Review of HW J 3 Aaron Bloomfield CS

Review of HW J 3 Aaron Bloomfield CS 101 -E

Strand representation Strand - nucleotide. Sequence = + public. Strand() slice(int i 1, int i 2) + public … Strand(String s)s) + public Object clone() + public String to. String() + public void splice(Strand other. Strand) + public. Strandslice(int i 1, intint i 2) + publicint inthas(Strand s)s) + public. Strandsubstrand(int i 1) + public. Strandsubstrand(int i 1, int i 2) String - “tgccagt” text = “tgccagt” - length = 7 + length (): int + char. At (int i): char + sub. String (int m, int n): String + index. Of (String s, int m): int +. . .

splice (…) void splice (Strand other. Strand) “ 0123456789” leading trailing this “ 012345” “ 6789” Strand - nucleotide. Sequence = other. Strand + public void splice (Strand ( other. Strand)) +… + public void splice (Strand other. Strand) +… public void splice(Strand other. Strand) { // determine the concatenation components String leading = this. nucleotide. Sequence ; nucleotide. Sequence; String trailing = other. Strand. nucleotide. Sequence; // update strand to be a concatenation this. nucleotide. Sequence = leading + trailing; } Not required! Not an member variable!!!

public Strand () this Strand - nucleotide. Sequence = + public Strand () +… // Strand(): default constructor public Strand() { this. nucleotide. Sequence = ""; } “”

public Strand (String s) this Strand - nucleotide. Sequence = desired. Sequence + public Strand ((String desired. Sequence) ) +… “ 012345” // Strand(): specific constructor public Strand(String desired. Sequence) { this. nucleotide. Sequence = desired. Sequence; }
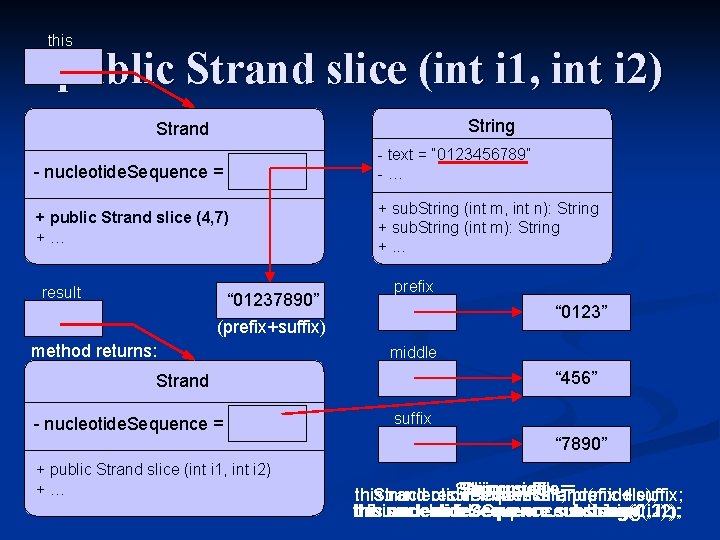
this public Strand slice (int i 1, int i 2) String Strand - nucleotide. Sequence = - text = “ 0123456789” -“ 01237890” … + public Strandslice(int (4, 7) i 1, int i 2) +… + sub. String (int m, int n): String + sub. String (int m): String +. . . result “ 01237890” prefix “ 0123” (prefix+suffix) method returns: middle “ 456” Strand - nucleotide. Sequence = + public Strand slice (int i 1, int i 2) +… suffix “ 7890” String middle prefix suffix this. nucleotide. Sequence Strand result return = new result; Strand(middle); ===prefix + suffix; this. nucleotide. Sequence. substring(i 1, this. nucleotide. Sequence. substring(0, this. nucleotide. Sequence. substring(i 2); i 1); i 2);

public Strand slice (int i 1, int i 2) // slice(): extract portion of the Strand with indices i 1 - i 2 -1 public Strand slice(int i 1, int i 2) { // decompose the nucleotide sequence into three parts String prefix = this. nucleotide. Sequence. substring(0, i 1); String middle = this. nucleotide. Sequence. substring(i 1, i 2); String suffix = this. nucleotide. Sequence. substring(i 2); // update strand to be concatenation of prefix and suffix this. nucleotide. Sequence = prefix + suffix; // return the extracted strand Strand result = new Strand(middle); return result; }
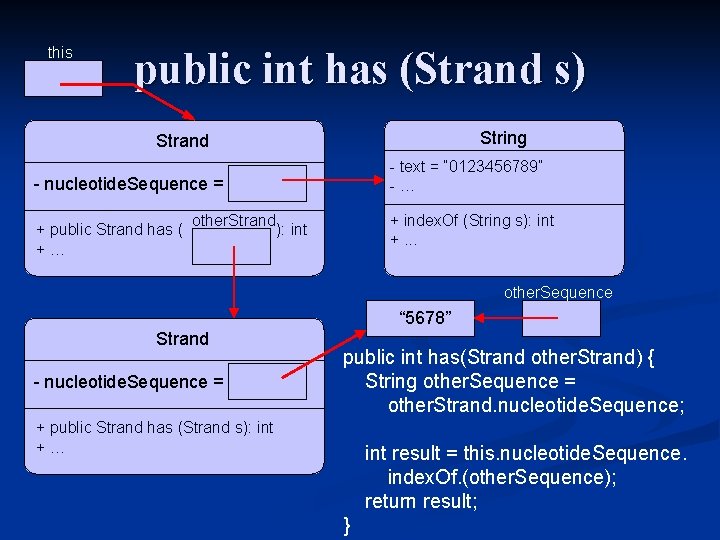
this public int has (Strand s) String Strand - nucleotide. Sequence = - text = “ 0123456789” -… other. Strand + public Strand has ( other. Strand): ): int +… + index. Of (String s): int +. . . other. Sequence “ 5678” Strand - nucleotide. Sequence = public int has(Strand other. Strand) { String other. Sequence = other. Strand. nucleotide. Sequence; + public Strand has (Strand s): int +… int result = this. nucleotide. Sequence. index. Of. (other. Sequence); return result; }

public Strand substrand (int i 1, int i 2) this String Strand - nucleotide. Sequence = + public. Strandsubstrand (int(2, 6) i 1, int i 2) +… - text = “ 0123456789” -… + substring (int i 1, int i 2): String +. . . result middle “ 2345” Strand - nucleotide. Sequence = + public Strand substrand (int i 1, int i 2) +… public Strand substrand (int i 1, int i 2) { String middle = this. nucleotide. Sequence. substring(i 1, i 2); Strand result = new Strand (middle); return result; }

public Strand substrand (int i 1) public Strand substrand (int i) { return this. substrand (i, i + 1); }
- Slides: 10