A Prionlike Mechanism in Amyotrophic Lateral Sclerosis Dr

- Slides: 12

A Prion-like Mechanism in Amyotrophic Lateral Sclerosis Dr. Eamonn F. Healy Professor of Chemistry St. Edward’s University Austin, Tx.

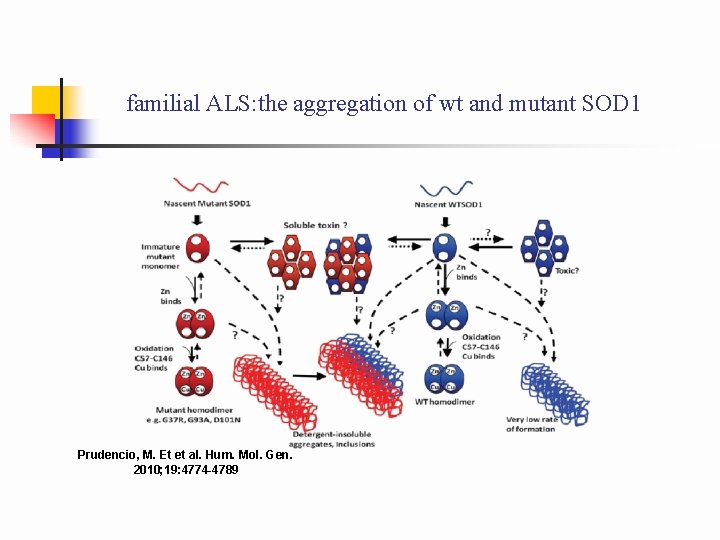
familial ALS: the aggregation of wt and mutant SOD 1 Prudencio, M. Et et al. Hum. Mol. Gen. 2010; 19: 4774 -4789

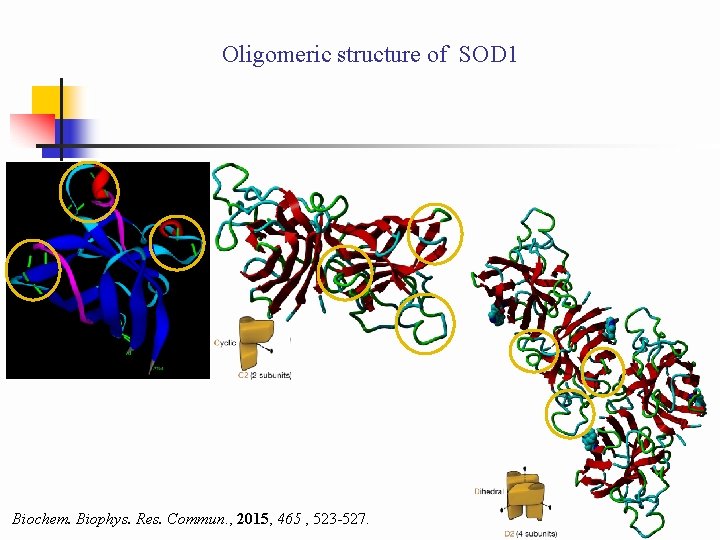
Oligomeric structure of SOD 1 Biochem. Biophys. Res. Commun. , 2015, 465 , 523 -527.

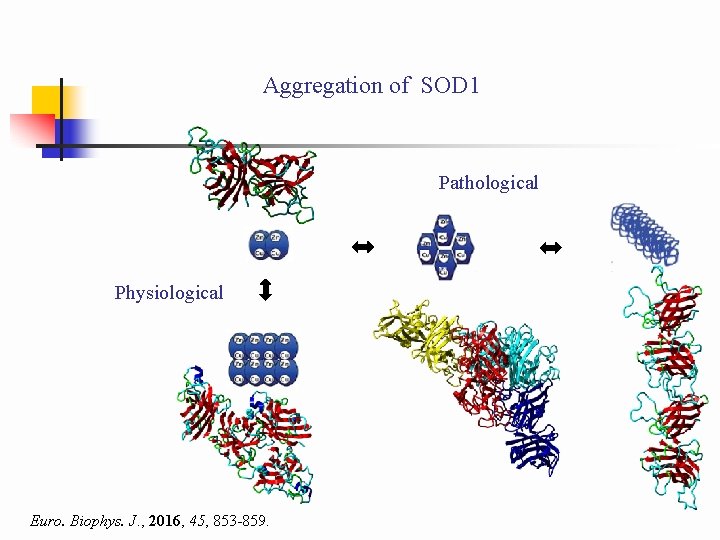
Aggregation of SOD 1 Pathological Physiological Euro. Biophys. J. , 2016, 45, 853 -859.

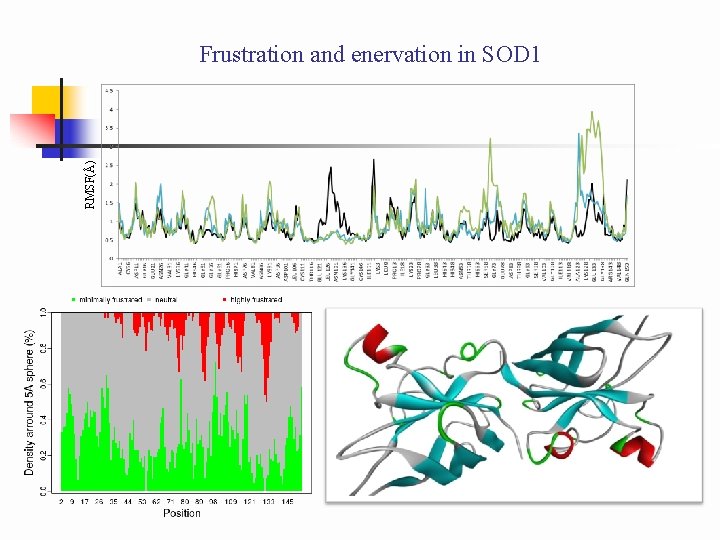
RMSF(Å) Frustration and enervation in SOD 1

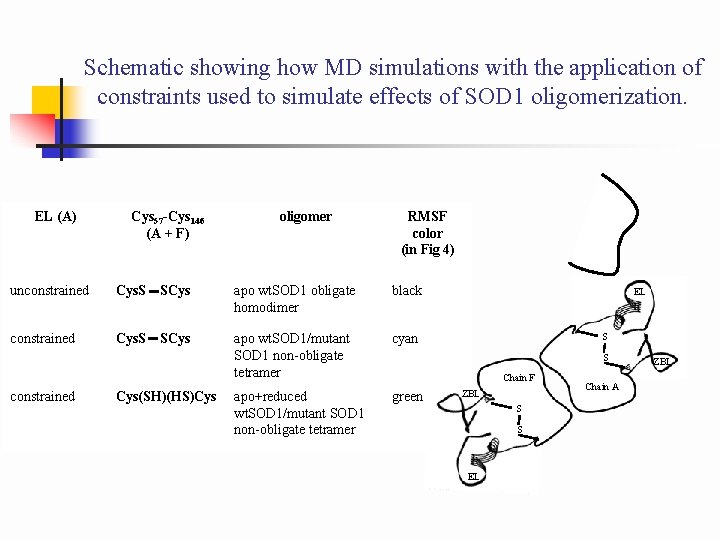
Schematic showing how MD simulations with the application of constraints used to simulate effects of SOD 1 oligomerization. EL (A) Cys 57 -Cys 146 (A + F) oligomer RMSF color (in Fig 4) unconstrained Cys. S SCys apo wt. SOD 1 obligate homodimer black constrained Cys. S SCys apo wt. SOD 1/mutant SOD 1 non-obligate tetramer cyan apo+reduced wt. SOD 1/mutant SOD 1 non-obligate tetramer green constrained Cys(SH)(HS)Cys EL S S Chain F ZBL S S EL Chain A ZBL

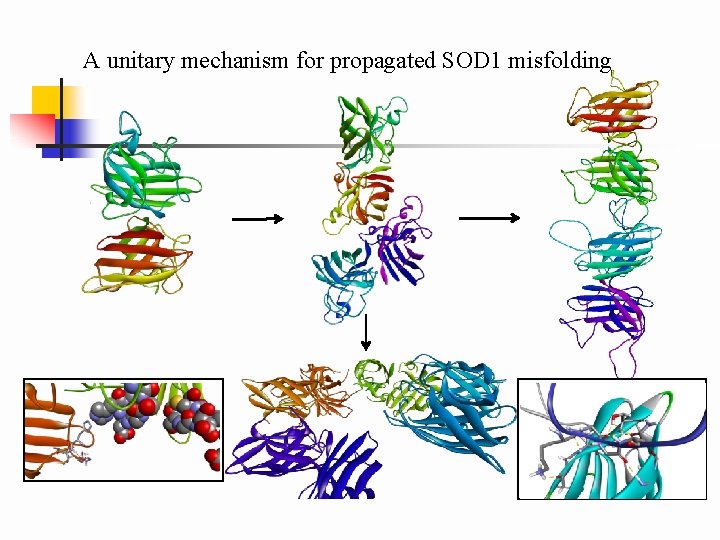
A unitary mechanism for propagated SOD 1 misfolding

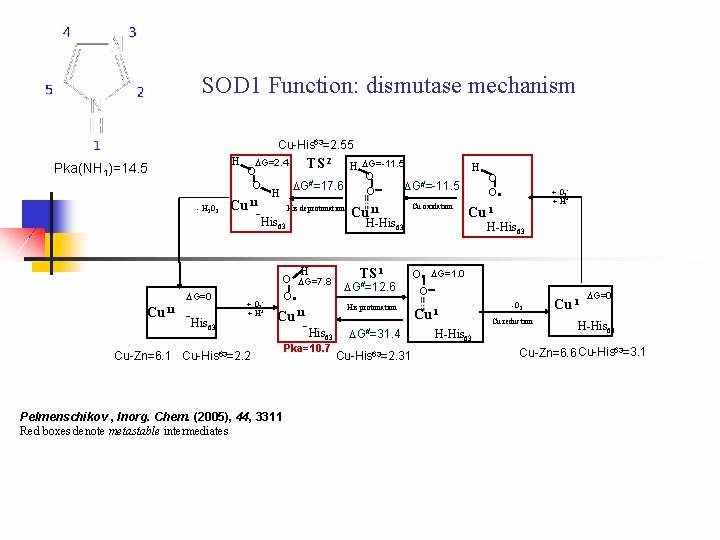
SOD 1 Function: dismutase mechanism Cu-His 63=2. 55 H Pka(NH 1)=14. 5 - H 2 O 2 DG=2. 4 O O Cu II H DG=-11. 5 O DG#=17. 6 DG#=-11. 5 O - H -His TS 2 His deprotonation Cu II - His 63 + O 2 -. + H+ H O DG=7. 8 O . Cu II -His 63 Cu-Zn=6. 1 Cu-His 63=2. 2 Pelmenschikov , Inorg. Chem. (2005), 44, 3311 Red boxes denote metastable intermediates Cu oxidation H-His 63 63 DG=0 Cu II Pka=10. 7 TS 1 DG#=12. 6 His protonation DG#=31. 4 Cu-His 63=2. 31 H O O . + O 2 -. + H+ Cu I H-His 63 . O DG=1. 0 O - Cu I H-His 63 -O 2 Cu reduction Cu I DG=0 H-His 63 Cu-Zn=6. 6 Cu-His 63=3. 1

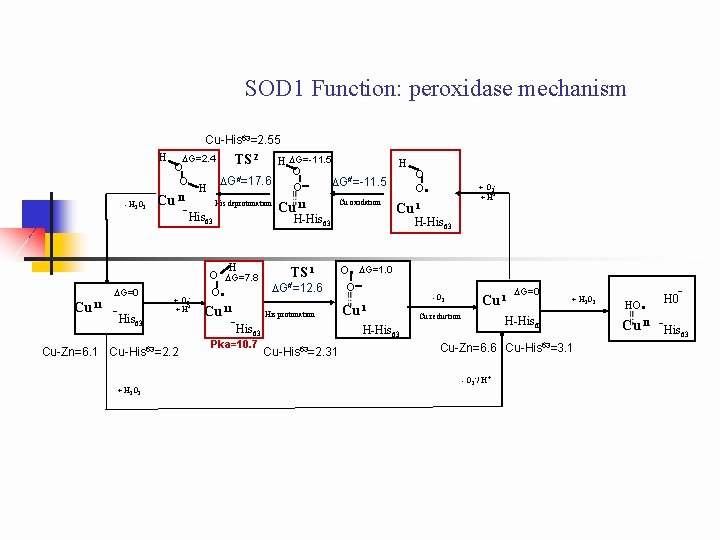
SOD 1 Function: peroxidase mechanism Cu-His 63=2. 55 H - H 2 O 2 DG=2. 4 O O Cu II H DG=-11. 5 O DG#=17. 6 DG#=-11. 5 O His deprotonation Cu II - His 63 + O 2 -. + H+ Cu II H O DG=7. 8 O . Cu II -His TS 1 DG#=12. 6 His protonation + H 2 O 2 Pka=10. 7 O O . + O 2 -. + H+ Cu I H-His 63 . O DG=1. 0 O - Cu I -O 2 Cu reduction Cu-His 63=2. 31 DG=0 + H 2 O 2 H-His 63 63 Cu-Zn=6. 1 Cu-His 63=2. 2 Cu oxidation H-His 63 63 DG=0 H -His TS 2 Cu-Zn=6. 6 Cu-His 63=3. 1 - O 2 -/ H+. HO . - H 0 Cu II -His 63

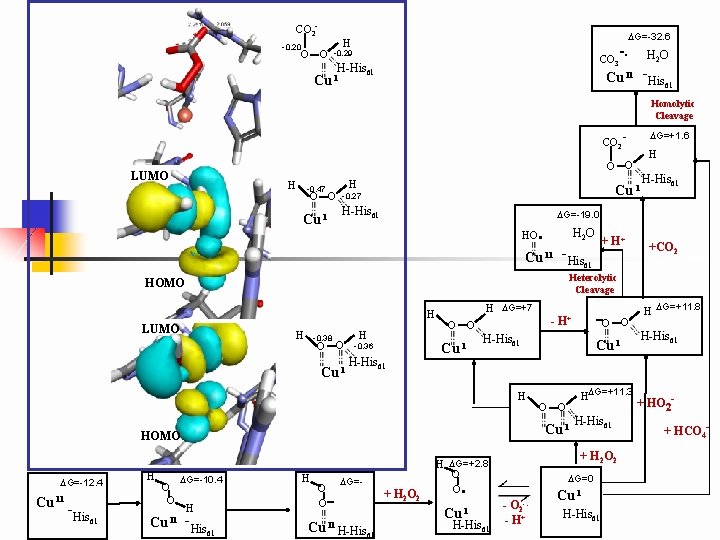
CO 2 - DG=-32. 6 H -0. 20 -. CO O O -0. 29 H 2 O 3 H-His 61 Cu II -His 61 Cu I Homolytic Cleavage - DG=+1. 6 CO 2 LUMO H H -0. 47 O O -0. 27 Cu I H-His 61 HO . H 2 O + H+ H LUMO H H -0. 38 O O -0. 36 Cu I H-His 61 H DG=+7 O O Cu I -His 61 Cu II H -His 61 H O O - DG=- Cu II H-His 61 + H 2 O 2 O O . H-His 61 + H 2 O 2 H DG=+2. 8 O O Cu I O HDG=+11. 3 Cu I DG=-10. 4 -O Cu I HOMO Cu II - H+ H-His 61 H O O +CO 2 Heterolytic Cleavage HOMO H H-His 61 DG=-19. 0 Cu II -His 61 DG=-12. 4 H O O DG=0 - O 2 - H+ -. Cu I H-His 61 H DG=+11. 8 H-His 61 + HO 2+ HCO 4 -

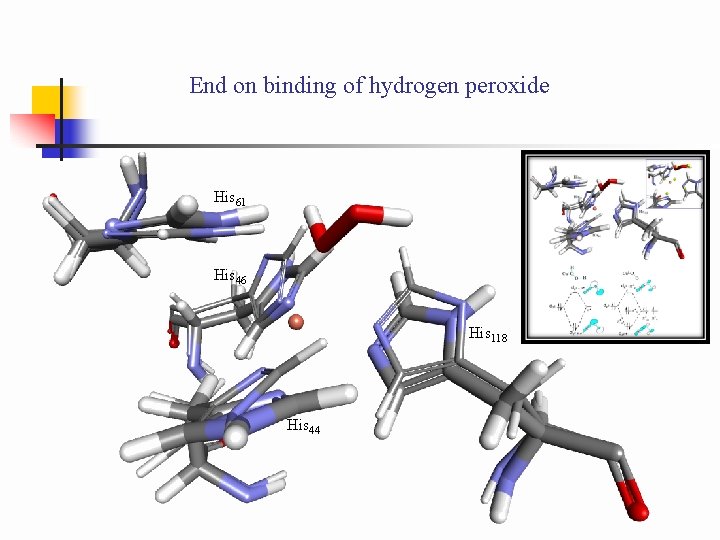
End on binding of hydrogen peroxide His 61 His 46 His 118 His 44

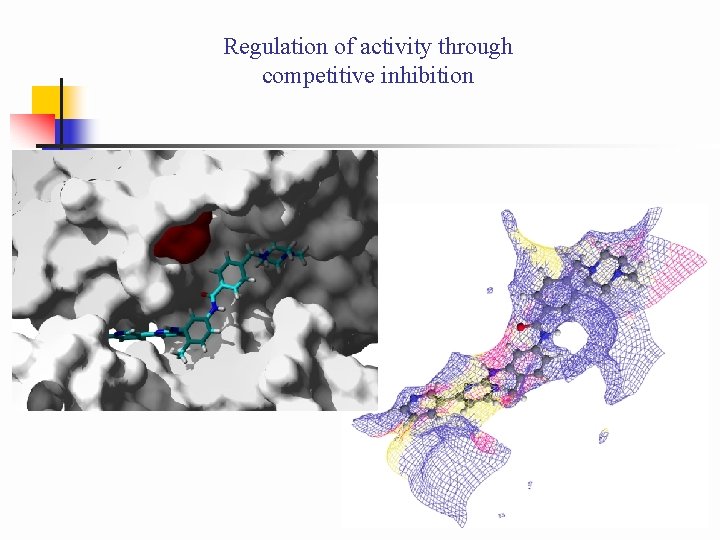
Regulation of activity through competitive inhibition