EE 194BIO 196 Modeling simulating and optimizing biological

EE 194/BIO 196: Modeling, simulating and optimizing biological systems Spring 2018 Tufts University Instructor: Joel Grodstein joel. grodstein@tufts. edu Objects 1

Why do we care about objects • We’ve learned about variables – Variables are great, but there’s only one of each • We’ve learned about arrays – Arrays are great, but … – each member of an array has the same data type and is, conceptually, an instance of the same “thing. ” • Often we want to treat a group of dissimilar things as one big group. – Example: A student has name, UTLN, year, grades, but it's still one student – and there are many students. – We have no good way to do that so far EE 194/Bio 196 Joel Grodstein 2

More examples? • What other examples can you think of? • An item in a store – has price, bar code, shelf location, weight • A Manduca sexta – might have one 2 D array for its leg-locked pattern and another for its muscle-activation pattern – and a variable for its name, perhaps EE 194/Bio 196 Joel Grodstein 3

Demo • Demo using Lunchables to introduce objects. EE 194/Bio 196 Joel Grodstein 4

Impressive terminology • Other courses will take this idea much further: – A student, or a lunch, or a hornworm will be an object. – Nobody has to know what’s inside of it. – You just know what you can do to it; you can make() your lunch, eat() it, or trade() it. • We’re taking the first steps in object-oriented programming. EE 194/Bio 196 Joel Grodstein 5

Objects • Computer programs can get big. Really big. – Mac OS/X is about 85 million of lines of code – Nobody can do that alone; big projects have big teams • Big teams of people don’t really help if everyone has to understand all 85 M lines of code. – We’ve talked about using functions and packages to divide up the work – Objects are the next big tool divide up the work. • One team of people implements a Manduca object – like what you’ll see in the HW. – It can be created, mate and mutate. It can be printed and simulated. • Another team of people uses Manduca. E. g. , put together a genetic algorithm that uses the abilities just described. – They don’t have to know how Manduca is implemented – They need only know what its capabilities are • Object-oriented programming is a Big Deal in Computer Science. EE 194/Bio 196 Joel Grodstein 6

Objects • Let’s start writing code for a Manduca object says that the rest of this code will define a class of object called a “Manduca” says what leg and muscle patterns go into the new Manduca class Manduca: this function def __init__(self, legs, muscles): self. legs=np. copy(legs) tells how to make a new self. muscles=np. copy(muscles) Manduca self. name = "Bob" copy/save the parameters into this Manduca says which of the many Manducas to work with Fine, all worms have the same name! EE 194/Bio 196 Joel Grodstein 7
![The basics my_legs = np. array ([[1, 1, 0, 0, 0], …]) my_musc = The basics my_legs = np. array ([[1, 1, 0, 0, 0], …]) my_musc =](http://slidetodoc.com/presentation_image_h2/0049d108e2da6a4d6ff161cbeec4a21b/image-8.jpg)
The basics my_legs = np. array ([[1, 1, 0, 0, 0], …]) my_musc = np. array ([[100, 0, 0, 100], …]) m 1 = Manduca (my_legs, my_musc) def __init__(self, legs, muscles): self. legs=np. copy(legs) self. muscles=np. copy(muscles) self. name= "Bob" my_legs my_musc legs m 1 muscles name [100, 0, …] [1, 1, 0, …] [100, 0, …] EE 194/Bio 196 Joel Grodstein "Bob" self 8

Next steps • Great, we can make a Manduca. In fact, we can make as many of them as we want: my_legs = np. array ([[1, 1, 0, 0, 0], …]) my_musc = np. array ([[100, 0, 0, 100], …] m 1 = Manduca (my_legs, my_musc) my_legs[3, : ]=[1, 0, 0, 0, 1]; m 2 = manduca (my_legs, my_musc) • But what else can we do with them? – Print them out EE 194/Bio 196 Joel Grodstein 9

Printing an object class Manduca: def __init__(self, legs, muscles, name): … def __repr__(self): str = self. name + " legs | musclesn" for r in range(10): # for each row str+= np. array 2 string (self. legs[r, : ]) + " | " + np. array 2 string (self. muscles[r, : ]) + "n" return(str) • __repr__() is a special function that Python calls whenever you print an object. – It lets you control how an object gets printed. EE 194/Bio 196 Joel Grodstein 10
![Printing example m 1 = Manduca (my_legs, my_musc) my_legs[0, : ]=[1, 0, 0, 0, Printing example m 1 = Manduca (my_legs, my_musc) my_legs[0, : ]=[1, 0, 0, 0,](http://slidetodoc.com/presentation_image_h2/0049d108e2da6a4d6ff161cbeec4a21b/image-11.jpg)
Printing example m 1 = Manduca (my_legs, my_musc) my_legs[0, : ]=[1, 0, 0, 0, 1] m 2 = manduca (my_legs, my_musc) print (m 1) print (m 2) legs muscles [1 1 0 0 0] | [100 0 0 100] … the other 9 rows … legs muscles [1 0 0 0 1] | [100 0 0 100] … the other 9 rows … EE 194/Bio 196 Joel Grodstein 11

What else can you do with a worm? • Mutate it class Manduca: def __init__(self, legs, muscles): … def __repr__(self): … def mutate (self): self. legs[3, 3] = 1 – self. legs[3, 3] Real mutations might be more random and perhaps more widespread • How do we use this? child = parent child. mutate() • Now we have a mutated child – What about the parent. Is it mutated too? Unfortunately, yes Objects are also too big to fit into a cup. EE 194/Bio 196 Joel Grodstein 12
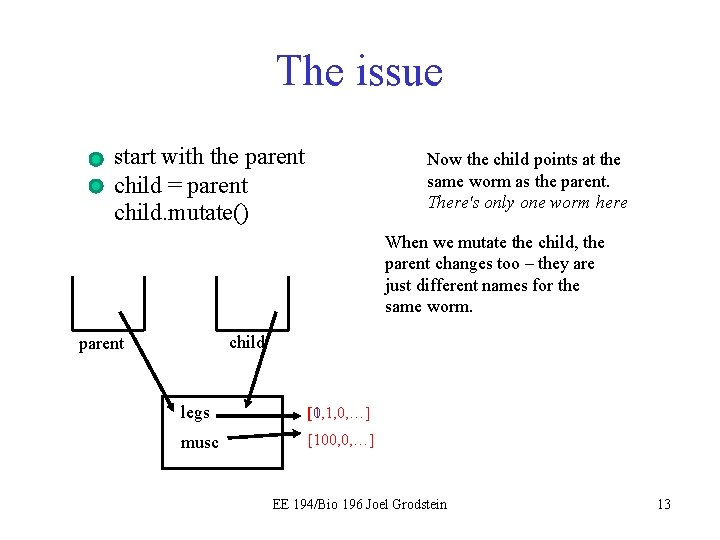
The issue start with the parent child = parent child. mutate() Now the child points at the same worm as the parent. There's only one worm here When we mutate the child, the parent changes too – they are just different names for the same worm. child parent legs [1, 1, 0, …] [0, 1, 0, …] musc [100, 0, …] EE 194/Bio 196 Joel Grodstein 13

More fun with a worm • Add a copy function: class Manduca: def __init__(self, legs, muscles, name): … def __repr__(self): … def mutate (self): … def copy (self): return (Manduca (self. legs. copy(), self. muscles. copy()) child = parent. copy() child. mutate() parent EE 194/Bio 196 Joel Grodstein child 14
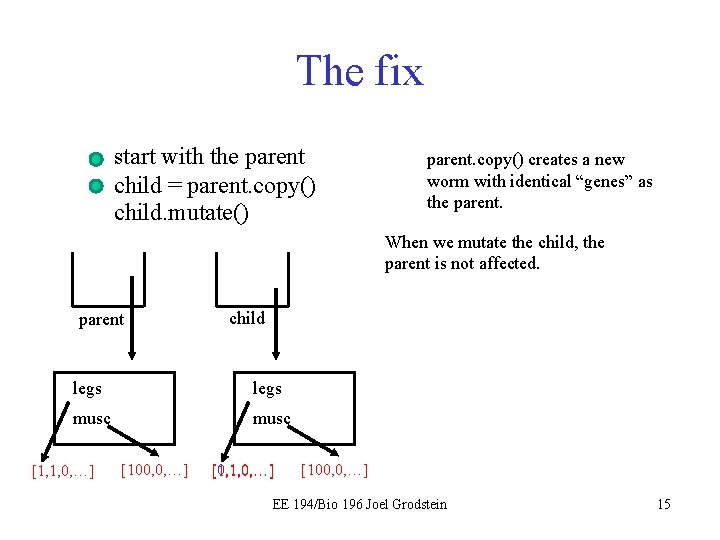
The fix start with the parent child = parent. copy() child. mutate() parent. copy() creates a new worm with identical “genes” as the parent. When we mutate the child, the parent is not affected. parent child legs musc [1, 1, 0, …] [100, 0, …] [0, 1, 0, …] [100, 0, …] EE 194/Bio 196 Joel Grodstein 15

More exercises • https: //www. w 3 resource. com/pythonexercises/class-exercises/ problems #10 and #11. • Play with the homework: – Look at the Manduca homework program manduca. Ev. py. – Make sure you understand the class it uses. – Write your own mutate() or mate() function, and print the resulting object to see if your function worked. – The __repr__ function is a bit complicated. Write your own, and see if you can get it to print differently. EE 194/Bio 196 Joel Grodstein 16
- Slides: 16