Platzhalter fr Bild Bild auf Titelfolie hinter das

Platzhalter für Bild, Bild auf Titelfolie hinter das Logo einsetzen Exercise Systems & Synthetic Biotechnology elementary flux mode analysis Veronique Beckers veronique. beckers@uni-saarland. de

Todays exercise 1. Update network construction 2. What to do with a metabolic network? 3. What are elementary flux modes? 4. What knowledge can we gain from elementary mode analysis? 5. Processing of flux modes in MATLAB® -2 -

Update network construction Project: 5 groups • Corynebacterium glutamicum • Aspergillus niger • Saccharomyces cerevisiae • Escherichia coli • Pseudomonas putida substrates: • Glucose • Xylose • Saccharose • Fructose • Glycerin • Lactate Network construction: § Which pathways? § Information required: stoichiometry, reversibility, localization, biomass synthesis requirements § Biomass synthesis: protein composition and precursor demand -3 -
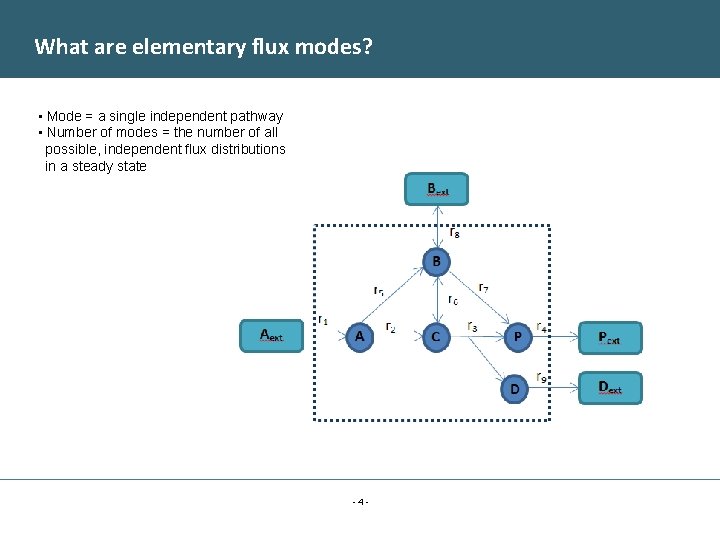
What are elementary flux modes? • Mode = a single independent pathway • Number of modes = the number of all possible, independent flux distributions in a steady state -4 -
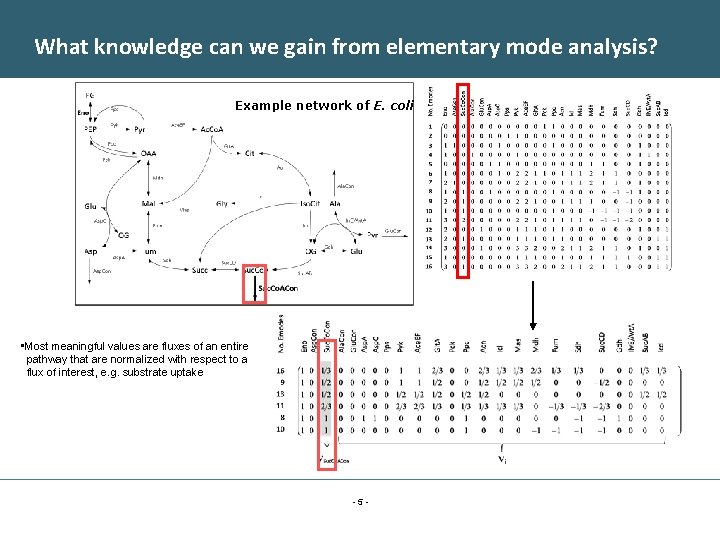
What knowledge can we gain from elementary mode analysis? Example network of E. coli • Most meaningful values are fluxes of an entire pathway that are normalized with respect to a flux of interest, e. g. substrate uptake -5 -
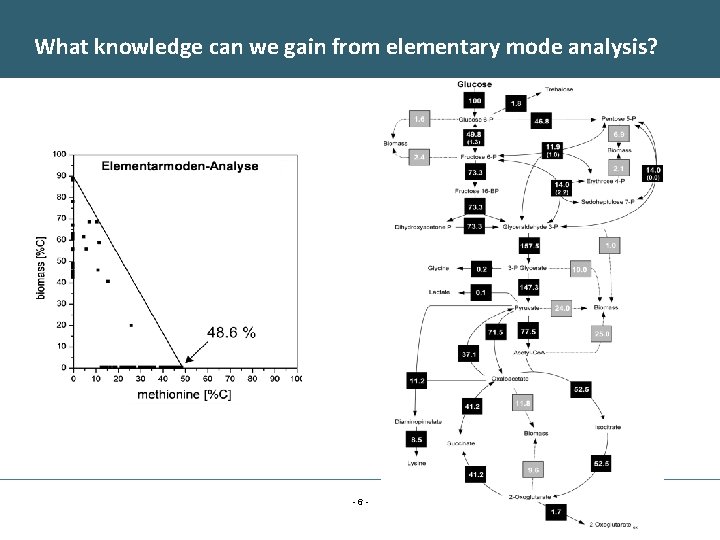
What knowledge can we gain from elementary mode analysis? -6 -
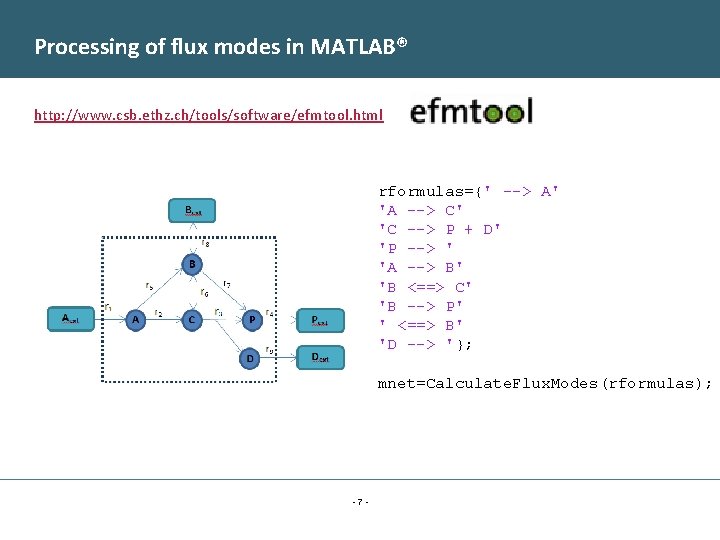
Processing of flux modes in MATLAB® http: //www. csb. ethz. ch/tools/software/efmtool. html rformulas={' --> A' 'A --> C' 'C --> P + D' 'P --> ' 'A --> B' 'B <==> C' 'B --> P' ' <==> B' 'D --> '}; mnet=Calculate. Flux. Modes(rformulas); -7 -
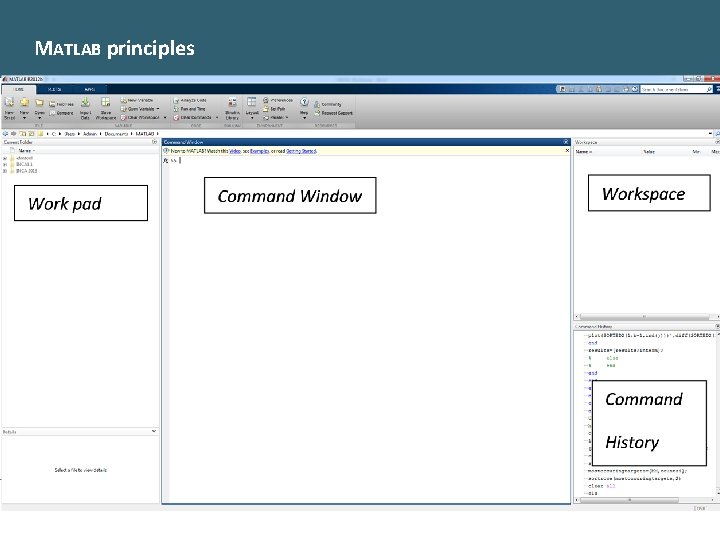
MATLAB principles -8 -
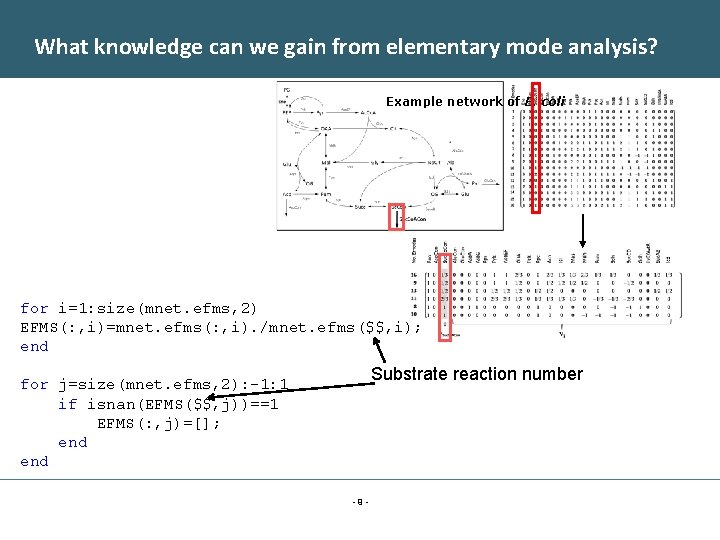
What knowledge can we gain from elementary mode analysis? Example network of E. coli for i=1: size(mnet. efms, 2) EFMS(: , i)=mnet. efms(: , i). /mnet. efms($$, i); end Substrate reaction number for j=size(mnet. efms, 2): -1: 1 if isnan(EFMS($$, j))==1 EFMS(: , j)=[]; end -9 -
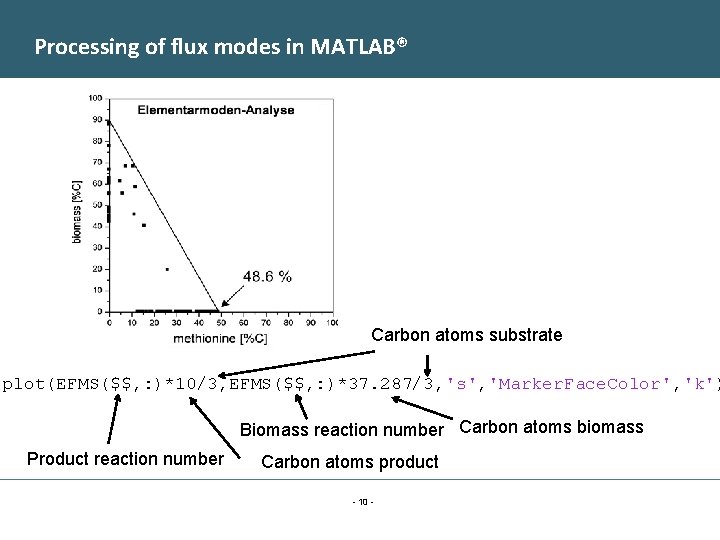
Processing of flux modes in MATLAB® Carbon atoms substrate plot(EFMS($$, : )*10/3, EFMS($$, : )*37. 287/3, 's', 'Marker. Face. Color', 'k') Biomass reaction number Carbon atoms biomass Product reaction number Carbon atoms product - 10 -

Processing of flux modes in MATLAB® Product reaction number indices=find(EFMS($$, : )==max(EFMS($$, : ))); Max. Yield=EFMS(: , indices); - 11 -
- Slides: 11