KDDCup 2004 Chairs Rich Caruana Thorsten Joachims Web

KDD-Cup 2004 Chairs: Rich Caruana & Thorsten Joachims Web Master++: Lars Backstrom Cornell University

KDD-Cup Tasks Goal: Optimize learning for different performance metrics Task 1: Particle Physics – – Accuracy Cross-Entropy ROC Area SLAC Q-Score Task 2: – – Protein Matching Squared Error Average Precision Top 1 Rank of Last

Competition Participation Timeline – April 28: tasks and datasets available – July 14: submission of predictions Participation – – – 500+ registrants/downloads 102 teams submitted predictions Physics: 65 submissions Protein: 59 submissions Both: 22 groups Demographics – Registrations from 49 Countries (including. com) – Winners from China, Germany, India, New Zealand, USA – Winners half from companies, half from universities

Task 1: Particle Physics Data contributed by Charles Young et al, SLAC (Stanford Linear Accelerator) Binary classification: distinguishing B from B-Bar particles Balanced: 50 -50 B/B-Bar 78 features (most real-valued) describing track Some missing values Train: 50, 000 cases Test: 100, 000 cases

Task 1: Particle Physics Metrics 4 performance metrics: – – Accuracy: had to specify threshold Cross-Entropy: probabilistic predictions ROC Area: only ordering is important SLAC Q-Score: domain-specific performance metric from SLAC Participants submit separate predictions for each metric – About half of participants submitted different predictions for different tasks – Winner submitted four sets of predictions, one for each task Calculate performance using PERF software we provided to participants
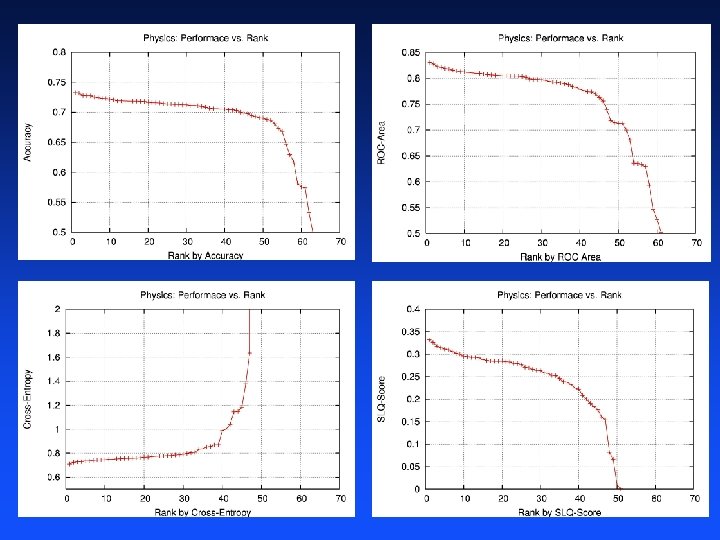

Determining the Winners For each performance metric – Calculate performance using same PERF software available to participants – Rank participants by performance – Honorable mention for participant ranked first Overall winner is participant with best average rank across all metrics

and the winners are…

Task 1: Physics Winners Rank Accuracy Cross Entropy ROC Area SLQ Score Average Rank 1 st 0. 7326 0. 7095 0. 8305 0. 3328 1. 25 2 nd 0. 7319 0. 7246 0. 8275 0. 3265 1. 75 3 rd 0. 7278 0. 7280 0. 8225 0. 3175 3. 25 Christophe Lambert (Golden Helix Inc. ): 3 rd place overall (out of 65) Lalit Wangikar et al. (Inductis Inc. ): 2 nd place overall, HM Acc David Vogels et al. (MEDai Inc. /University of Central Florida): 1 st place overall, HM ROC, HM Cross-Entropy, HM SLQ

Bootstrap Analysis of Results How much does selection of winner depend on specific test set (100 k)? Algorithm: – Repeat many times: Ç Take 100 k bootstrap sample (with replacement) from test set Ç Evaluate performance on bootstrap sample and re-rank participants – What is probability of winning/placing?
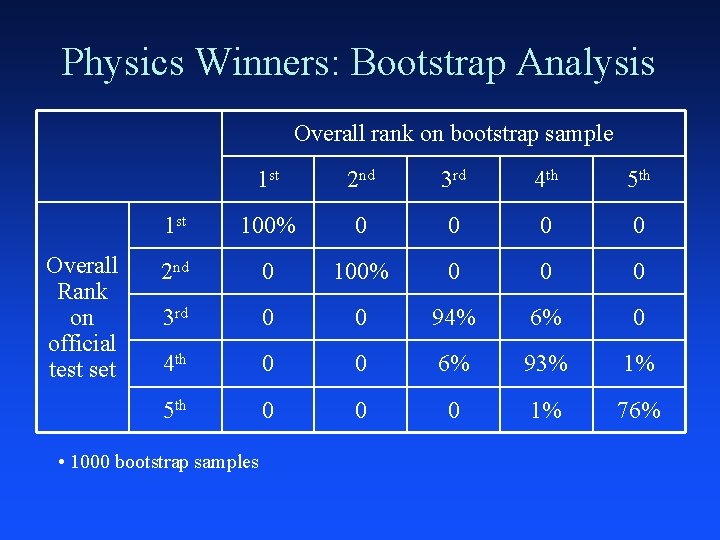
Physics Winners: Bootstrap Analysis Overall rank on bootstrap sample Overall Rank on official test set 1 st 2 nd 3 rd 4 th 5 th 1 st 100% 0 0 2 nd 0 100% 0 0 0 3 rd 0 0 94% 6% 0 4 th 0 0 6% 93% 1% 5 th 0 0 0 1% 76% • 1000 bootstrap samples

MEDai Talk

Task 2: Protein Matching Data contributed by Ron Elber, Cornell University Finding homologous proteins (structural similarity) 74 real-valued features describing match between two proteins Data comes in blocks Pr 1 Pr 2 … Pr N – + … – Unbalanced: typically < 10 homologs (+) per – – … – block of 1000 + – … + Train: 153 Proteins (145, 751 cases) – + … – Test: 150 Proteins (139, 658 cases) … … – – … –

Task 2: Protein Matching Metrics Four performance metrics: – Mean Squared Error: probabilistic predictions – Mean Average Precision: only ordering within each block is important – Mean Top 1: best predicted match is true homolog in each block – Mean Rank of Last: finding all homologs Again metric participants submitted separate predictions for each – Again, about half of participants submitted multiple sets of predictions – 19/20 top participants submitted multiple sets of predictions – Optimizing to each metric separately helped more on Protein than on Physics
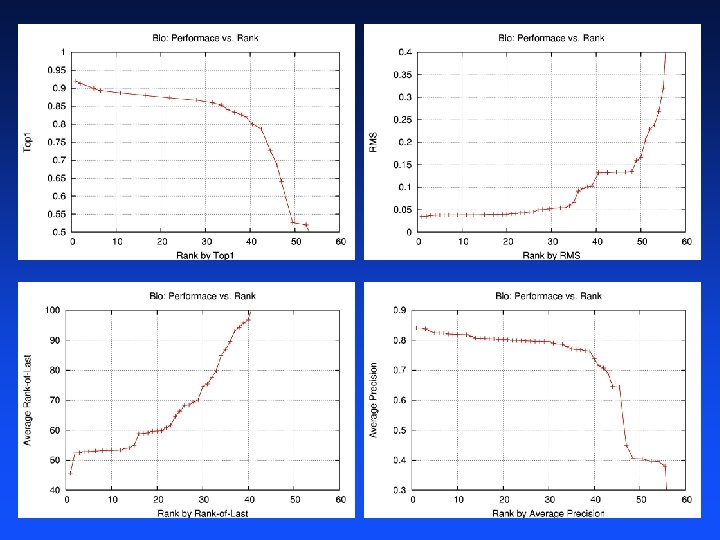
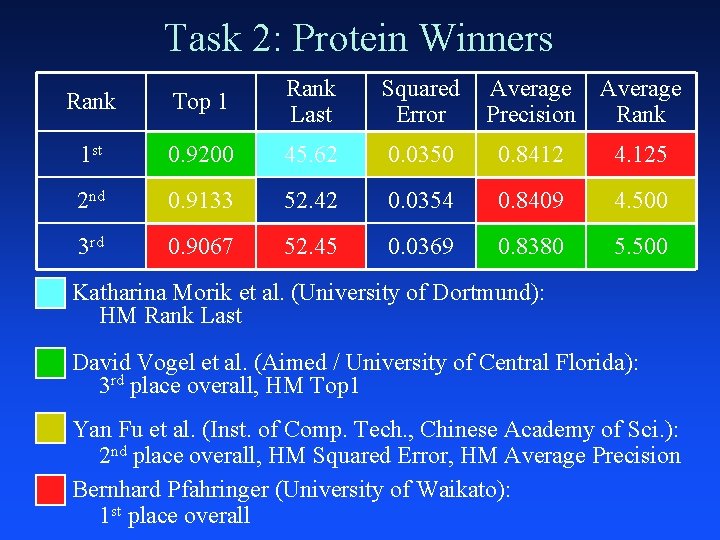
Task 2: Protein Winners Rank Top 1 Rank Last Squared Error Average Precision Average Rank 1 st 0. 9200 45. 62 0. 0350 0. 8412 4. 125 2 nd 0. 9133 52. 42 0. 0354 0. 8409 4. 500 3 rd 0. 9067 52. 45 0. 0369 0. 8380 5. 500 Katharina Morik et al. (University of Dortmund): HM Rank Last David Vogel et al. (Aimed / University of Central Florida): 3 rd place overall, HM Top 1 Yan Fu et al. (Inst. of Comp. Tech. , Chinese Academy of Sci. ): 2 nd place overall, HM Squared Error, HM Average Precision Bernhard Pfahringer (University of Waikato): 1 st place overall
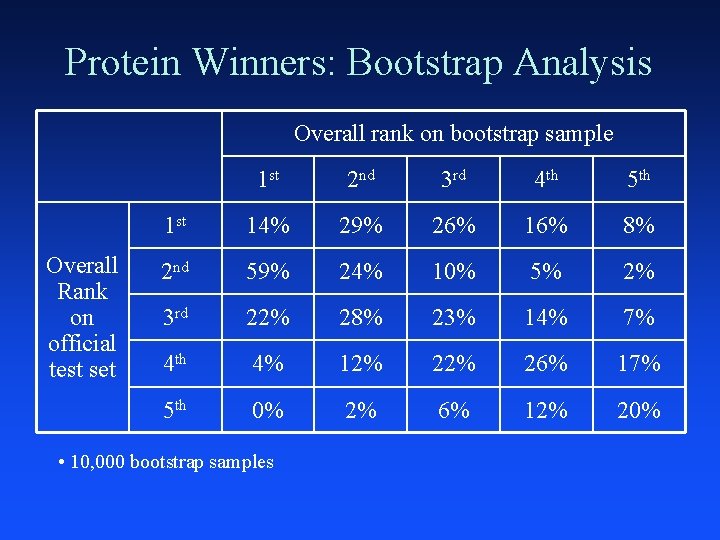
Protein Winners: Bootstrap Analysis Overall rank on bootstrap sample Overall Rank on official test set 1 st 2 nd 3 rd 4 th 5 th 1 st 14% 29% 26% 16% 8% 2 nd 59% 24% 10% 5% 2% 3 rd 22% 28% 23% 14% 7% 4 th 4% 12% 26% 17% 5 th 0% 2% 6% 12% 20% • 10, 000 bootstrap samples

Talk by Bernhard Pfahringer
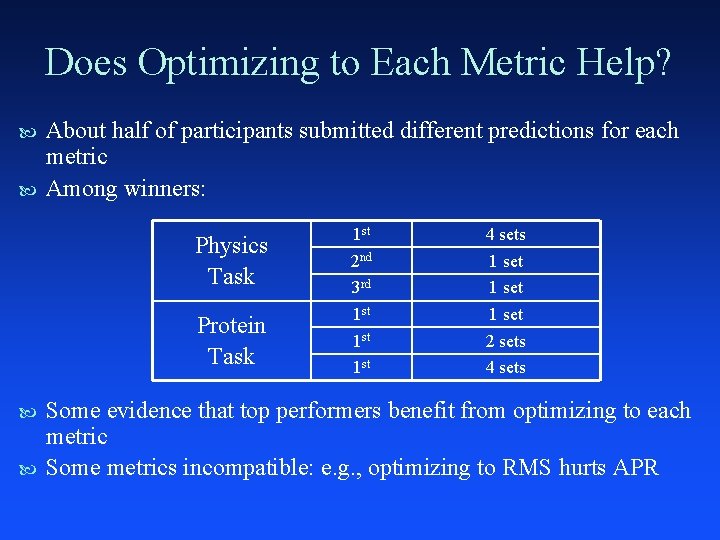
Does Optimizing to Each Metric Help? About half of participants submitted different predictions for each metric Among winners: Physics Task Protein Task 1 st 2 nd 3 rd 1 st 1 st 4 sets 1 set 2 sets 4 sets Some evidence that top performers benefit from optimizing to each metric Some metrics incompatible: e. g. , optimizing to RMS hurts APR
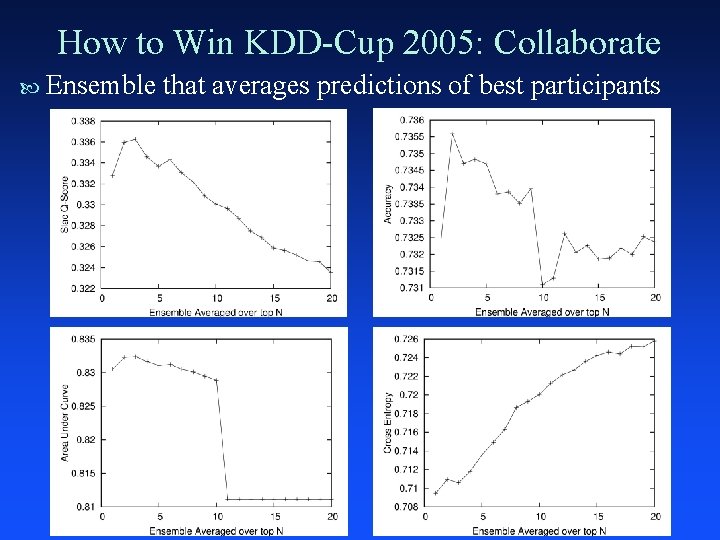
How to Win KDD-Cup 2005: Collaborate Ensemble that averages predictions of best participants
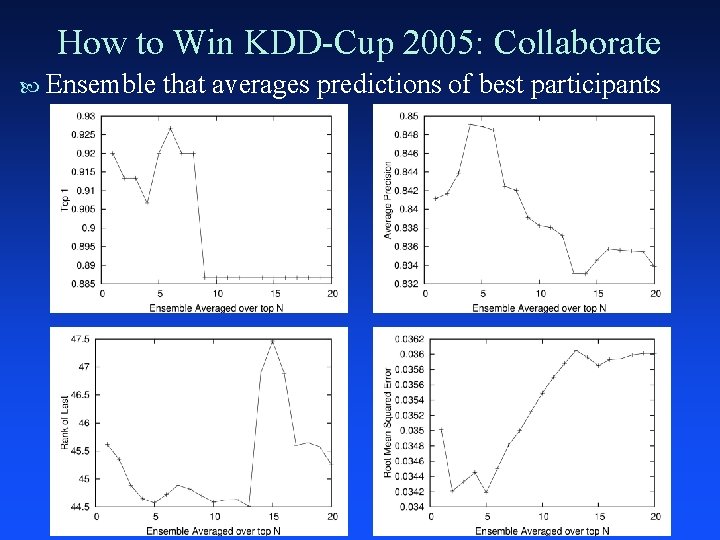
How to Win KDD-Cup 2005: Collaborate Ensemble that averages predictions of best participants

Closing Data and all results available online: http: //kodiak. cs. cornell. edu/kddcup PERF software download: http: //www. cs. cornell. edu/~caruana Thanks to: – – Web Master++: Lars Backstrom (Cornell) Physics Data: Charles Young (SLAC) Protein Data: Ron Elber (Cornell) PERF: Alex Niculescu (Cornell), Filip Radlinski (Cornell), Claire Cardie (Cornell), … Thanks to participants who found bugs in the PERF software: – Chinese Academy of Sciences – University of Dortmund And of course, thanks to everyone who participated!
- Slides: 22