Taverna and Soap Lab Experience Elda Rossi CINECA

Taverna and Soap. Lab Experience @ Elda Rossi – CINECA (Italy)

R & Bioconductor is an open source and open development software project for the analysis and comprehension of genomic data. ¢ It is based on R , a language and environment for statistical computing and graphics. ¢

R & Bioconductor ¢ ¢ l Bio. Conductor is a collection of “packages” Two main types: 1. provides basic infrastructure support. 2. Provides innovative methodology We chose a function in the affy package (type 2. )

The affy package ¢ Package: affy Description: The package contains functions for exploratory oligonucleotide array analysis. The dependance to tk. Widgets only concerns few convenience functions. 'affy' is fully functional without it. Version: 1. 5. 8 -1 Author: Rafael A. Irizarry , Laurent Gautier , Benjamin Milo Bolstad , and Crispin Miller with contributions from … Maintainer: Rafael A. Irizarry Dependencies: R (>= 1. 9. 0), Biobase (>= 1. 4. 22), repos. Tools Suggests: tk. Widgets (>= 1. 2. 2), affydata System. Requirements: None License: LGPL version 2 or newer URL: None available ¢ Function: Expresso. From raw probe intensities to expression values

The expresso function Expression measures The most common operation is certainly to convert probe level data to expression values. 1. 2. 3. 4. 5. reading in probe level data background correction 4 methods Normalization 7 methods probe specific background correction, e. g. subtracting MM 3 methods summarizing the probe set values into one expression measure and, in some cases, a standard error for this summary 5 methods
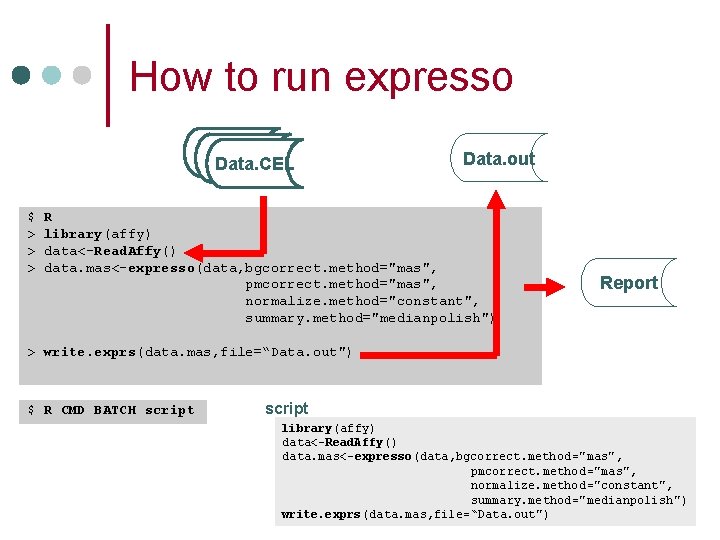
How to run expresso Data. CEL $ > > > Data. out R library(affy) data<-Read. Affy() data. mas<-expresso(data, bgcorrect. method="mas", pmcorrect. method="mas", normalize. method="constant", summary. method="medianpolish") Report > write. exprs(data. mas, file=“Data. out") $ R CMD BATCH script library(affy) data<-Read. Affy() data. mas<-expresso(data, bgcorrect. method="mas", pmcorrect. method="mas", normalize. method="constant", summary. method="medianpolish") write. exprs(data. mas, file=“Data. out")
![The files OUT file CEL file [CEL] Version=3 [HEADER] Cols=126 Rows=126 Total. X=126 Total. The files OUT file CEL file [CEL] Version=3 [HEADER] Cols=126 Rows=126 Total. X=126 Total.](http://slidetodoc.com/presentation_image_h/1a884f974b82036ec61b4b6c61963dba/image-7.jpg)
The files OUT file CEL file [CEL] Version=3 [HEADER] Cols=126 Rows=126 Total. X=126 Total. Y=126 Baseline=Not normalized Dat. Header=ctrl 150: CLS=1167 … [INTENSITY] Number. Cells=15876 Cell. Header=X Y MEAN 0 0 551. 0 10651. 0 2 0 642. 0 3 0 10855. 0 4 0 278. 0 5 0 452. 0 6 0 11139. 0 100084_at 101482_at 31962_at 32466_at 35201_at 36189_at 36678_at 37001_at 37029_at 37046_at 37189_at 37719_at 37725_at 38437_at 38730_at 39425_at 40276_at Sample 001. cel 2. 68016528652511 2. 41830136307405 12. 3667390890414 12. 4078453130306 6. 73875347104673 6. 91195864883172 10. 0269997503136 8. 7690698709579 7. 58176898579828 4. 7250160934765 7. 08125646141077 5. 33679629782696 7. 634367429284 7. 54693596951725 7. 61959398527742 6. 07663839694708 6. 33983152588017 Sample 002. cel 2. 75619854567269 2. 19230548692681 12. 4534076075796 12. 5262787728982 6. 36824635919863 6. 77835938949316 9. 76893096184106 8. 57322443505215 7. 24297853600119 4. 7250160934765 7. 0999566997911 5. 33679629782696 7. 41050271151406 7. 16216316289552 7. 65907193898742 6. 03298499862286 6. 21300599988174 Sample 003. cel 3. 82550383255225 3. 4173900695363 12. 8658623516881 13. 2129784659009 7. 53465018481639 7. 94585515997792 11. 1443619988943 9. 80956768540462 8. 67002397585278 5. 68254863921313 7. 92512679504857 6. 39140386282694 8. 85664197069339 8. 3816810916508 9. 00657184492387 7. 14769809957403 6. 85968858773872

Setting up Soap. Lab A linux based server was chosen ¢ Tomcat was installed ¢ Java was upgraded ¢ Axis was installed ¢ Soap. Lab was installed ¢ Up to here: No Problems !!! Vega. cineca. it Tomcat 5. 0. 28 Java 1. 4 Axis 1. 1 Soap. Lab precompiled for Suse Linux

Defining the Application 1. 2. 3. 4. 5. Write the application wrapper Write the ACD file for the application Convert ACD to XML Start up the Soap. Lab server Deploy the new service

1. Write the application wrapper /biotools/services/affy-expresso. pl # R code to run analysis #!/usr/bin/perl use Getopt: : Long; open(AFFY, ">$datadir/affy"); print AFFY <<EOF ; library(affy) data<-Read. Affy() data. mas<-expresso(data, bgcorrect. method="$bgcorrect", pmcorrect. method="mas", normalize. method="$normalize", summary. method="medianpolish") write. exprs(data. mas, file ="data. txt") EOF close(AFFY); # command arguments (with default) Get. Options("bgcorrect=s"=>$bgcorrect, "normalize=s"=>$normalize); $bgcorrect="mas" if $bgcorrect eq ""; $normalize="constant" if $normalize eq ""; # location of R executable $rexe="/biotools/R/R-2. 1. 0/bin/R"; # data directory $datadir=“/biotools/services/data"; # now run program system "cd $datadir; $rexe CMD BATCH affy"; # print output open(OUT, "$datadir/data. txt "); while (<OUT>) {print $_; } close(OUT);

2. Write the ACD file /biotools/soapbin/analysis-interfaces/metadata/affy. acd appl: bioconductor [ documentation: "affy/expresso function of Bio. Conductor" version: "1. 0" groups: "Microarrays" nonemboss: "Y" executable: affy-expresso. pl The path is defined in the shell ] string: bgcorrect [ additional: "Y" Input 1: Background correction parameter: "Y" default: "mas" ] string: normalize [ additional: "Y" Input 1: Normalization method parameter: "Y" default: "constant" ] outfile: output [ additional: "Y" Output: standard output default: “stdout"

3, 4, 5: Final steps 3. Convert ACD to XML /biotools/soapbin/analysis-interfaces/generator/acd 2 xml From: To: 4. . . /metadata/affy. acd. . /metadata/microarrays/affy-al. xml Start up the Soap. Lab server /biotools/soapbin/analysis-interfaces/run-App. Lab-server 5. Deploy the new service /biotools/soapbin/analysis-interfaces/ws/deploy-web-services How to shut down the server?
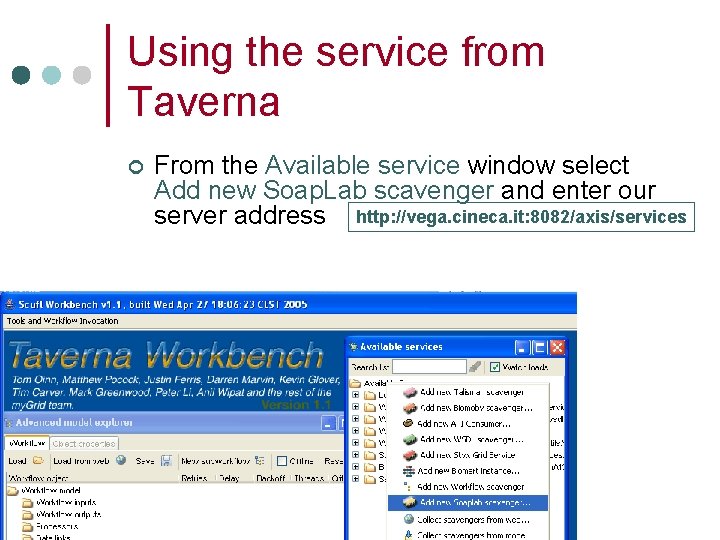
Using the service from Taverna ¢ From the Available service window select Add new Soap. Lab scavenger and enter our server address http: //vega. cineca. it: 8082/axis/services

Using the service … (2) ¢ ¢ ¢ The new processor appears in the microarrays folder you can find the affy service After connecting input & output ports, the service can be launched
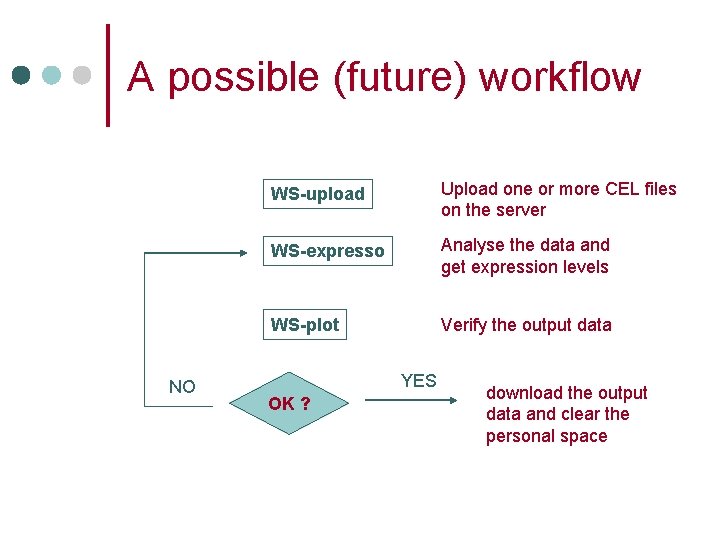
A possible (future) workflow NO WS-upload Upload one or more CEL files on the server WS-expresso Analyse the data and get expression levels WS-plot Verify the output data YES OK ? download the output data and clear the personal space
- Slides: 15