GLOBAL BIODIVERSITY INFORMATION FACILITY Challenges operating a global

GLOBAL BIODIVERSITY INFORMATION FACILITY Challenges operating a global biodiversity Portal Tim Robertson Information Systems Architect September 2010 www. gbif. org

About GBIF • • An operational network Connecting hundreds of institutions Thousands of data sources Free and open access to information • Achieved through globally recognised standards – Not “standards”, “interoperability” (Dr. Michael J. Ackerman)

The Data Portal • Provides services – Search (real time) – Browse (taxonomic, geographic, by publisher etc) – Pre-processed reports – Visualisations – Various export capabilities – Means to access the original source of data • An index of content available through the GBIF Network Status: Live since 2007 http: //data. gbif. org
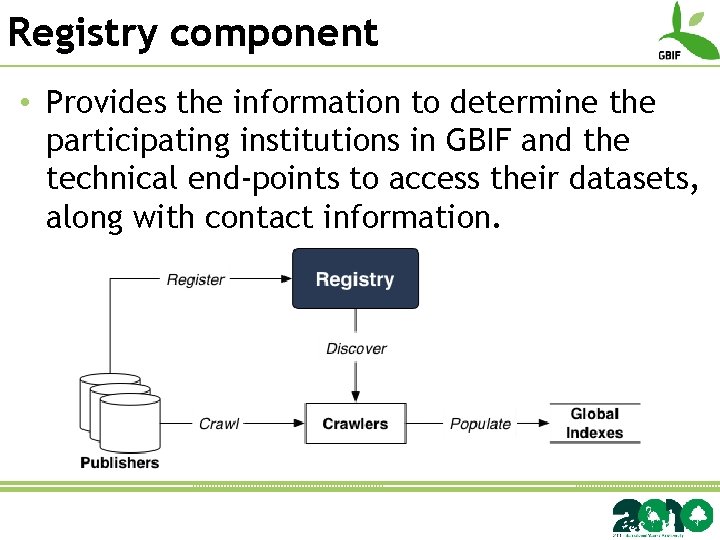
Registry component • Provides the information to determine the participating institutions in GBIF and the technical end-points to access their datasets, along with contact information.

Registry component • Previously implemented using an open industry business registry known as UDDI – 2 -tier model of “data publisher having several datasets” • The GBIF network is more complicated than this. – Datasets are shared or published through multiple channels. Results in complex attribution chains.

Registry component • Developed a graph based model to handle this information • Challenge is now to open the management of content – Wikipedia style open access curation? – Facebook style request / confirmation? – Complex rules for editing permissions? Status: Prototype available http: //gbrds. gbif. org
Registry component http: //gbrds. gbif. org Status: Prototype available http: //gbrds. gbif. org

Metadata catalogue • Data portal currently provides – Contact information – Basic attribution • Limited means to – To understand the nature of dataset creation – Difficult to assess fitness-for-use of data – To discover undigitised content or content in non standard forms Status: Under construction

Metadata catalogue • Recent work focusing on: – Accommodate existing metadata standards (ISO, FGDC, EML, DIF, DC, NCD etc) • Limit use of “lossy” transformations – Support OAI-PMH protocols for harvesting – Provide OAI-PMH services for wider participation – Developing a GBIF metadata profile • Based on the EML 2. 1. 0 profile • http: //rs. gbif. org/schema/eml-gbif-profile/ – Prototype basic and structured search Status: Under construction

Checklist. Bank • Unified access to multiple checklists – Taxonomic, nomenclatural, thematic • Provide dictionaries to help improve services for parsing and name finding • Name based services – Treatment of names – Classification services for names – Vernacular names – Identifiers used by sources of checklists • E. g. Catalogue of Life LSIDs • Lexical and nomenclatural grouping Status: Prototype available http: //ecat-dev. gbif. org
Checklist. Bank http: //ecat-dev. gbif. org Status: Prototype available http: //ecat-dev. gbif. org

Annotating content • Correcting mistakes • Aligning to standardised vocabularies • Completing missing terms (e. g. reverse georeferencing) • Complementing with additional information • Invasive indicator • Protected area identifier • etc. • Lesson learnt: Calculate once, store along with record Annotation Interest Group

Annotating content • Not all annotations are of interest to the data holder • Are all annotations from trustworthy source? • Challenge is to design an infrastructure that supports – Widespread quality control – Brokerage of annotations for reuse • Investigate open access to help foster innovation and research Annotation Interest Group

Performance • An index should be: – Fast in operation – Relevant • Provide means to search that suit the users – Accurate • Reflect changes in the network quickly • “Changes made by the data holder should be reflected in index within 1 month”

Performance • Transfer stage: – No robust mechanism to follow changes • dwc: date. Last. Modified not often usable – TAPIR / Di. GIR / Bio. CASe • Inefficient transfer for full dataset harvesting • No mechanism to inform of deletion of records • Need to do a complete dataset harvest each time • Darwin Core text guidelines one means for simplifying this – 1 month saw a 13 million record increase over usual 1 -2 million due to Darwin. Core archives

Challenge: performance • Post-harvesting stage: 215, 000 records refreshed in a month 7, 000 per day 300, 000 per hour (24/7 is a challenge in itself!) 5000 per minute 83 per second (with no growth…) • Clearly parallelisation is key… … and database becomes a bottleneck

Challenge: consistency • The more “batch” processing one does, the higher the risk of inconsistencies • Aim for eventual consistency? • Can be mitigated – through careful data process planning – through clear explanations to users of when a “view” was last produced

Roadmap 2011 • Consolidate existing work – Unified data entry to (Data API) • Institutions, collections, occurrences, names… • Rich metadata where available • Multiple indexes to the content – Marine, botany, invasive etc – Service offerings (Service API) • • Registration services Name services Mapping services Annotation services
- Slides: 18