Genes to Trees Daniel Ayres and Adam Bazinet

Genes to Trees Daniel Ayres and Adam Bazinet CMSC 858 P - Project 2 Proposal
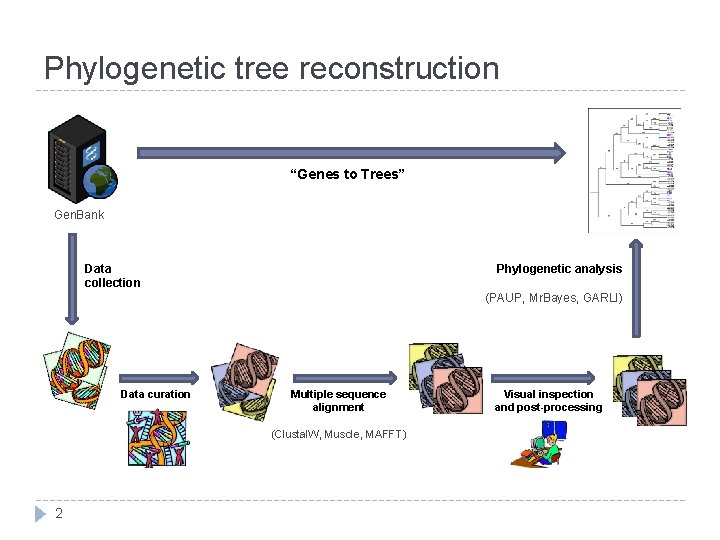
Phylogenetic tree reconstruction “Genes to Trees” Gen. Bank Data collection Phylogenetic analysis (PAUP, Mr. Bayes, GARLI) Data curation Multiple sequence alignment (Clustal. W, Muscle, MAFFT) 2 Visual inspection and post-processing

How does it work? User inputs: Set of DNA or amino acid sequences Taxonomic constraints Workflow Homologous sequences obtained from Gen. Bank Smaller groups eliminated Multiple alignment of each group made Uninformative columns removed “Super-matrix” of all sequences created Phylogenetics analysis performed Output: 3 Phylogenetic tree of closely related organisms

Is it feasible? Scripting will be done with Perl Extensive use of Bio. Perl libraries Collection of modules for bioinformatics programming 4 Accessing sequence data from local and remote databases Manipulating individual sequences Searching for similar sequences Creating and manipulating sequence alignments

Why is this relevant? Results can serve as a starting point for further analysis Multiple analyses can be run in parallel Workflow is modular A step towards robust, high-throughput phylogenetics 5
- Slides: 5