolog code! prolog does most of the work) ● Like makefiles, but with function constructs (aka skolems) ● Works with filesystem, databases, GH-style datastores (URLs? ) ● Sync and Async dependency logic – works with remote execution e. g PBS ● Fully programmable + other nice features (see later) ● As generic as makefiles – no bioinformatics logic – Bio. Make ● A set of rules useful for bioinformatics pipelines

Skam is based on skolem function constructs – May be recursively applied; functions as trees – A skolem function uniquely identifies a target / piece of data; for example: xml 2 html(bop(genscan(rmask(My. Seq. ID)))) ● – This skolem identifies the data that comes from repeatmasking My. Seq. ID, putting it through genscan, feeding the results to bop, then transforming the resulting GAME XML to HTML In the example above, My. Seq. ID is a variable

Tangent: Other uses of skolems not necessarily related to pipelines ● Uniquely identifying features typed by SO – – general form for splices sites, exons and introns ● donor(Gene. ID, Number) ● acceptor(Gene. ID, Number) ● intron(Donor. ID, Acceptor. ID) ● exon({Acceptor. ID, init}, {Donor. ID, term}) } primitive } derived features can be uniquely and persistently identified with nested functions ● intron(donor(CG 1234, 4), acceptor(CG 1234, 7)) ● exon(acceptor(CG 1234, 7), term)

Example Skolem Rule mblastx(Seq, DB) flat: blast_results/Seq. DB. blastx req: formatdb(DB) rmask(Seq) srun: blastx -filter SEG+XNU DB rmask(Seq) comment: this target is for the results of running blastx on a masked input genomic sequence (wublast) – skolem function followed by tag: value metadata – variables always begin with Upper. Case

formatdb(DB) req: DB run: formatdb DB comment: prepares blastdb for blasting (wublast) rmask(Seq) flat: masked_seqs/Seq. masked req: Seq srun: Repeat. Masker -lib $(LIB) Seq comment: masks out repeats from input sequence mblastx(Seq, DB) flat: blast_results/Seq. DB. blastx req: formatdb(DB) rmask(Seq) srun: blastx -filter SEG+XNU DB rmask(Seq) comment: this target is for the results of running blastx on a masked input genomic sequence (wublast)

Skolem Specifications ● Each rule concerns a skolem function construct – ● eg sim 4(Seq, DB) [similar to makefile target] Each goal has metadata tag: value pairs, including – flat: How to “flatten” skolem function (to map to filesystem/URL/whatever) ● – req: required Sub. Goals (also specified as functions) ● – [just like makefile dependencies] (s)run: Shell command for generating target ● – [unix makefiles conflate this with the target] [just like makefile action] Other meta tags allowed (. . . )

Bio. Make Data ● relational (ie tabular) data can be included – use xml-like <data>. . . </data> tags ● implemented as prolog facts ● useful for – specifying metadata on local fasta databases ● logical name, file path, SO type, alphabet, species ● we can use this to intelligently automate analyses

Example data section <data relation="fastadb" delimiter="ws" spec="fastadb(ID, Seq. Alphabet, SOType, Bop. Type, Taxon)" comment="formal description of local multifasta datasources"> na_pe. dros dmel ne_EST. all_nr. dros dmel na_gb. dros dmel na_c. DNA. dros dmel na_DGC. dros dmel na_ARGs. dros dmel na_users. dros dmel </data> na transposable_element P_Element na EST m. RNA na m. RNA c. DNA na EST na m. RNA EST

Controlling Bio. Make logic with Rules ● Data can be queried with prolog rules ● Useful for ● – Adding logic/decisions/code to specifications – figuring out which blast program to run against which fastadb – Or for determinig bop type – Querying ontologies – eg SO Skolem targets can be qualified with prolog constraints in squiggly braces {. . . }

<prolog> exprdb(D): - fastadb(D, na, 'm. RNA'). % SO type m. RNA is expressed exprdb(D): - fastadb(D, na, 'sn. RNA'). % SO type sn. RNA is expressed exprdb(D): - fastadb(D, na, 'EST'). % SO type EST is expressed % only use sim 4 against fastadbs of expressed seq from dmel sim 4 db(D): - exprdb(D), fastadb(D, _, _, _, dmel). </prolog> all_sim 4(Seq) req: L {setof(sim 4(S, D)), sim 4 db(D), L)} comment: the requirement for this target is to sim 4 the sequence S against all databases D such that sim 4 db(D) is true; i. e. All databases of expressed drosophila melanogaster sequence. We do this with the prolog predicate setof, which finds all solutions to a problem.

Runmodes ● Default runmode is 'sync' (runs locally, waits until sh command is done before proceeding – just like makefiles) ● PBS wrapper also provided ● Extensible – you can add other runmodes – run in background, locally – rsh to remote machine and run

Specifying Runmodes % specify runmode as a USER-VARIABLE; can % be overridden on the command line $(RUNMODE) = async(submit('qsub -q qresearch 1@bam')) mblastx(Seq, DB) flat: blast_results/Seq. DB. blastx req: formatdb(DB) rmask(Seq) srun: blastx -filter SEG+XNU DB rmask(Seq) runode: $(RUNMODE)

How skam executes actions ● ● Actions are specified with the run: tag ● Actions are unix shell (sh) commands ● Function arguments are flattened ● sh command is wrapped in a shell mini-pipeline Check status functions for goal; run if no status – mini-pipeline can be run locally/remotely, synchronously/asynchronously (eg PBS) ● shell pipe deals with copying/moving files and setting status flags
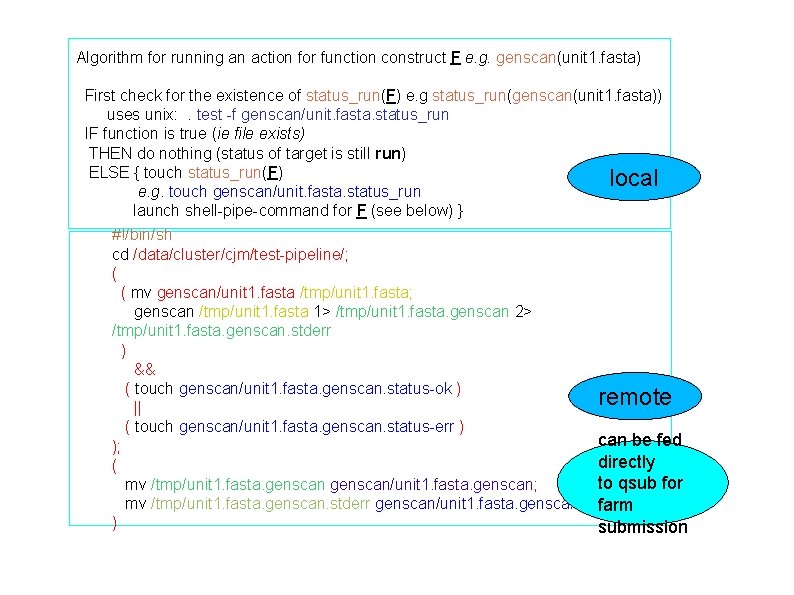
Algorithm for running an action for function construct F e. g. genscan(unit 1. fasta) First check for the existence of status_run(F) e. g status_run(genscan(unit 1. fasta)) uses unix: . test -f genscan/unit. fasta. status_run IF function is true (ie file exists) THEN do nothing (status of target is still run) ELSE { touch status_run(F) local e. g. touch genscan/unit. fasta. status_run launch shell-pipe-command for F (see below) } #!/bin/sh cd /data/cluster/cjm/test-pipeline/; ( ( mv genscan/unit 1. fasta /tmp/unit 1. fasta; genscan /tmp/unit 1. fasta 1> /tmp/unit 1. fasta. genscan 2> /tmp/unit 1. fasta. genscan. stderr ) && ( touch genscan/unit 1. fasta. genscan. status-ok ) remote || ( touch genscan/unit 1. fasta. genscan. status-err ) can be fed ); directly ( to qsub for mv /tmp/unit 1. fasta. genscan/unit 1. fasta. genscan; mv /tmp/unit 1. fasta. genscan. stderr genscan/unit 1. fasta. genscan. stderr; farm ) submission

Data. Shifters ● Bio. Make defaults to the local filesystem ● ● ● function constructs flattened to make relative file paths files automatically shifted between working dir and temporary dir Other Data. Shifters possible – just provide a wrapper script that implements simple basic data-atom operations (fetch, store, test-id) – My. SQL wrapper provided (tiny fast perl script) ● copies files back and forth between tmpdir and db
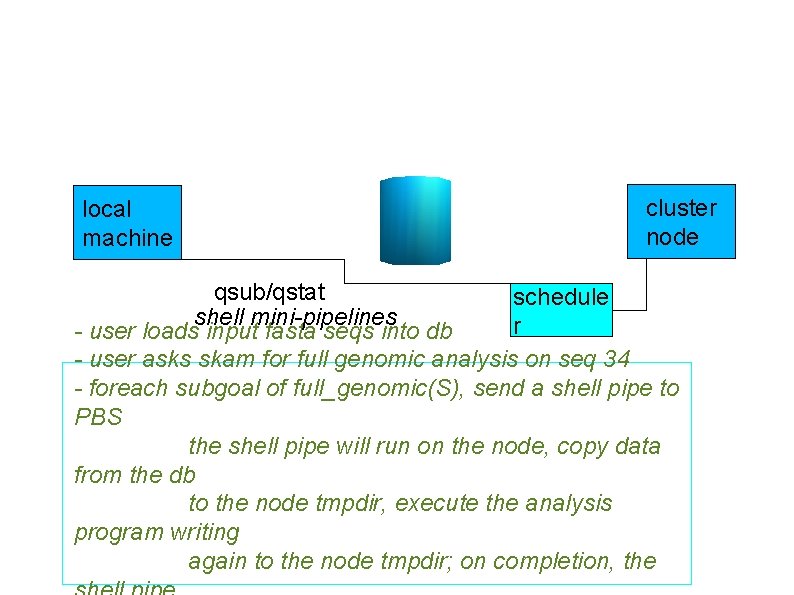
local machine cluster node qsub/qstat schedule shell mini-pipelines r - user loads input fasta seqs into db - user asks skam for full genomic analysis on seq 34 - foreach subgoal of full_genomic(S), send a shell pipe to PBS the shell pipe will run on the node, copy data from the db to the node tmpdir, execute the analysis program writing again to the node tmpdir; on completion, the

Status/Discussion ● Almost ready for public consumption – memory/garbage collection problem ● No fancy user interface – just command line ● Prolog: extremely powerful and concise – may be abstruse to some ● ● . . . but should be hidden behind the scenes Almost in use for GO/OBO – db construction, release management and file transformations
- Slides: 33