Eran Yanowski Eran Hornsteins Monitor drug impact on














- Slides: 23

Eran Yanowski, Eran Hornstein’s: Monitor drug impact on the transcriptome of mouse beta cells (primary and cell-line) using Transeq/RNA-Seq 17. 08. 15 Report I

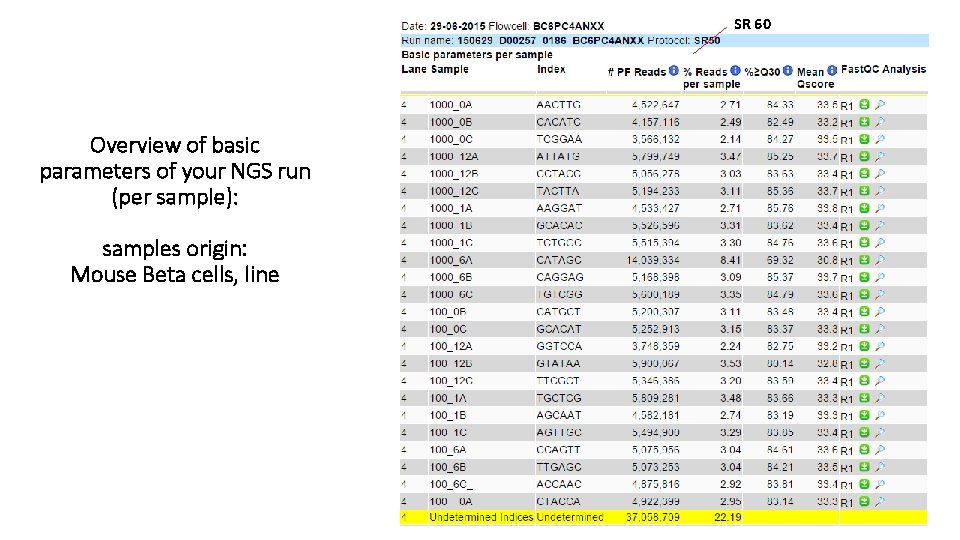
SR 60 Overview of basic parameters of your NGS run (per sample): samples origin: Mouse Beta cells, line

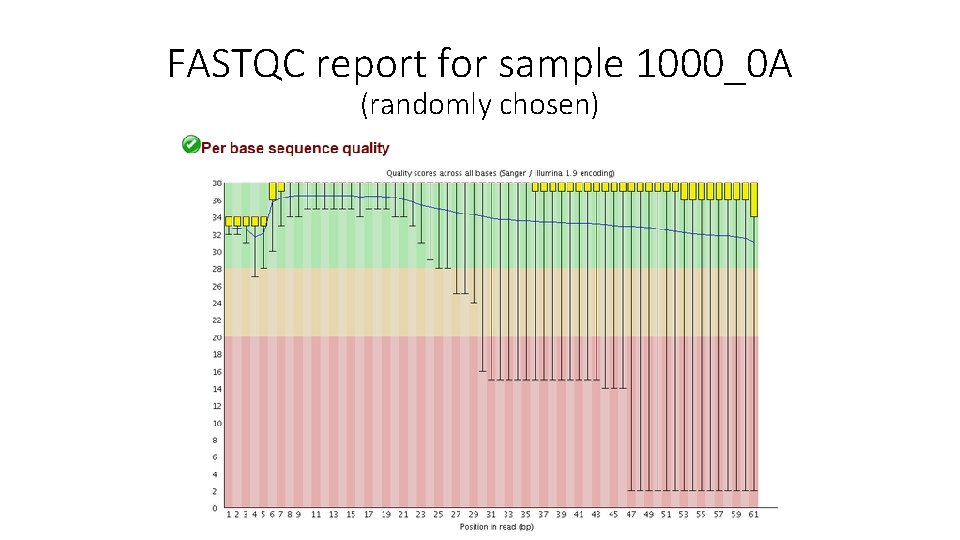
FASTQC report for sample 1000_0 A (randomly chosen)

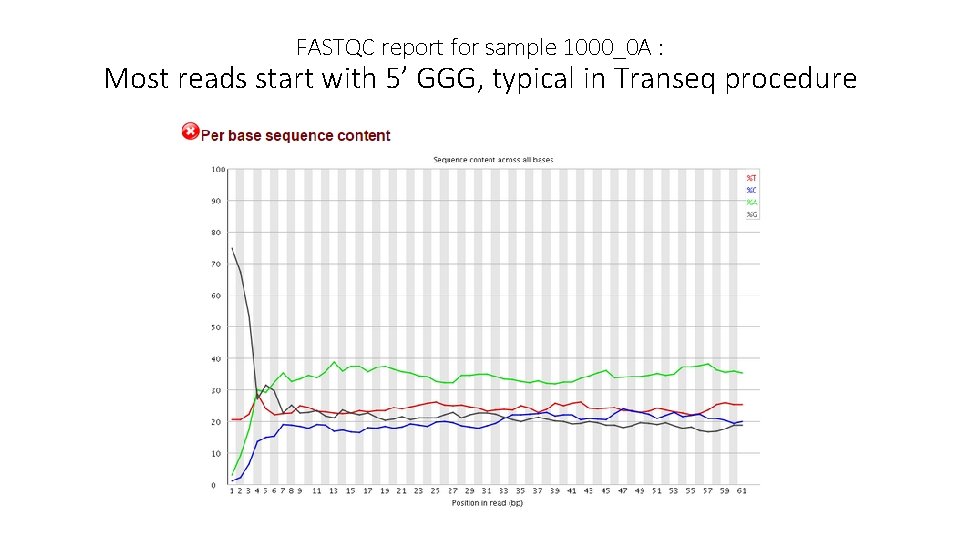
FASTQC report for sample 1000_0 A : Most reads start with 5’ GGG, typical in Transeq procedure

FASTQC report for sample : overrepresentation of Ins 2 reads

Pre-Pipeline data processing Þ Need to remove 5’GGG sequences ÞNeed to remove reads containing either poly-A or poly-T

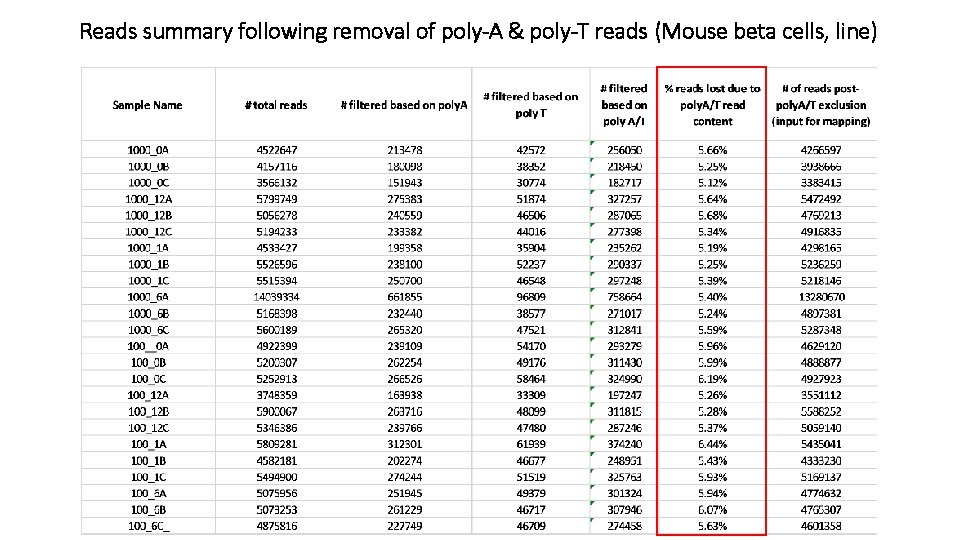
Reads summary following removal of poly-A & poly-T reads (Mouse beta cells, line)

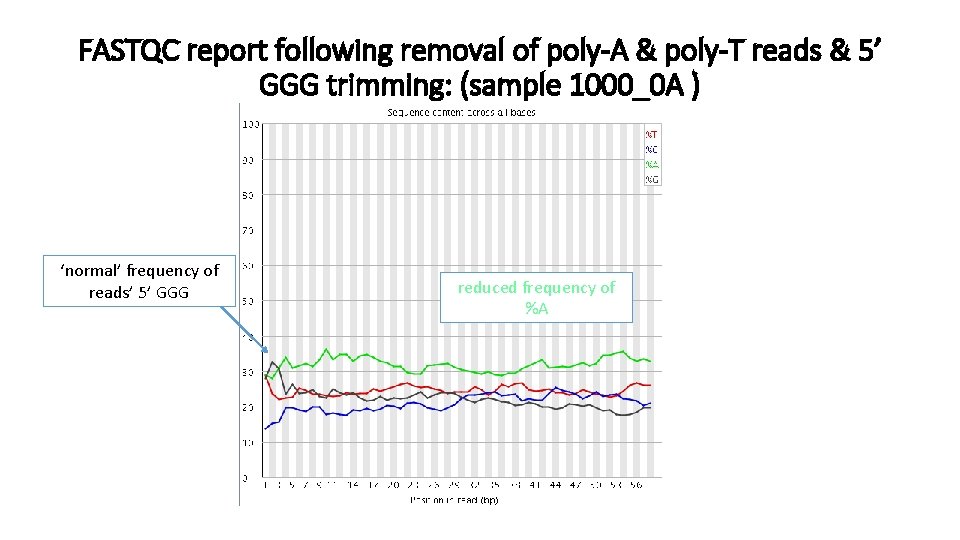
FASTQC report following removal of poly-A & poly-T reads & 5’ GGG trimming: (sample 1000_0 A ) ‘normal’ frequency of reads’ 5’ GGG reduced frequency of %A

Data processing (pipeline) workflow (done using Mouse_mm 9_v 1 base repository) 1. If each sample has more than one fastq file (per sequencing read) then fastq files merging-step is performed 2. Transeq Reads pre-processing (5’ GGG trimming & poly. A & poly. T removal) 3. Processed-reads Mapping (using Top. Hat) 4. TES reads coverage profile (Transeq protocol QC step) 5. Reads Count (per 3’UTR) (using HTSeq-count) 6. Data Normalization and Differential Gene Expression (using DESeq 2) 7. QC: Principal Component Analysis (PCA) & Hierarchical Clustering

Reads mapping summary_Exp 4_1_Mouse Beta cells, line

Reads mapping summary_Exp 4_1_Mouse Beta cells, line: 23 -24% of total reads count were mapped to Ins 2/Ins 1 genes

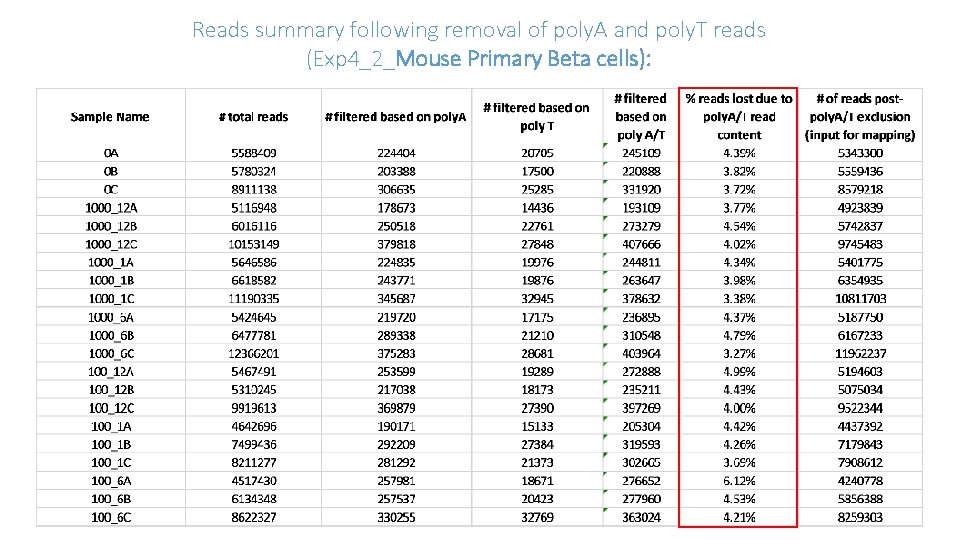
Reads summary following removal of poly. A and poly. T reads (Exp 4_2_Mouse Primary Beta cells):

Reads mapping summary_Exp 4_2_Mouse Primary Beta cells:

Reads mapping summary_Exp 4_2_Mouse Primary Beta cells: % of total reads count were mapped to Ins 2/Ins 1 genes

Hierarchical clustering: Mouse beta cells, line Drug con=100 Drug Con=1000

Hierarchical clustering: Mouse Primary beta cells Drug con=100 Separated by processing day: A/B/C ? Drug Con=1000

• Differential gene expression data is assessed by DESeq 2 • DESeq output is summarized in a single sheet per experiment • Genes differentially-expressed during each time series were called by two independent means: I. Using pairwise comparison vs. time zero II. Using a tool named: ma. Sig. Pro

RNA-Seq drug dose response (1000 and 100)/ time series (0, 1, 6 and 12 hrs) gene filtering criteria • The data filters used during the last analysis performed (12. 08. 15): I. ma. Sig. Pro: • Norm. Counts of genes meeting Max. Raw. Count>50 served as input; • ma. Sig. Pro output was further filtered against potential outlier genes (flagged by ma. Sig. Pro) • Genes showing FC greater than 1. 5 (at least in one of the paired-comparisons); • By default ma. Sig. Pro requires (BH) adjusted p-value <0. 05 II. Pairwise comparison criteria: Max. Raw. Count>50, adjusted-p-value<0. 05 and FC greater than 1. 5 (at least in one of the paired-comparisons);

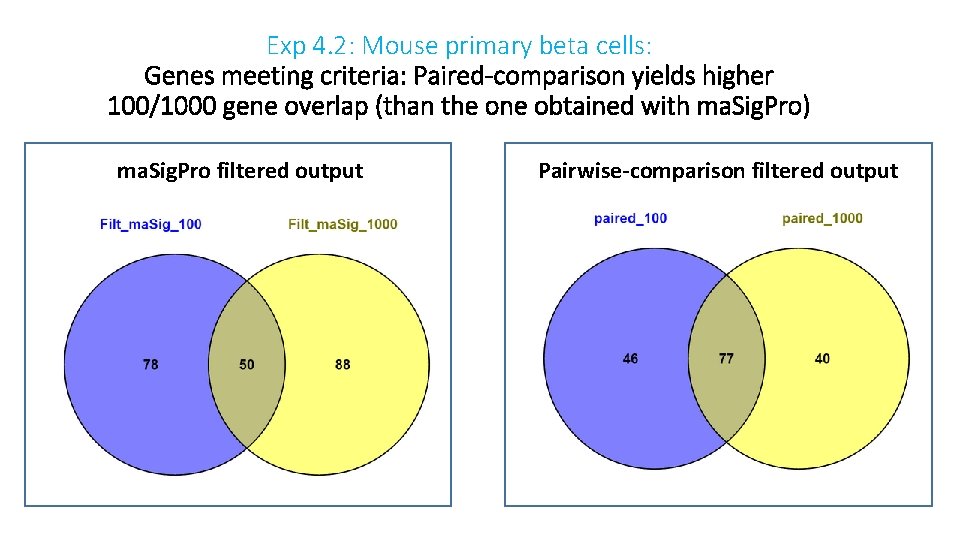
Exp 4. 2: Mouse primary beta cells: Genes meeting criteria: Paired-comparison yields higher 100/1000 gene overlap (than the one obtained with ma. Sig. Pro) ma. Sig. Pro filtered output Pairwise-comparison filtered output

Exp 4. 2: Mouse primary beta cells: Most of ma. Sig. Pro shared-genes are included in the group of paired shared genes To determine which output is preferred data validation using an orthogonal method is essential

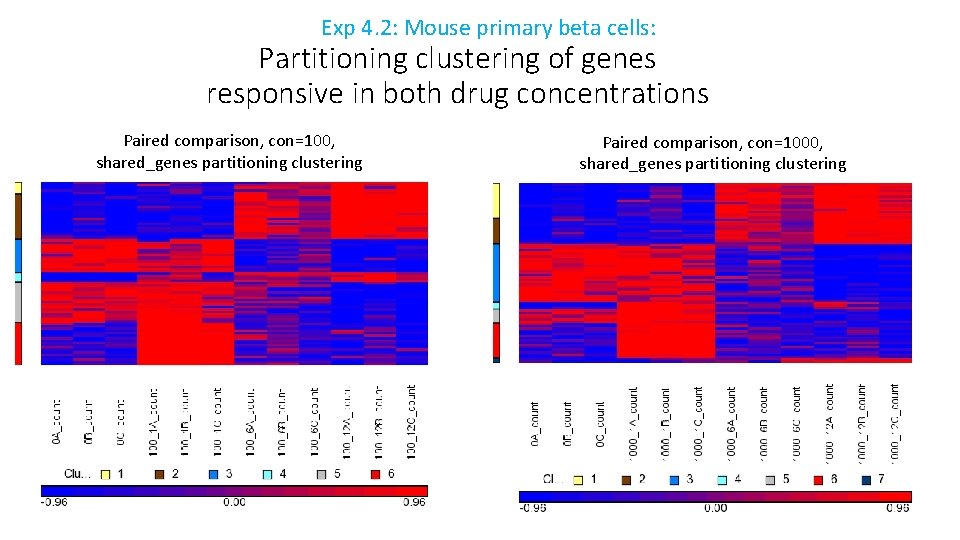
Exp 4. 2: Mouse primary beta cells: Partitioning clustering of genes responsive in both drug concentrations Paired comparison, con=100, shared_genes partitioning clustering Paired comparison, con=1000, shared_genes partitioning clustering

Exp 4. 1: Mouse Beta Cells, cell line • Generally this experiment yielded less significant results • When applying the same filters used for the primary beta-cells datasets, very few genes pass; • The possibility of using p-value (instead of adjusted-p-value should be tested by the investigator) • The level of 100/1000 intersection (shared-genes) is lower here compared to the one observed in the primary cells experiment

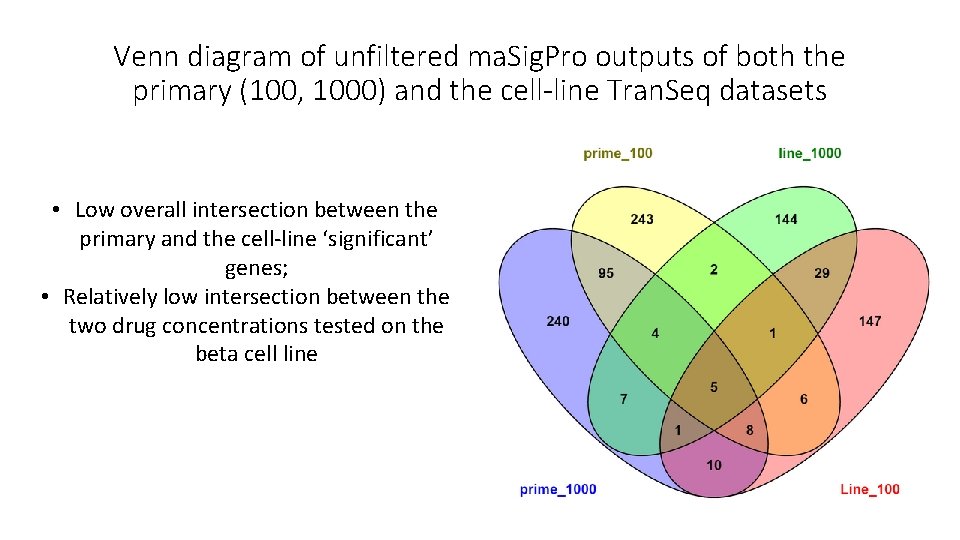
Venn diagram of unfiltered ma. Sig. Pro outputs of both the primary (100, 1000) and the cell-line Tran. Seq datasets • Low overall intersection between the primary and the cell-line ‘significant’ genes; • Relatively low intersection between the two drug concentrations tested on the beta cell line