Stubbs Lab Bioinformatics 4 Alignment Summary Report Count

Stubbs Lab Bioinformatics – 4 Alignment Summary Report & Count files with htseq-count Nov 29, 2016 Joe Troy

Agenda • One new Linux command - htop • The RNA-Seq Analysis so far • Creating an alignment summary report • Creating count files with htseq-count
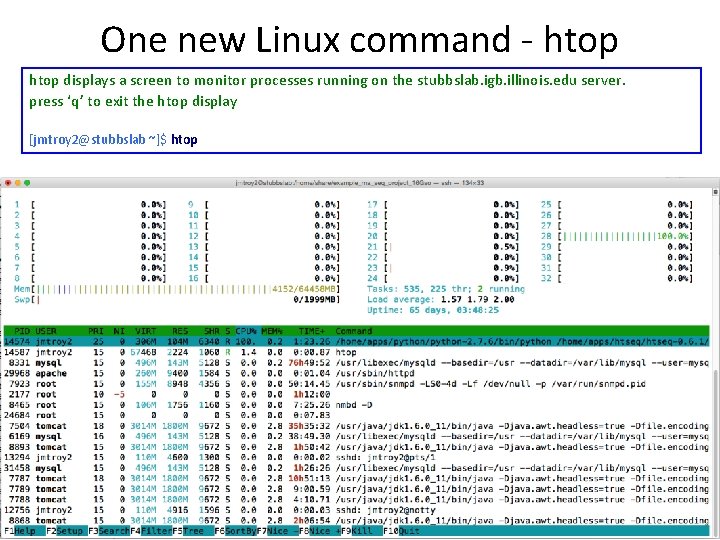
One new Linux command - htop displays a screen to monitor processes running on the stubbslab. igb. illinois. edu server. press ‘q’ to exit the htop display [jmtroy 2@stubbslab ~]$ htop
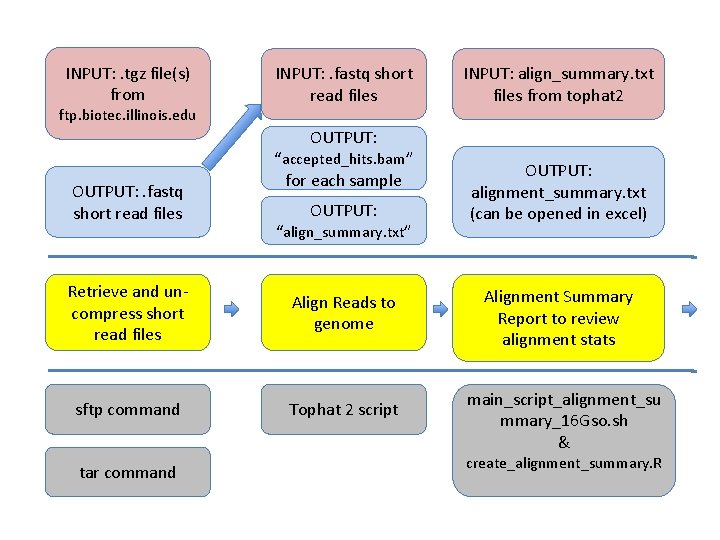
INPUT: . tgz file(s) from ftp. biotec. illinois. edu INPUT: . fastq short read files INPUT: align_summary. txt files from tophat 2 OUTPUT: . fastq short read files “accepted_hits. bam” for each sample OUTPUT: “align_summary. txt” Retrieve and uncompress short read files Align Reads to genome sftp command Tophat 2 script tar command OUTPUT: alignment_summary. txt (can be opened in excel) Alignment Summary Report to review alignment stats main_script_alignment_su mmary_16 Gso. sh & create_alignment_summary. R
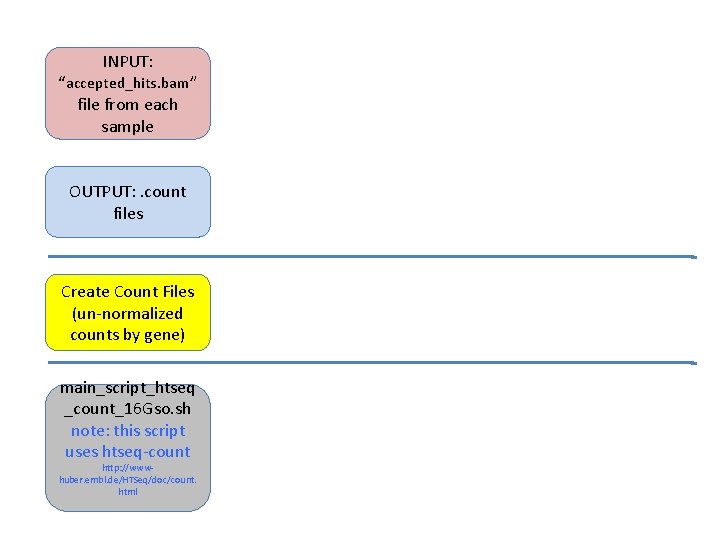
INPUT: “accepted_hits. bam” file from each sample OUTPUT: . count files Create Count Files (un-normalized counts by gene) main_script_htseq _count_16 Gso. sh note: this script uses htseq-count http: //wwwhuber. embl. de/HTSeq/doc/count. html

Instructions to create an alignment summary report INSTRUCTION SLIDE 1 of 2 – no need to use ‘screen’ as it runs quick! Josephs-Mac. Book-Pro: ~ josephtroy$ ssh jmtroy 2@stubbslab. igb. illinois. edu's password: Last login: Mon Nov 21 20: 15: 51 2016 from c-73 -73 -226 -74. hsd 1. il. comcast. net [jmtroy 2@stubbslab ~]$ df -h Filesystem Size Used Avail Use% Mounted on /dev/sda 1 4. 6 T 4. 2 T 150 G 97% / /dev/sda 2 95 G 14 G 77 G 16% /var /dev/sdb 1 289 M 246 M 11% /boot tmpfs 32 G 0% /dev/shm /dev/sdb 2 275 G 116 G 145 G 45% /var/lib/ mysql [jmtroy 2@stubbslab ~]$ cd /home/share/example_rna_seq_project_16 Gso/ [jmtroy 2@stubbslab example_rna_seq_project_16 Gso]$ cd code_020_alignment_summary_report/ [jmtroy 2@stubbslab code_020_alignment_summary_report]$ ls -lh total 16 K -rw-rw-r-- 1 jmtroy 2 598 Nov 27 12: 00 alighment_report_heading. txt -rw-rw-r-- 1 jmtroy 2 3. 9 K Nov 26 19: 04 create_alignment_summary. R -rw-rw-r-- 1 jmtroy 2 4. 8 K Nov 27 12: 14 main_script_alignment_summary_16 Gso. sh [jmtroy 2@stubbslab code_020_alignment_summary_report]$ sh main_script_alignment_summary_16 Gso. sh [1] "start parameter loop" [1] "end parameterloop" end script main_script_alignment_summary_16 Gso. sh at Sun Nov 27 12: 20: 46 CST 2016 [jmtroy 2@stubbslab code_020_alignment_summary_report]$
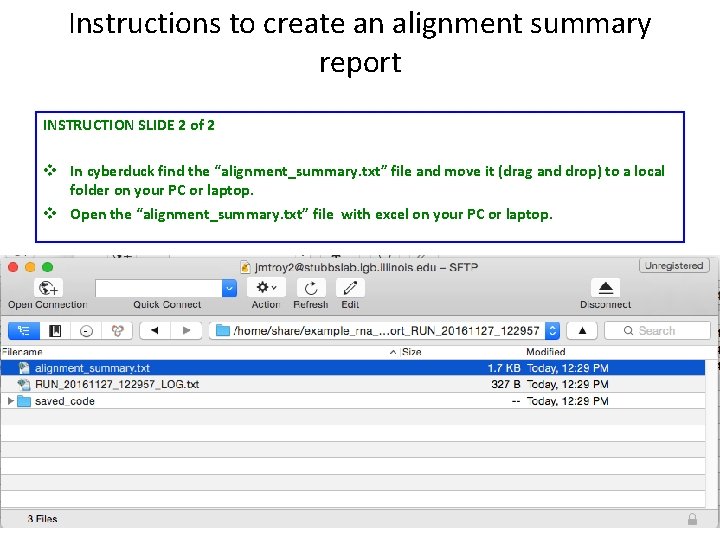
Instructions to create an alignment summary report INSTRUCTION SLIDE 2 of 2 v In cyberduck find the “alignment_summary. txt” file and move it (drag and drop) to a local folder on your PC or laptop. v Open the “alignment_summary. txt” file with excel on your PC or laptop.
Instructions to create count files with htseq-count http: //www-huber. embl. de/HTSeq/doc/count. html INSTRUCTION SLIDE 1 of 3 Josephs-Mac. Book-Pro: ~ josephtroy$ ssh jmtroy 2@stubbslab. igb. illinois. edu's password: Last login: Mon Nov 21 20: 15: 51 2016 from c-73 -73 -226 -74. hsd 1. il. comcast. net [jmtroy 2@stubbslab ~]$ df -h Filesystem Size Used Avail Use% Mounted on /dev/sda 1 4. 6 T 4. 2 T 150 G 97% / /dev/sda 2 95 G 14 G 77 G 16% /var /dev/sdb 1 289 M 246 M 11% /boot tmpfs 32 G 0% /dev/shm /dev/sdb 2 275 G 116 G 145 G 45% /var/lib/mysql [jmtroy 2@stubbslab example_rna_seq_project_16 Gso]$ screen

Instructions to create count files with htseq-count http: //www-huber. embl. de/HTSeq/doc/count. html INSTRUCTION SLIDE 2 of 3 [jmtroy 2@stubbslab ~]$ cd /home/share/example_rna_seq_project_16 Gso/ [jmtroy 2@stubbslab example_rna_seq_project_16 Gso]$ cd code_025_htseq_count/ [jmtroy 2@stubbslab code_025_htseq_count]$ ls main_script_htseq_count_16 Gso. sh [jmtroy 2@stubbslab code_025_htseq_count]$ sh main_script_htseq_count_16 Gso. sh … NOW HOLD DOWN THE CONTROL KEY AND PRESS a, THEN PRESS d, TO DETACH FROM THE SCREEN SESSION Options for monitoring a running process: v Use cyberduck (don’t forget to keep refreshing the screen) to watch the output folder. v Use the htop command to monitor system activity. v Reattach the screen session for your process. Press ENTER a few times, and when the command prompt appears the process is done.

Instructions to create count files with htseq-count http: //www-huber. embl. de/HTSeq/doc/count. html INSTRUCTION SLIDE 3 of 3 Remember: the process number ‘ 11559’ will be different when you run this exercise [jmtroy 2@stubbslab ~]$ screen -ls There is a screen on: 11559. pts-2. stubbslab (Detached) 1 Socket in /var/run/screen/S-jmtroy 2. [jmtroy 2@stubbslab ~]$ screen -r 11559 [jmtroy 2@stubbslab code_025_htseq_count]$ exit [jmtroy 2@stubbslab ~]$ screen -ls No Sockets found in /var/run/screen/S-jmtroy 2.
- Slides: 10