ZOO 405 Week 6 ZOO 405 by Rania

ZOO 405, Week 6 ZOO 405 by Rania Baleela is licensed under a Creative Commons Attribution. Non. Commercial-Share. Alike 3. 0 Unported License

This week • East Africa genome • Benefits of human genome project • Miscellaneous Issues & definitions

East Africa • A strategic region to study human genetic diversity, why? 1. ethnically diverse populations 2. Linguistically diverse populations 3. geographically diverse populations.

Northeast African genomic variation • Genotyping was performed on an Illumina Human Omni 5 MExome SNP-array (~3. 5 Million SNPs) • 244 unrelated individuals from 18 population Hollfelder N, Schlebusch CM, Günther T, Babiker H, Hassan HY, Jakobsson M (2017) Northeast African genomic variation shaped by the continuity of indigenous groups and Eurasian migrations. PLo. S Genet 13(8): e 1006976.
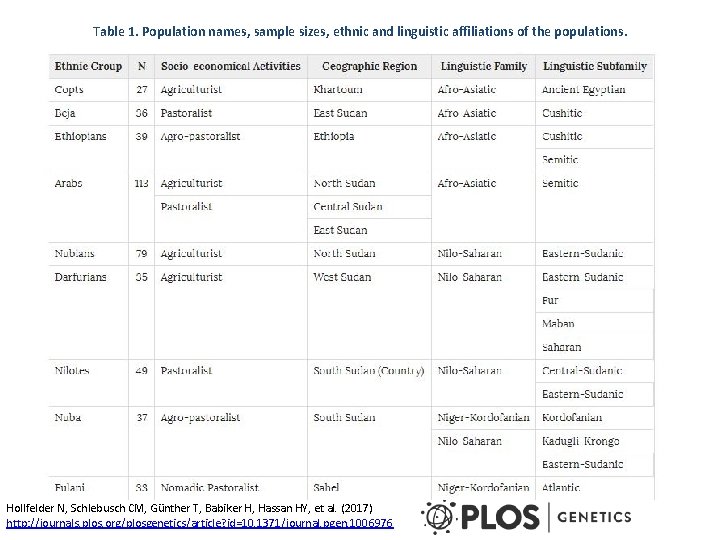
Table 1. Population names, sample sizes, ethnic and linguistic affiliations of the populations. Hollfelder N, Schlebusch CM, Günther T, Babiker H, Hassan HY, et al. (2017) http: //journals. plos. org/plosgenetics/article? id=10. 1371/journal. pgen. 1006976
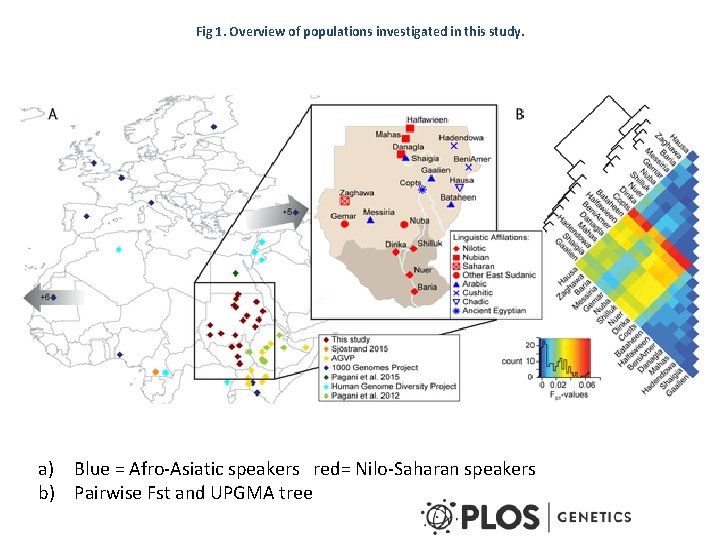
Fig 1. Overview of populations investigated in this study. a) Blue = Afro-Asiatic speakers red= Nilo-Saharan speakers b) Pairwise Fst and UPGMA tree

Findings • Genetic variation correlated with geography to a greater extent than to linguistic classification indicating that geography drives population structure in the area. • Several populations, in particular from the North and East of Sudan displayed genetic affinities to non. Africans, which is consistent with recent admixture into these groups. • This admixture unifies the Nubian, Arabic and Beja populations from the north • Admixture is almost completely absent in the western Sudanese and South Sudanese populations.

1. Nilotic groups • Baria, Nuer, Dinka and Shilluk • One of the most common source populations for other Sudanese and South Sudanese populations • Strongly endogamic => appear to have remained relatively isolated over time. • They emerged from an ancestral group of East Africa • No significant signal of gene flow with external populations for the Nuer and Baria • Indications of external gene flow from West Africa (YRI) into Dinka and TSI to Shilluk
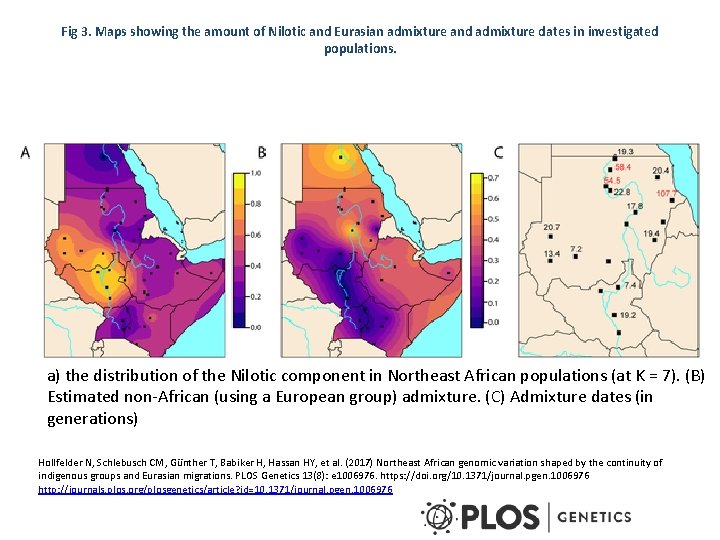
Fig 3. Maps showing the amount of Nilotic and Eurasian admixture and admixture dates in investigated populations. a) the distribution of the Nilotic component in Northeast African populations (at K = 7). (B) Estimated non-African (using a European group) admixture. (C) Admixture dates (in generations) Hollfelder N, Schlebusch CM, Günther T, Babiker H, Hassan HY, et al. (2017) Northeast African genomic variation shaped by the continuity of indigenous groups and Eurasian migrations. PLOS Genetics 13(8): e 1006976. https: //doi. org/10. 1371/journal. pgen. 1006976 http: //journals. plos. org/plosgenetics/article? id=10. 1371/journal. pgen. 1006976

2. West Africa component • NO west African or Bantu component in Sudanese and South Sudanese populations, except in the Hausa. • The Bantu migration that swept over most of sub. Saharan Africa 3– 4 thousand years ago (kya) did not cause massive admixture in northeast Africa, contrary to what has been found in many other sub-Saharan African regions, e. g. East Africa and southern Africa. • Nilotic populations and climatic conditions could have acted as a migration barrier for northeast Africa. • Although there a few Bantu speaking populations in South Sudan that likely migrated during the Bantu expansion, they do not appear to have mixed much with local Nilotic groups.

3. Nubians are an admixed group with gene-flow from outside of Africa • Eastern Sudanic languages of the Nilo-Saharan linguistic family • Little genetic differentiation among individuals and groups, • Maximum pairwise FST of 0. 004513 between the Mahas and the Halfawieen. • The FST values to the surrounding Arabic and Beja populations were also low, which hints at geneflow or shared ancestry with the neighboring populations

Even though the Nubians and the Nilotes are linguistically closer to each other than to the Afro-Asiatic groups, the Nubians showed the greatest genetic differentiation (FST between 0. 02 and 0. 04) to the Nilotes

• The strongest signal of admixture into Nubian populations came from Eurasian populations • Nubians showed the highest level of allelic richness, number of private alleles and shared private alleles (between Danagla and Halfawieen)

3. Hausa from Sudan are relatively isolated • Afro-Asiatic speaking, migrated 300 years ago to Sudan • They cluster in between the West African Yoruba and Nzime, and the Darfurian/Kordofanian and Nilotic populations. • They show some level of local Nilotic genetic material • Show Eurasian source estimated to 31. 2 ± 9. 3 generations ago (i. e. before the historically documented settlement) • It is unknown if the Hausa populations of West Africa also show Eurasian admixture signal.

Northeastern Sudanese populations especially Nubians and Beja were strongly affected by Eurasian migrations since the introduction of Islam from the Arabian Peninsula through Egypt and the Red Sea starting around 651 A. D

4. Assuming that the Nubian population is a mixture of an incoming Eurasian • 1 st contact is dated using patterns of LD-decay to roughly 56 generations ago for the Danagla and the Mahas • The Halfawieen have received Eurasian admixture later, around 19 generations ago

5. Shaigia, Gaalien and Bataheen • Eurasian migrations admixed with local populations • Populations in central Sudan that self-identify as Arab were originally a local northeast African population (similar to the Nubians and the Beja) that mixed with a Eurasian population during the Arab expansion, or possibly earlier. However, the mixed groups kept the language and culture of the incoming migrants.

6. Beja groups • Show high non-African admixture • The Beni Amer: strong admixture signal with a Eurasian population as well as a shared ancestry component with the Somali population => suggest admixture with the East African Cushitic-speaking populations (~ 59 generations ago for the Hadendowa and ~ 68– 75 generations for the Beni Amer). • Admixture of non-Africans into the Beni Amer: ~ 107. 7 ± 24. 4 generations ago and a younger event, 34. 2 generations ago => suggesting an early migration from Eurasian into these coastal African populations, possibly across the sea. • Admixture events could be driven by admixture from the Cushitic-speaking populations of the Horn of Africa,

7. Copts • Generally practicing Christianity, migrated from Egypt to Sudan around 200 years ago • They are an admixed population of at least one sub-Saharan population and one Eurasian population • Estimated to be of 69. 54% ± 2. 57 European ancestry • They have remained relatively isolated since the arrival to Sudan with only low levels of admixture with local northeastern Sudanese groups.

8. Messeiria, Gamar, Zaghawa and Nuba • The Messiria (Semitic speaking ), are nomads who inhabit a wide area in the Darfur and Kordofan regions. • They were genetically closer to other Darfurian/Kordofanian populations than to the Arab populations of central Sudan. • Clearly genetically differentiated from the Arab populations of northeastern Sudan • The Messiria showed a significant signal of admixture between Nuer and Eurasians

8. Messeiria, Gamar, Zaghawa and Nuba • The Eurasian fraction in the Messiria ~ 15% compared to the (40%-48%) in the northeastern Arabic populations. • The admixture was dated to about 7 generations ago. • Messiria = a local Kordofanian population that has acquired the language and culture from an incoming Semitic population that they mixed with some 200 years ago (190– 244 years ago). • The Gemar, a Nilo-Saharan speaking population of Darfur and Kordofan also showed signals of ~13% Eurasian admixture , dated ~ 13 generations ago • The Zaghawa and the Nuba: very little Eurasian admixture and low genetic differentiation to the Gemar and the Messiria as well as to the Nilotic populations suggesting common ancestry of Nilotic, Darfurian and Kordofanian populations.

• Northeast African Nilotes showed some distinction from an ancient Ethiopian individual (Mota, found in the Mota Cave in the southern Ethiopian highlands), which suggests population structure between northeast and eastern Africa already 4, 500 years ago. • The modern-day Nilotic groups are likely direct descendants of past populations living in northeast Africa many thousands of years ago

HGP

Benefits of Human Genome Project research • • • Improved diagnosis. Research for fuel and environmental cleanup. DNA forensics. Improved agriculture and livestock. Better understanding of evolution and human migration. More accurate risk assessment.

• Disease Genes Discovered: at least one diseaserelated mutation has been identified for 1100 genes • Clinical disorders and gene mutations: Different mutations in the same gene can give rise to more or less distinct disorders (number of diseases for which there are known mutations is ~1500) • Functional Classifications: Disease genes classed by function and their relative representations

Some diseases involve polygenic effects • A number of classic “genetic diseases” are caused by mutations of a single gene: – Huntington’s, Cystic Fibrosis, Tay-Sachs, PKU, etc. • Many diseases are the result of the interactions of many genes: – Asthma, Heart disease, Cancer Groups of SNP markers may be associated with a disease without determining mechanism

There are many socio-ethical implications of the HGP Three of the most important ones 1. Behavioral Genetics If genes that indicate susceptibility for criminality, intelligence, or homosexuality are discovered, how should we respond? 2. Gene Testing/Therapy 3. Privacy Issues

Ethical, legal & social implications of the Human Genome Project • Some questions to consider: • Fairness & privacy: who should have access to your genetic information? • Psychological stigmatization: how does knowing your predisposition to disease affect an individual? • Genetic testing: should screening be done when there is no treatment available? • Reproductive issues: use of genetic information in decision making. • Clinical issues: implementation of standards and quality control measures in testing procedures.

MISCELLANEOUS ISSUES & DEFINITIONS

Complementation analyses allow us to determine whether a certain trait is determined by just one gene or by more than one gene. e. g. inherited deafness in humans. If the 2 mutants show complementation we can conclude that the mutations in the parents are in different genes.

Chromosome walking • A method used to move systematically along a chromosome from a known location. • Is used to find adjacent genes or parts of a gene which are missing in the original clone as well as to analyse long stretches of eukaryotic DNA. • It is necessary to use DNA probes of singlecopy sequences, otherwise if the probe used is a repeated sequence, then several unrelated recombinants could be identified.

Lampbrush chromosomes (LBCs) Giant (polytene) chromosomes • LBCs are found inside amphibian and bird oocytes (immature egg cells) • Is thought to reflect the underlying organization of chromosomes • Result of continuous replication of DNA without nuclear division.
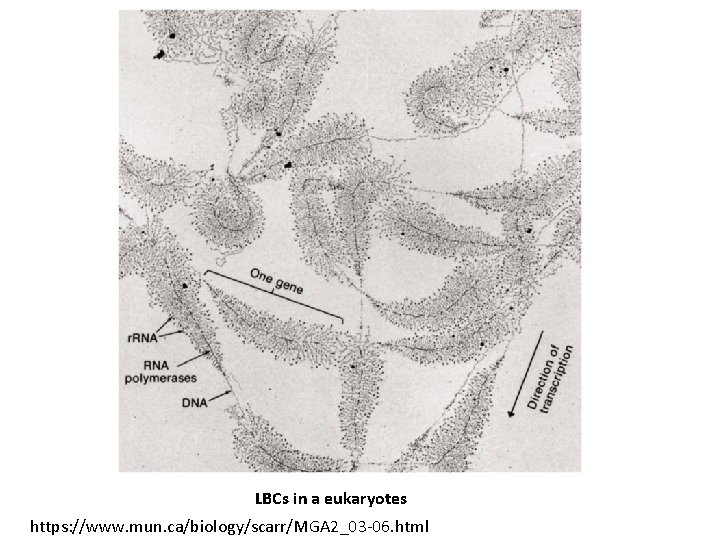
LBCs in a eukaryotes https: //www. mun. ca/biology/scarr/MGA 2_03 -06. html
- Slides: 33