Progress report on Crank Experimental phasing Biophysical Structural

Progress report on Crank: Experimental phasing Biophysical Structural Chemistry Leiden University, The Netherlands

Crank developments available in CCP 4 6. 1 • “Greatly enhanced” – better tested • Underlying programs haven’t changed (much), but crank almost completely re-written from version in 6. 0. 2 • Better ccp 4 i interface • Support for more programs (PIRATE, BUCCANEER, RESOLVE, COOT) • Faster substructure detection – Use BP 3 to (quickly) check trials and look at deviations between different CRUNCH 2 trials significantly decreases the time required for successful substructure detection.
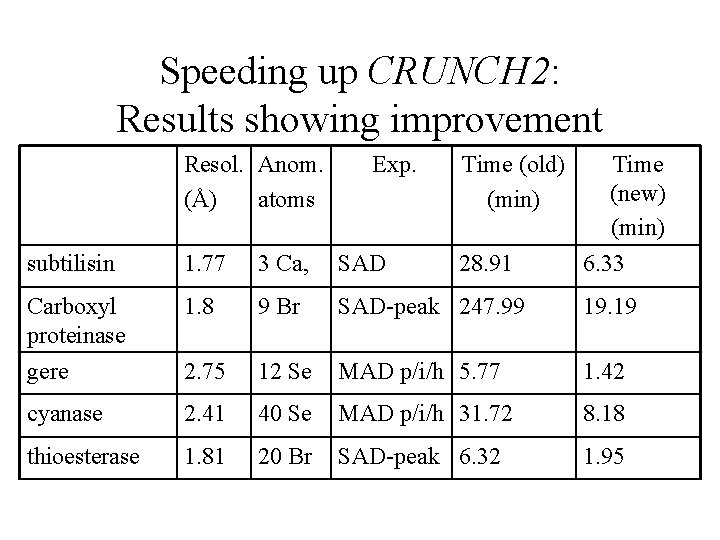
Speeding up CRUNCH 2: Results showing improvement Resol. Anom. (Å) atoms Exp. Time (old) (min) 28. 91 Time (new) (min) 6. 33 subtilisin 1. 77 3 Ca, SAD Carboxyl proteinase gere 1. 8 9 Br SAD-peak 247. 99 19. 19 2. 75 12 Se MAD p/i/h 5. 77 1. 42 cyanase 2. 41 40 Se MAD p/i/h 31. 72 8. 18 thioesterase 1. 81 20 Br SAD-peak 6. 32 1. 95
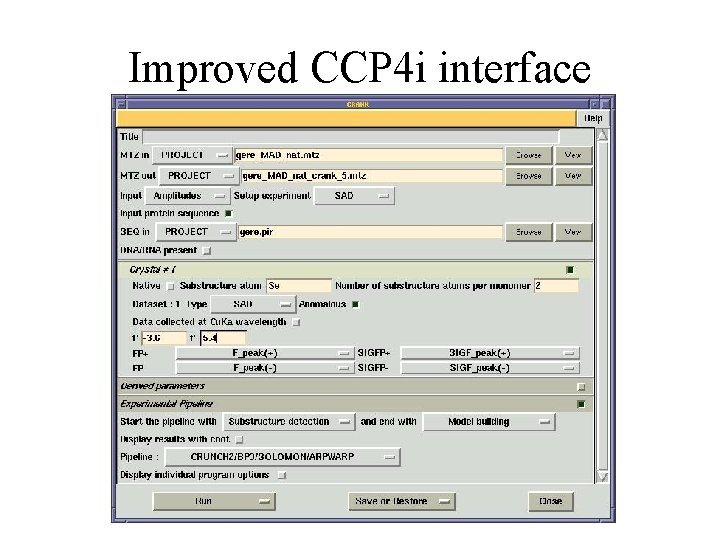
Improved CCP 4 i interface

Preliminary substructure detection results from JCSG test cases • 144 mostly MAD Se-Met data sets • Defaults only: the only input was number of Se. Met per monomer (number of monomer was guessed). Mtz files, f’, f”. • Some data sets had f” < 1 (solved by MR) • Some data sets had incorrectly labelled X-PLOR files as mtz. • DISCLAIMER: 1 st logfiles produced analyzed yesterday after dinner (until 4 a. m. ).
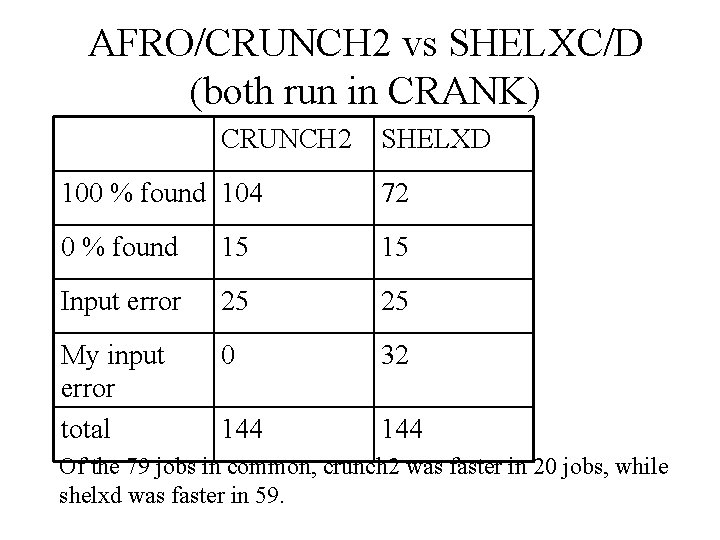
AFRO/CRUNCH 2 vs SHELXC/D (both run in CRANK) CRUNCH 2 SHELXD 100 % found 104 72 0 % found 15 15 Input error 25 25 My input error total 0 32 144 Of the 79 jobs in common, crunch 2 was faster in 20 jobs, while shelxd was faster in 59.

Comparison not fair • Same algorithm to identify solution with BP 3 can be used in SHELXD • SHELXD uses much better Fa values (i. e. using the MAD data – at the moment, Afro just uses delta F from the data set with the greatest anomalous signal).

Improving FA values • An early step in solving a structure by SAD/MAD or SIRAS is to determine FA values. • FA is the structure factor amplitude corresponding to the substructure to input to direct methods and/or Patterson programs (i. e. SHELXD or CRUNCH 2)

Current FA estimation • FA is currently estimated by | |F+| - |F-| | for SAD data in most programs. • Direct method programs are very sensitive to FA values. • Improving estimates can improve hit rates of direct methods and solve substructures that can not previously been solved.

Multivariate SAD equation E(|FA|, |F+|, |F-|) = |FA| P(|FA|, αA, | |F+|, α+, |F-|, α-) d|FA| dαA dα+dα • Giacovazzo previously proposed multivariate FA estimation, with an implementation assuming Bijvoet phases are equal. • An equation can be obtained without the equal phase assumption requiring only one numerical integration. • The equation has been implemented – which reduces to Giacovazzo’s equation if Bijvoet phases are equal.

Covariance matrix properties • The covariance matrix considers experimental sigmas and correlations between F+, F- and FA. • Problem: Covariance matrix also depends on (overall) substructure occupancy and b-factor. • Solution: Obtain a multivariate likelihood estimate for unknown parameters.

Refining overall substructure parameters • Initial guess of number of substructure atoms per monomer obtained from user. • Initial guess of B-factor obtained from likelihood estimate of overall B-factor of data set. • Result: Refinement is stable and maximizes correlation with calculated final E’s. • Another possible application: Use refined overall occupancy and B-factor for anomalous signal estimation.
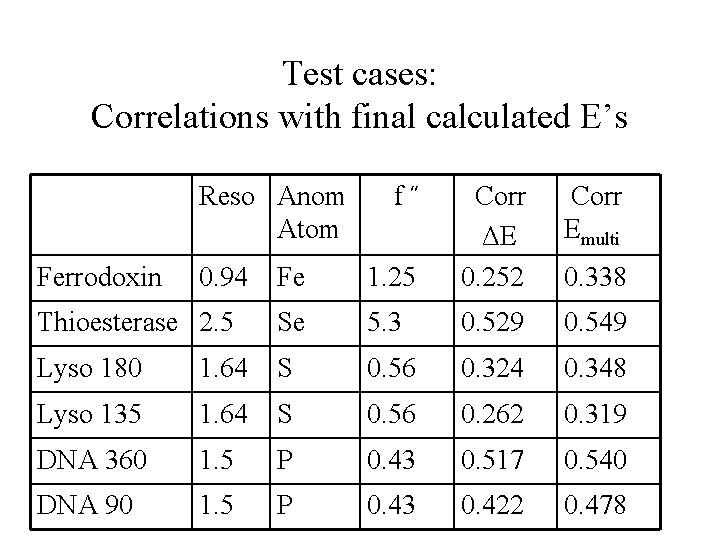
Test cases: Correlations with final calculated E’s Reso Anom Atom Fe 1. 25 Corr ΔE 0. 252 Thioesterase 2. 5 Se 5. 3 0. 529 0. 549 Lyso 180 1. 64 S 0. 56 0. 324 0. 348 Lyso 135 1. 64 S 0. 56 0. 262 0. 319 DNA 360 1. 5 P 0. 43 0. 517 0. 540 DNA 90 1. 5 P 0. 43 0. 422 0. 478 Ferrodoxin 0. 94 f˝ Corr Emulti 0. 338
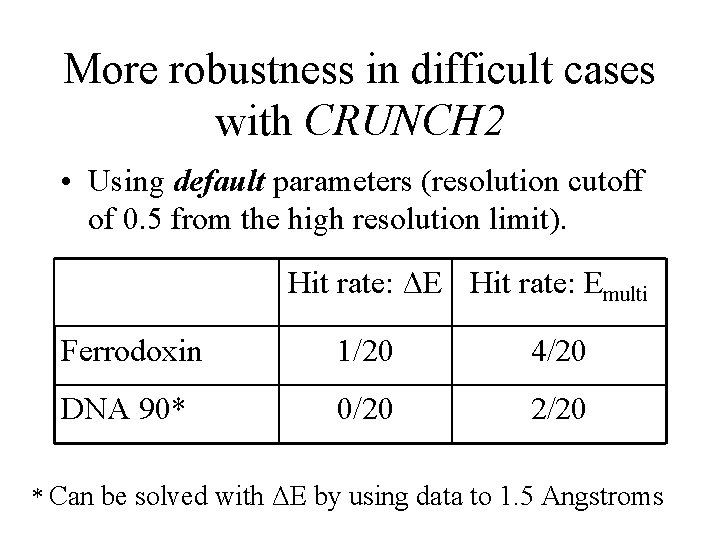
More robustness in difficult cases with CRUNCH 2 • Using default parameters (resolution cutoff of 0. 5 from the high resolution limit). Hit rate: ΔE Hit rate: Emulti Ferrodoxin 1/20 4/20 DNA 90* 0/20 2/20 * Can be solved with ΔE by using data to 1. 5 Angstroms
- Slides: 14