M roreri DE NOVO GENOME ASSEMBLY USING ABYSS1

M. roreri DE NOVO GENOME ASSEMBLY USING ABYSS/1. 9. 0 MAXK 96 GROUP 5 Hyeim Jung Pedro Pablo Parra Diana Vanessa Sarria Zuniga Jacob Shoemake
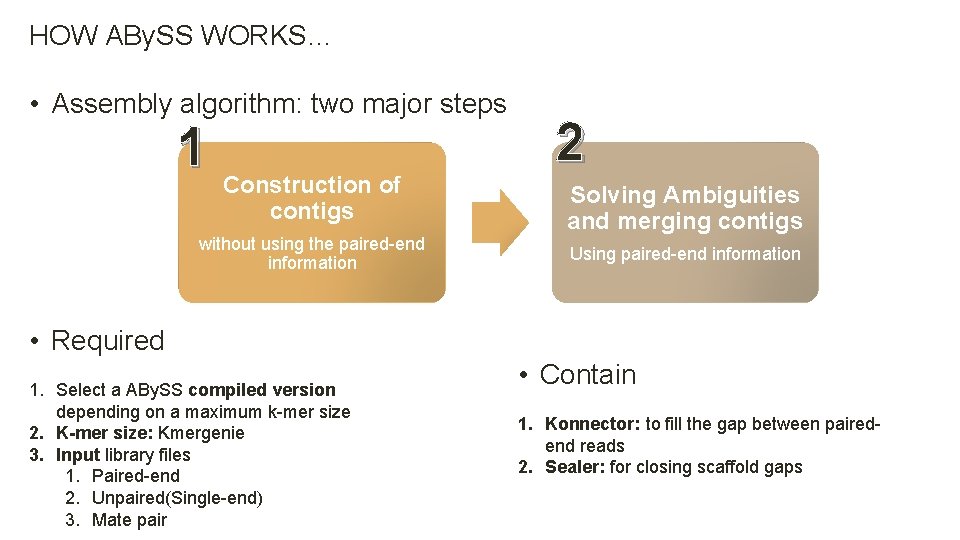
HOW ABy. SS WORKS… • Assembly algorithm: two major steps 1 Construction of contigs without using the paired-end information • Required 1. Select a ABy. SS compiled version depending on a maximum k-mer size 2. K-mer size: Kmergenie 3. Input library files 1. Paired-end 2. Unpaired(Single-end) 3. Mate pair 2 Solving Ambiguities and merging contigs Using paired-end information • Contain 1. Konnector: to fill the gap between pairedend reads 2. Sealer: for closing scaffold gaps

OUR ASSEMBLY STRATEGIES… • Two assembly types • Assembly 5 • Assembly 3 abyss-pe k=87 name=assembly 5 lib='pe 1' mp='mp 1' pe 1=‘paired PE. 1. fq paired PE 2. fq’ abyss-pe k=81 name=assembly 3 lib='pe 1 pe 2' mp='mp 1' pe 1=‘paired PE. 1. fq paired PE 2. fq’ pe 2=‘paired MP. 1. fq paired MP. 2. fq’ se=’unpaired PE-MP’ mp 1=‘paired MP. 1. fq paired MP. 2. fq’ Note: mp 1 is used for scaffolding. Do not contribute to the consensus sequence.
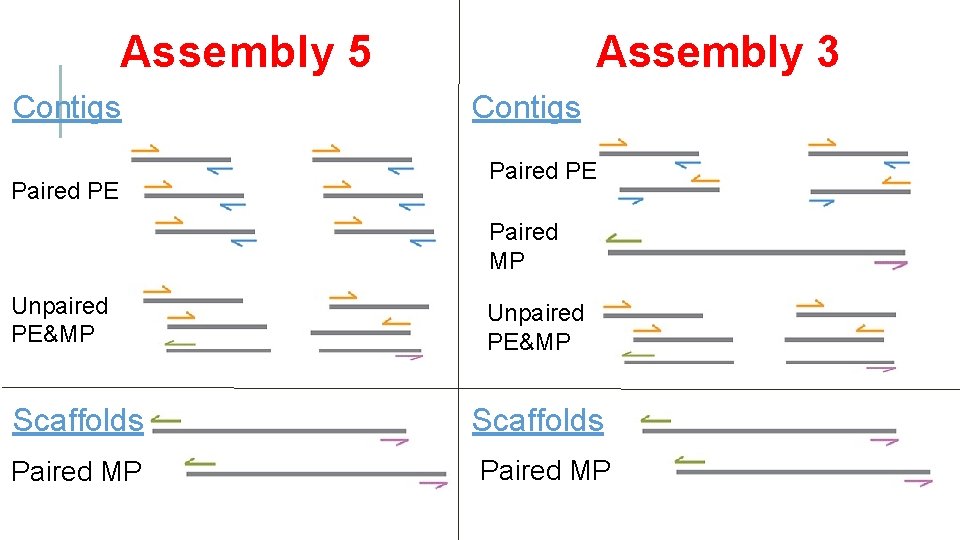
Assembly 5 Contigs Paired PE Assembly 3 Contigs Paired PE Paired MP Unpaired PE&MP Scaffolds Paired MP
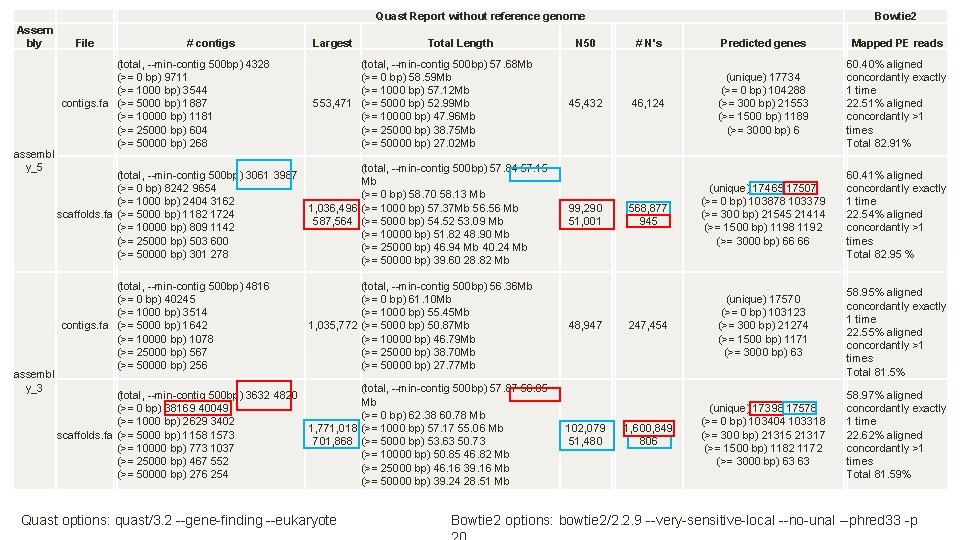
Quast Report without reference genome Assem bly assembl y_5 assembl y_3 File # contigs Largest Total Length N 50 Bowtie 2 # N's Predicted genes Mapped PE reads 46, 124 (unique) 17734 (>= 0 bp) 104288 (>= 300 bp) 21553 (>= 1500 bp) 1189 (>= 3000 bp) 6 60. 40% aligned concordantly exactly 1 time 22. 51% aligned concordantly >1 times Total 82. 91% 568, 877 945 (unique) 17465 17507 (>= 0 bp) 103878 103379 (>= 300 bp) 21545 21414 (>= 1500 bp) 1198 1192 (>= 3000 bp) 66 66 60. 41% aligned concordantly exactly 1 time 22. 54% aligned concordantly >1 times Total 82. 95 % (unique) 17570 (>= 0 bp) 103123 (>= 300 bp) 21274 (>= 1500 bp) 1171 (>= 3000 bp) 63 58. 95% aligned concordantly exactly 1 time 22. 55% aligned concordantly >1 times Total 81. 5% (unique) 17398 17578 (>= 0 bp) 103404 103318 (>= 300 bp) 21315 21317 (>= 1500 bp) 1182 1172 (>= 3000 bp) 63 63 58. 97% aligned concordantly exactly 1 time 22. 62% aligned concordantly >1 times Total 81. 59% EVALUATION OF BEST ASSEMBLIES (total, --min-contig 500 bp) 4328 (>= 0 bp) 9711 (>= 1000 bp) 3544 contigs. fa (>= 5000 bp) 1887 (>= 10000 bp) 1181 (>= 25000 bp) 604 (>= 50000 bp) 268 (total, --min-contig 500 bp) 57. 68 Mb (>= 0 bp) 58. 59 Mb (>= 1000 bp) 57. 12 Mb 553, 471 (>= 5000 bp) 52. 99 Mb (>= 10000 bp) 47. 96 Mb (>= 25000 bp) 38. 75 Mb (>= 50000 bp) 27. 02 Mb (total, --min-contig 500 bp) 57. 84 57. 15 (total, --min-contig 500 bp) 3061 3987 Mb (>= 0 bp) 8242 9654 (>= 0 bp) 58. 70 58. 13 Mb (>= 1000 bp) 2404 3162 1, 036, 496 (>= 1000 bp) 57. 37 Mb 56. 56 Mb scaffolds. fa (>= 5000 bp) 1182 1724 587, 564 (>= 5000 bp) 54. 52 53. 09 Mb (>= 10000 bp) 809 1142 (>= 10000 bp) 51. 82 48. 90 Mb (>= 25000 bp) 503 600 (>= 25000 bp) 46. 94 Mb 40. 24 Mb (>= 50000 bp) 301 278 (>= 50000 bp) 39. 60 28. 82 Mb (total, --min-contig 500 bp) 4816 (>= 0 bp) 40245 (>= 1000 bp) 3514 contigs. fa (>= 5000 bp) 1642 (>= 10000 bp) 1078 (>= 25000 bp) 567 (>= 50000 bp) 256 (total, --min-contig 500 bp) 56. 36 Mb (>= 0 bp) 61. 10 Mb (>= 1000 bp) 55. 45 Mb 1, 035, 772 (>= 5000 bp) 50. 87 Mb (>= 10000 bp) 46. 79 Mb (>= 25000 bp) 38. 70 Mb (>= 50000 bp) 27. 77 Mb (total, --min-contig 500 bp) 57. 87 56. 05 (total, --min-contig 500 bp) 3632 4820 Mb (>= 0 bp) 38169 40049 (>= 0 bp) 62. 38 60. 78 Mb (>= 1000 bp) 2629 3402 1, 771, 018 (>= 1000 bp) 57. 17 55. 06 Mb scaffolds. fa (>= 5000 bp) 1158 1573 701, 868 (>= 5000 bp) 53. 63 50. 73 (>= 10000 bp) 773 1037 (>= 10000 bp) 50. 85 46. 82 Mb (>= 25000 bp) 467 552 (>= 25000 bp) 46. 16 39. 16 Mb (>= 50000 bp) 276 254 (>= 50000 bp) 39. 24 28. 51 Mb Quast options: quast/3. 2 --gene-finding --eukaryote 45, 432 99, 290 51, 001 48, 947 102, 079 51, 480 247, 454 1, 600, 849 806 Bowtie 2 options: bowtie 2/2. 2. 9 --very-sensitive-local --no-unal --phred 33 -p
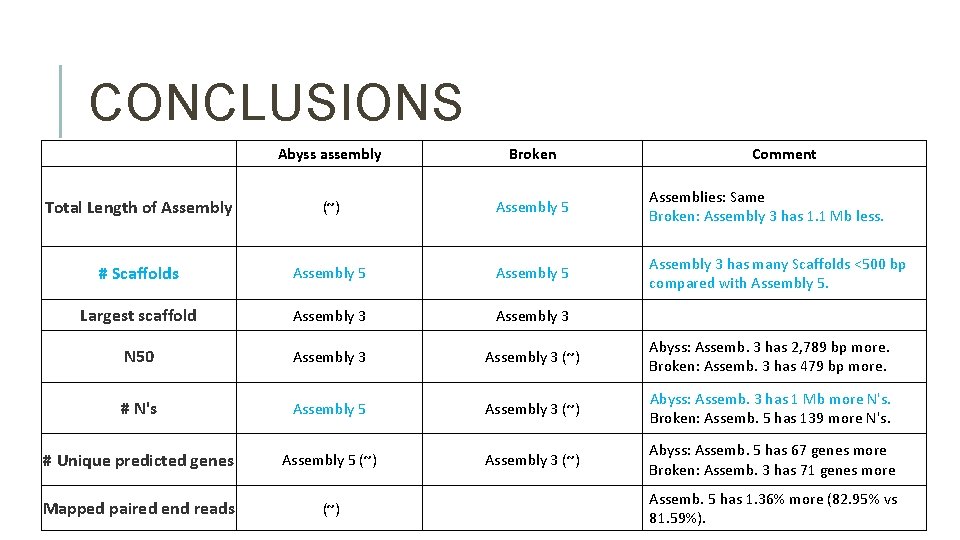
CONCLUSIONS Abyss assembly Broken Comment Total Length of Assembly (~) Assembly 5 Assemblies: Same Broken: Assembly 3 has 1. 1 Mb less. # Scaffolds Assembly 5 Assembly 3 has many Scaffolds <500 bp compared with Assembly 5. Largest scaffold Assembly 3 N 50 Assembly 3 (~) Abyss: Assemb. 3 has 2, 789 bp more. Broken: Assemb. 3 has 479 bp more. # N's Assembly 5 Assembly 3 (~) Abyss: Assemb. 3 has 1 Mb more N's. Broken: Assemb. 5 has 139 more N's. # Unique predicted genes Assembly 5 (~) Assembly 3 (~) Abyss: Assemb. 5 has 67 genes more Broken: Assemb. 3 has 71 genes more Mapped paired end reads (~) Assemb. 5 has 1. 36% more (82. 95% vs 81. 59%).


25298314 reads; of these: 25298314 (100. 00%) were paired; of these: 4322365 (17. 09%) aligned concordantly 0 times 15280202 (60. 40%) aligned concordantly exactly 1 time 5695747 (22. 51%) aligned concordantly >1 times --- 4322365 pairs aligned concordantly 0 times; of these: 2648376 (61. 27%) aligned discordantly 1 time --- 1673989 pairs aligned 0 times concordantly or discordantly; of these: 3347978 mates make up the pairs; of these: 37310 (1. 11%) aligned 0 times 725071 (21. 66%) aligned exactly 1 time 2585597 (77. 23%) aligned >1 times 99. 93% overall alignment rate
- Slides: 8