MEGAN analysis of metagenomic data Daniel H Huson











- Slides: 17

MEGAN analysis of metagenomic data Daniel H. Huson, Alexander F. Auch, Ji Qi, et al. Genome Res. 2007

Early metagenomic � Known phylogenetic markers and subsequent sequencing of clones � Analysis of paired-end reads � Complete sequences of environmental fosmid and BAC clones Rough annotation of the metabolic capacity � Environmental assemblies Distinguish between discrete species and population of closely related biotypes � Problem of using proven phylogenetic markers(ribosomal genes, coding sequences) � Slow-evolving genes : distinguishing between species at large evolutionary distances

What is MEGAN? � Metagenome Analyzer (MEGAN) � Free software. � Deviates from the analytical pattern of previous � Built on the statistical analysis of comparing random sequence intervals with unspecified phylogenetic properties against databases � Depends on the related sequences in the databases � Providing filter to adjust the level of stringency later to an appropriate level � Laptop analysis � Comparing � Graphical result (BLAST)-> laptop (MEGAN) and statistical output

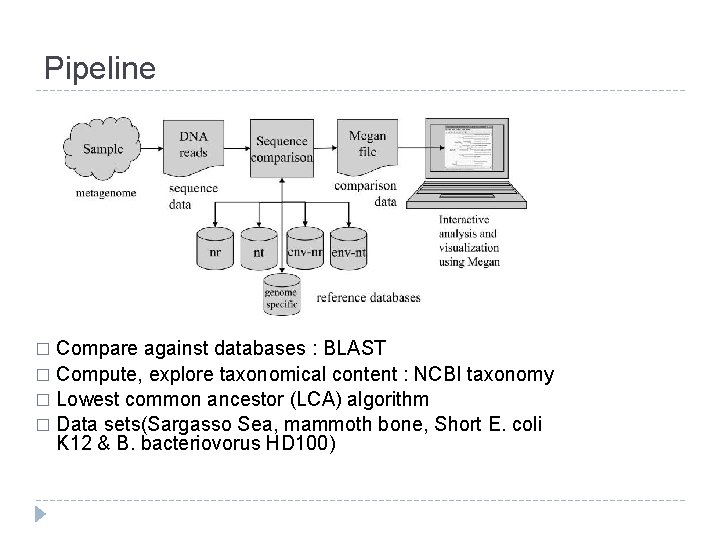
Pipeline Compare against databases : BLAST � Compute, explore taxonomical content : NCBI taxonomy � Lowest common ancestor (LCA) algorithm � Data sets(Sargasso Sea, mammoth bone, Short E. coli K 12 & B. bacteriovorus HD 100) �

What we can do with MEGAN � Species and strain identification through species-specific genes � Searching species or taxa by find tool � Distribution of strains of a species � Underlying sequence alignments

Experiments-1 � Sargasso � Sea data set � Sanger sequencing � Sample 1 -4 from DDBJ/EMBL/Gen. Bank 10000 reads from Sample 1 Randomly selected a pooled set of 10000 reads from samples 2 -4 � BLASTX->NCBI-NR � 1% no hits from sample 1, <3% no hits from sample 2 -4 � Filters � Min-score : bit-score threshold of 100 � Top-percent : bit scores lie within 5% of the best score � Min-support : isolated assignments it by one read) discarded

Analysis-Sargasso Sea data � 1. 66 M reads, AVG. 818 bp by Sanger sequensing � Species profile of 16 taxonomical groups � Environmental � By assemblies analyzing six specific phylogenetic markers r. RNA, Rec. A/Rad. A, HSP 70, Rpo. B, EF-Tu, and Ef-G

Result • Sample 1 • ~83% reads were assigned to taxa that were more speific than the kingdom level • Majority of (8298) were assigned to bacterial group • Sample 2 -4 • ~59% reads were assigned to taxa that were more specific than the kingdom level • Majority of (5709) were assigned to bacterial group • Alphaproteobacteria, Gammaproteobacteria by a factor of 2 -4 over the remaining 14 taxonomic groups • Eukaryotes & Viruses : size filtering • Archaea : May be there is 10 times as much vacterial sequence information in the public databases

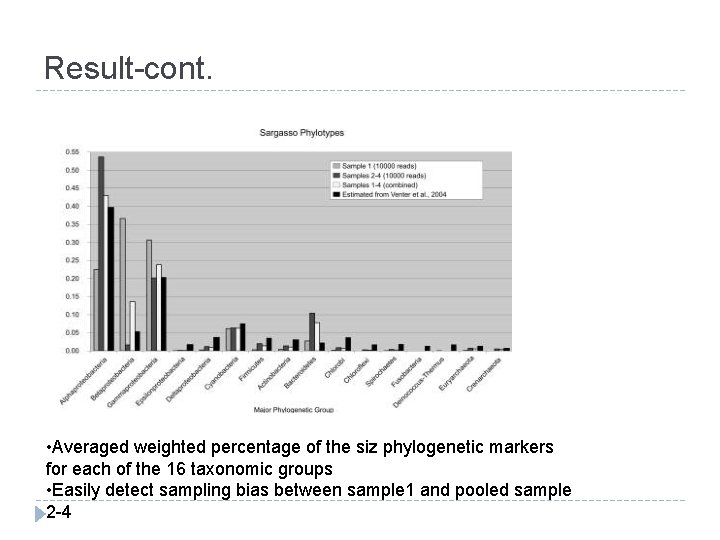
Result-cont. • Averaged weighted percentage of the siz phylogenetic markers for each of the 16 taxonomic groups • Easily detect sampling bias between sample 1 and pooled sample 2 -4

Experiments-2 � Mammoth � Data bone set � Roche GS 20 sequencing (Sequencing-by-synthesis) � Sample from 1 g of mammoth bone , 28000 years � ~300, 000 reads, 95 bp � BLASTZ-genome sequences (elephant, human, dog) � 45. 4% of the reads mammoth DNA, others are environmental organisms (bacteria, fungi, amoeba, nematodes) � BLASTX–NCBI-NR for environmental sequences � Filters : bit-score threshold 30, discard isolated assignment (filtered 2086 reads)

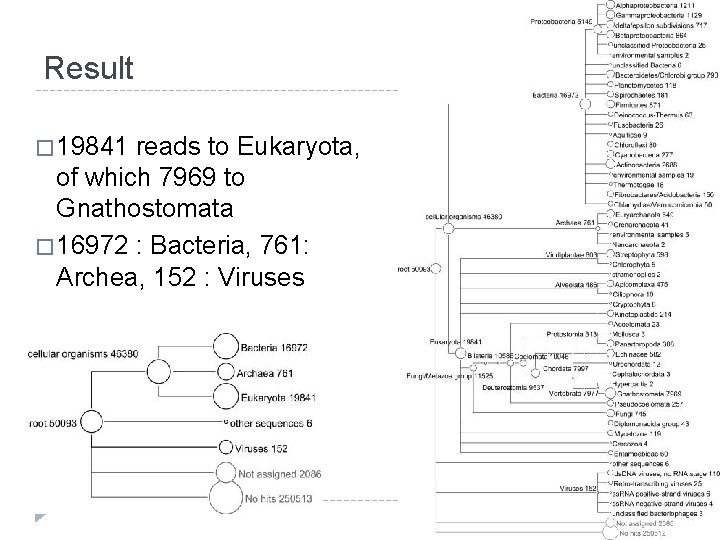
Result � 19841 reads to Eukaryota, of which 7969 to Gnathostomata � 16972 : Bacteria, 761: Archea, 152 : Viruses

Experiment 3 � Identifying � Short species from various lead length E. coli K 12 & B. bacteriovorus HD 100 simulation � 5000 random shotgun reads � BLASTX-NCBI-NR � Filters Bit-score threshold 35 20% of the best hit Discarded isolated assignments � Result : no false-positive assignment, short read can be used for metagenomic analysis, albeit at the cost of a high rate of underprediction

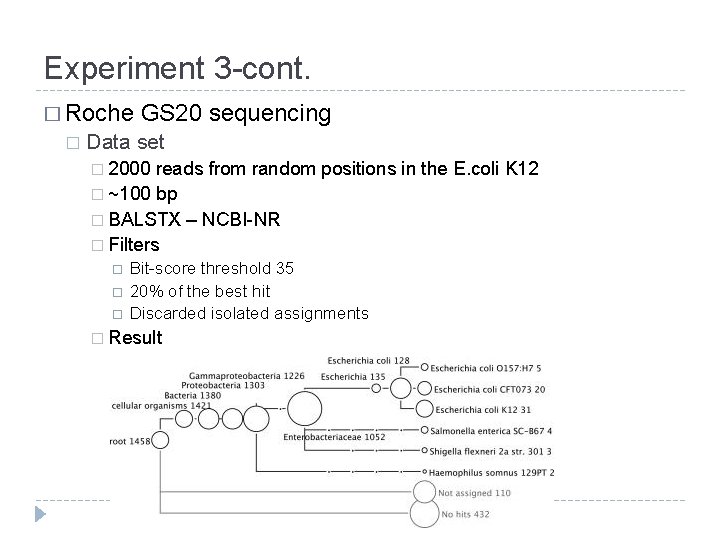
Experiment 3 -cont. � Roche � GS 20 sequencing Data set � 2000 reads from random positions in the E. coli K 12 � ~100 bp � BALSTX – NCBI-NR � Filters Bit-score threshold 35 20% of the best hit Discarded isolated assignments � Result

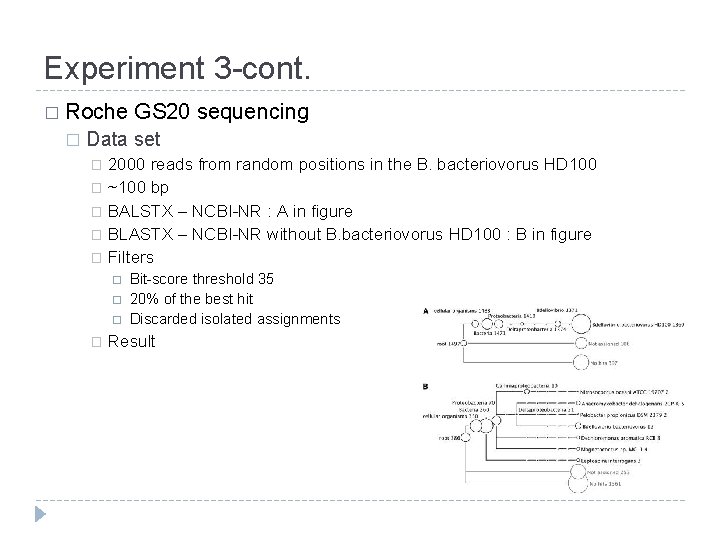
Experiment 3 -cont. � Roche � GS 20 sequencing Data set 2000 reads from random positions in the B. bacteriovorus HD 100 � ~100 bp � BALSTX – NCBI-NR : A in figure � BLASTX – NCBI-NR without B. bacteriovorus HD 100 : B in figure � Filters � � Bit-score threshold 35 20% of the best hit Discarded isolated assignments Result

MEGAN 3(June, 2009) � Suitable for very large datasets � Advances in the throughput and cost-efficiency of sequencing technology � Interests � From changed ‘which species present’ to ‘What’s different? ’ � Features � Visualization technique for multiple database � New statistical method for highlighting the difference in a pairwise comparison

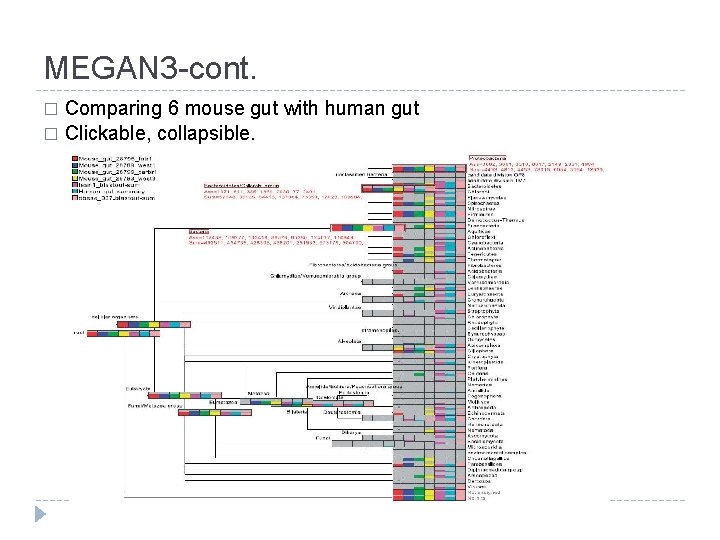
MEGAN 3 -cont. Comparing 6 mouse gut with human gut � Clickable, collapsible. �
