Abstract Molecular nanotechnology is the precise threedimensional control

Abstract Molecular nanotechnology is the precise, three-dimensional control of materials and devices at the atomic scale. An important part of nanotechnology is the design of molecules for specific purposes. This paper describes early results using genetic software techniques to automatically design molecules under the control of a fitness function. The fitness function must be capable of determining which of two arbitrary molecules is better for a specific task. The software begins by generating a population of random molecules. The population is then evolved towards greater fitness by randomly combining parts of the better individuals to create new molecules. These new molecules then replace some of the worst molecules in the population. The unique aspect of our approach is that we apply genetic crossover to molecules represented by graphs, i. e. , sets of atoms and the bonds that connect them. We present evidence suggesting that crossover alone, operating on graphs, can evolve any possible molecule given an appropriate fitness function and a population containing both rings and chains. Most prior work evolved strings or trees that were subsequently processed to generate molecular graphs. In principle, genetic graph software should be able to evolve other graph representable systems such as circuits, transportation networks, metabolic pathways, computer networks, etc. Al Globus, John Lawton, and Todd Wipke

Previous work David Weininger, Patent US 5434796, Daylight Chemical Information Systems, Inc. 1995 • two parameter crossover; fragments • some commercial success 4 Robert B. Nachbar, Merck Research Laboratories, "Molecular evolution: a hierarchical representation for chemical topology and its automated manipulation, " Proceedings of the. Third Annual Genetic Programming Conference, University of Wisconsin, Madison, Wisconsin, 22 -25 July 1998, pages 246 -253, 1998 • tree representation • no crossover within rings 4 Astro Teller, CMU, Neural Programming, personal communication 4 Acknowledgments • Creon Levit, NASA Ames • Jason Lohn, Caelum, Inc. at NASA Ames • Rich Mc. Clellan, UCSC • Subash Saini, NASA Ames • Meyyapan, NASA Ames 4

Genetic software Randomly generate a set of molecules 4 Many times: • Select parent molecules at random with bias towards better performance • Randomly rip copies of each parent in two • Mate opposite halves • Replace random molecules with bias towards worse performance 4 Repeat until satisfied 4 4 Algorithm properties • Stochastic, embarrassingly parallel • Robust to failure • No guaranteed outcome • Fitness function is crucial and non-trivial • Performs well as cycle-scavenger using Condor, University of Wisconsin, http: //www. cs. wisc. edu/condor
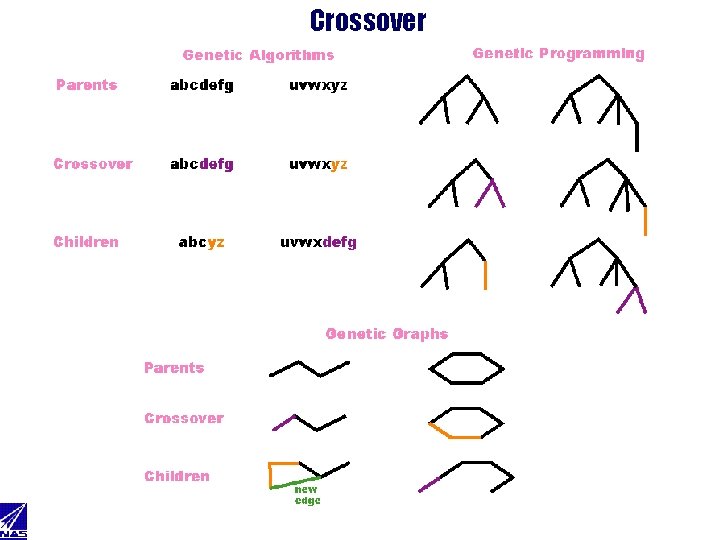
Crossover

Graph Crossover Rip in half • Choose random bond • Find the shortest path • Remove and remember random path bond • Repeat until cut set found 4 Mate halves • Select a random cut bond • If cut bond in other half exist – choose one at random – merge cut bonds, respect valence • else – flip coin – heads -- attach cut bond to random atom in other half respecting valence – tails -- discard cut bond • repeat until all cut bonds processed 4

Fitness function General • Given two molecules, decides which is better • Return a number for each molecule • Must operate on any molecule, including very bad ones • Must provide routes for evolution to reach good molecules • Must make fine distinctions 4 Evolve towards molecular target • All-pairs-shortest-path (APSP) distance – Assign extended types to each atom (element plus bond pattern) – Find shortest path between each pair of atoms – Create bag with one element per shortest path. Each element is the (sorted) extended types of the end point and the number of bonds in the (shortest) path – A bag is a set with duplicate elements – Tanimoto distance between bags – |intersection| / |union| 4

Time to find small molecules
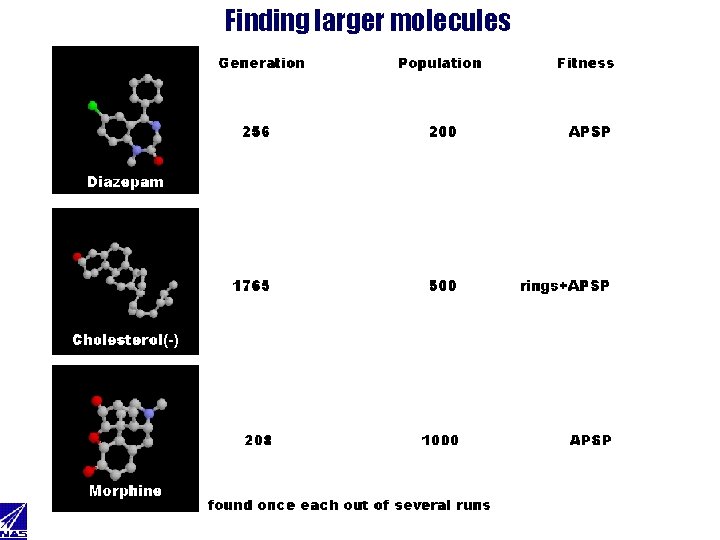
Finding larger molecules
- Slides: 8