ZOO 405 Week 2 ZOO 405 by Rania

ZOO 405, Week 2 ZOO 405 by Rania Baleela is licensed under a Creative Commons Attribution. Non. Commercial-Share. Alike 3. 0 Unported License

This week • • • Genome content Eukaryotic genome constitution Viruses Morphological types of viruses Retroviruses and their genome organization Retroviruses classification

Genome content

Size measurements in the molecular world • 1 mm (millimeter) = 1/1, 000 meter • 1 mm (“micron”) = 1/1, 000 of a meter (1 x 10 -6) • 1 nm (nanometer) = 1 x 10 -9 meter • 1 bp (base pair) = 1 nt (nucleotide pair) • 1, 000 bp = 1 kb (kilobase) • 1 million bp = 1 Mb (megabase) • 5 billion bp DNA ~ 1 meter • 5 thousand bp DNA ~ 1. 2 mm

The C-value enigma/paradox

“Although genes are made of DNA, much DNA is not genes” Doolittle, 1989
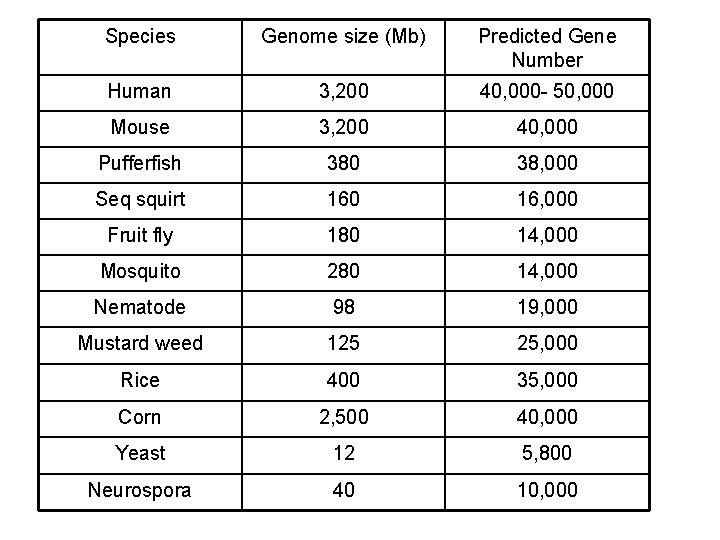
Species Genome size (Mb) Predicted Gene Number Human 3, 200 40, 000 - 50, 000 Mouse 3, 200 40, 000 Pufferfish 380 38, 000 Seq squirt 160 16, 000 Fruit fly 180 14, 000 Mosquito 280 14, 000 Nematode 98 19, 000 Mustard weed 125 25, 000 Rice 400 35, 000 Corn 2, 500 40, 000 Yeast 12 5, 800 Neurospora 40 10, 000

The C-value paradox Complexity does not correlate with genome size Dr Richard Horton 3. 4 109 bp Homo sapiens 1. 5 1010 bp Allium cepa 6. 8 1011 bp Amoeba dubia

Genome size changes Increase: (1) global increases (i. e. the entire genome or a major part of it is duplicated), (2) regional increases (i. e. a particular sequence is multiplied to generate repetitive DNA). Decreases: Loss of 1 chromosome (Aneuploidy).

Mechanisms for global genome size increase 1. Polyploidization = the addition of one or more complete sets of chromosomes to the original set. 2. Repetitive sequences: Ø Ribosomal RNA genes Ø Centromeres Ø Telomeres Ø TEs.

Transposable Elements and genome size • Variation in gene numbers cannot explain variation in genome size among eukaryotes • Most of variation in genome size is due to variation in the amount of repetitive DNA (mostly derived from TEs) • TEs accumulate in intergenic regions
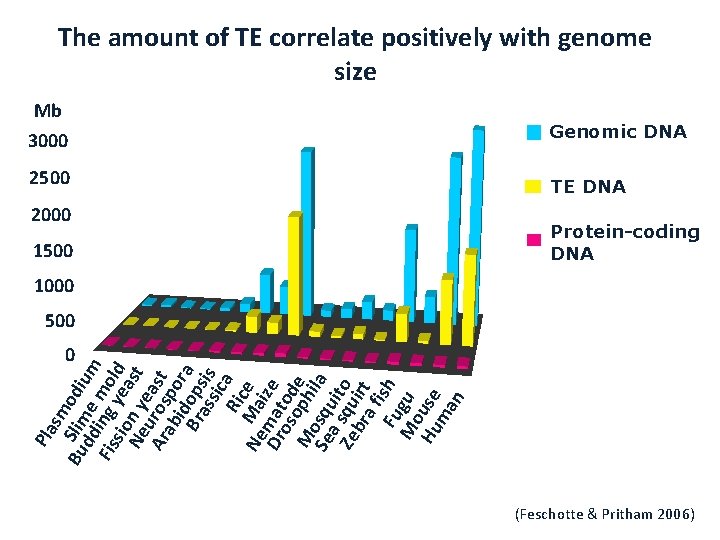
Pla sm Bu Slim odi dd e um Fis ing mol sio yea d Ne n y st e Ar uros ast ab po i r Br dops a as is sic Ric a Ne Ma e Dr mat ize os od M oph e Se osqu ila a s it Ze qu o br irt a f Fu ish M gu o Hu use ma n The amount of TE correlate positively with genome size Mb 3000 2500 2000 1500 Genomic DNA TE DNA Protein-coding DNA 1000 500 0 (Feschotte & Pritham 2006)
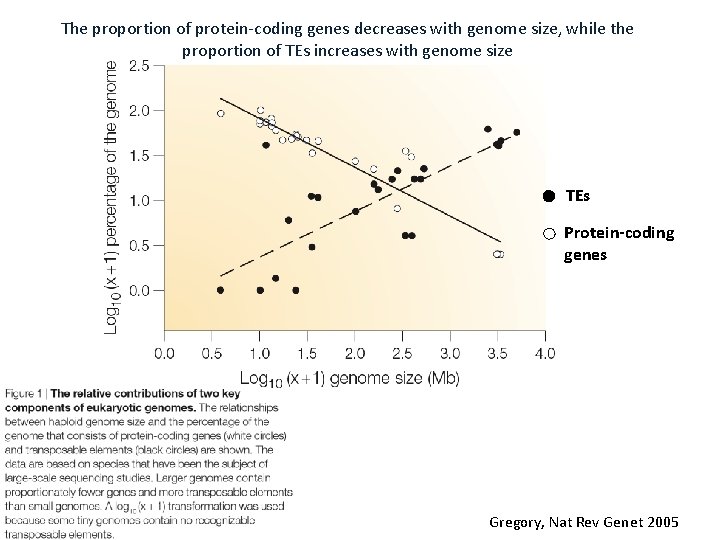
The proportion of protein-coding genes decreases with genome size, while the proportion of TEs increases with genome size TEs Protein-coding genes Gregory, Nat Rev Genet 2005

Contrasted Genome Landscapes Transposable Element
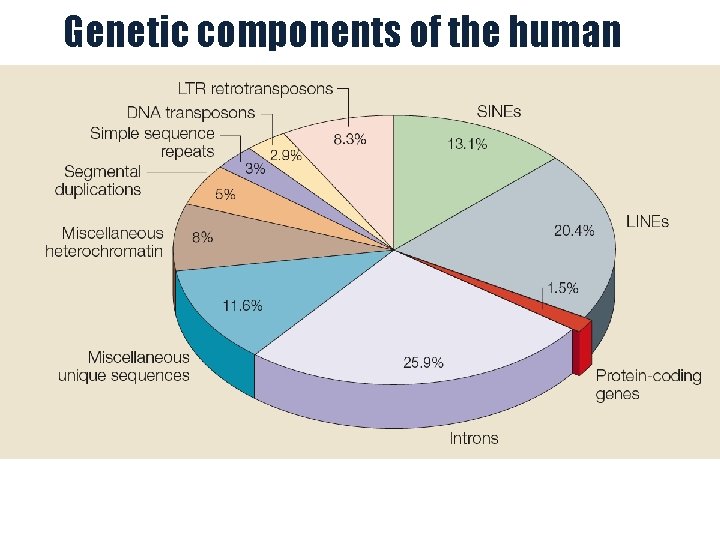
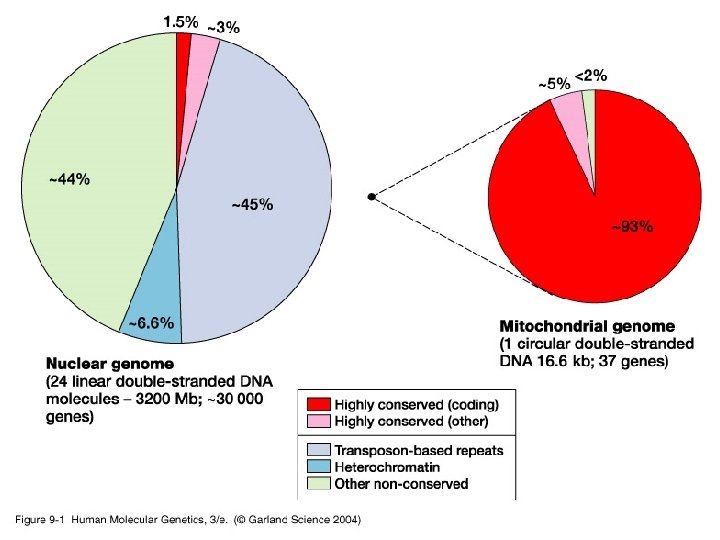
Genetic components of the human genome
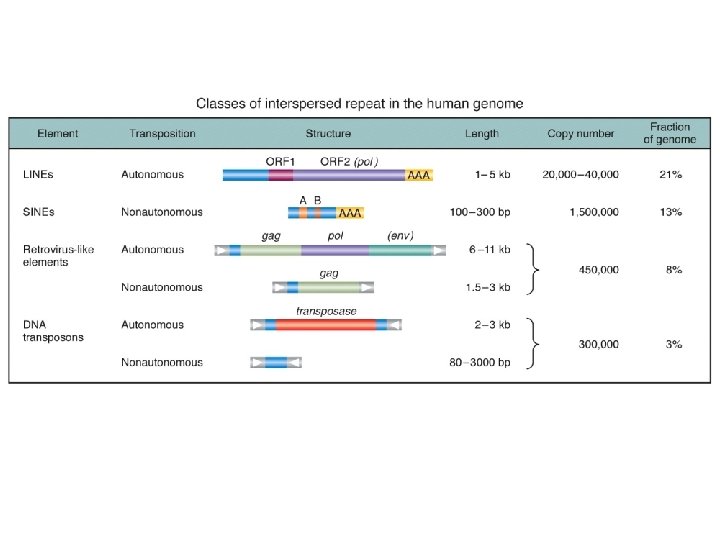

Noncoding DNA the end of the paradox

• Today, C-value differences are no longer paradoxical. • In spite of its label, the “paradox” was not the lack of a correlation with complexity, per se, but rather the inability of early researchers to reconcile the constancy of DNA content within species (which occurs because it is the stuff of genes) with the variation in quantity of DNA among species (which does not relate to the number of genes).

Excess transposition may provoke rapid changes in genome size e. g. grass genomes

Long Terminal Repeat (LTR) retrotransposons • Abundant and can impact gene and genome evolution. • Most are large elements (0. 4 kb) and are most often found in heterochromatic (gene poor) regions. • The smallest LTR retrotransposon = 292 bp (Gao et al. , 2012): • In rice, maize, sorghum and other grass genomes (indicates presence in the grass ancestor at least 50– 80 MYA). It may still be active in some genomes • The small LTR retrotransposons (SMARTs) => distributed throughout the genomes and are often located within or near genes=> can in a few instances alter both gene structures and gene expression.
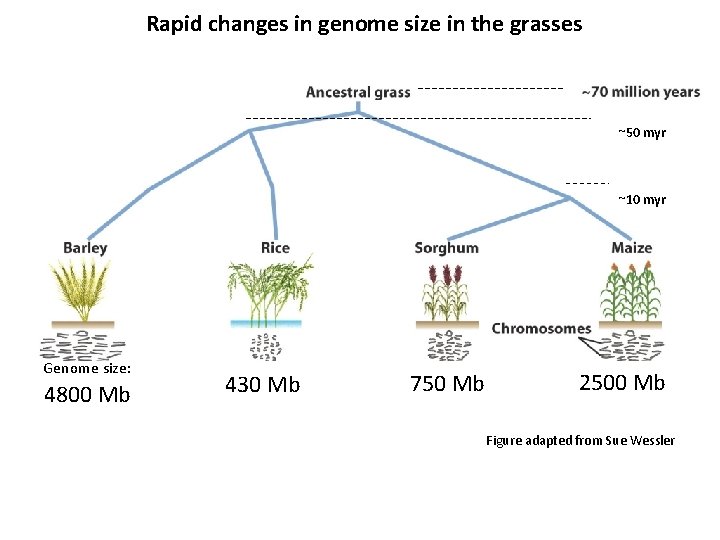
Rapid changes in genome size in the grasses ~50 myr ~10 myr Genome size: 4800 Mb 430 Mb 750 Mb 2500 Mb Figure adapted from Sue Wessler
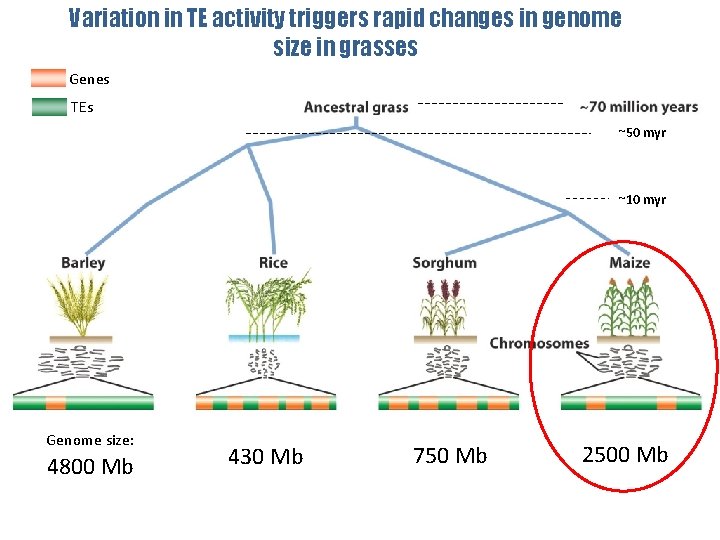
Variation in TE activity triggers rapid changes in genome size in grasses Genes TEs ~50 myr ~10 myr Genome size: 4800 Mb 430 Mb 750 Mb 2500 Mb
Retrotransposon amplification has resulted in the doubling of the maize genome in the last ~6 myr (San Miguel et al. 1998)
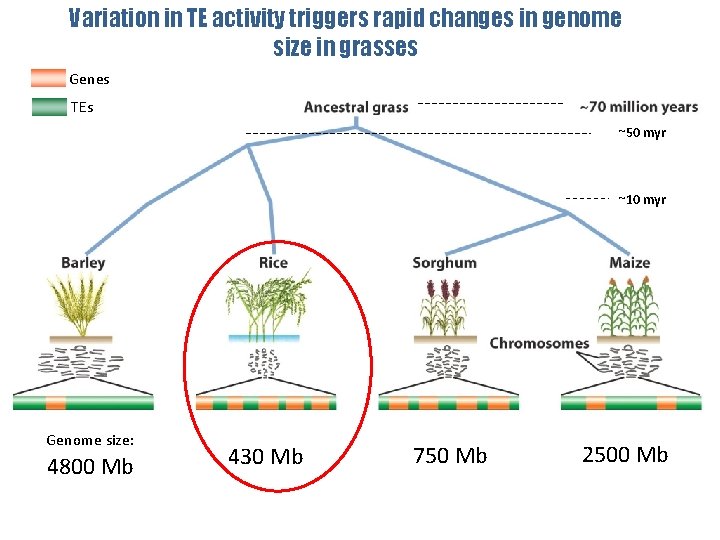
Variation in TE activity triggers rapid changes in genome size in grasses Genes TEs ~50 myr ~10 myr Genome size: 4800 Mb 430 Mb 750 Mb 2500 Mb
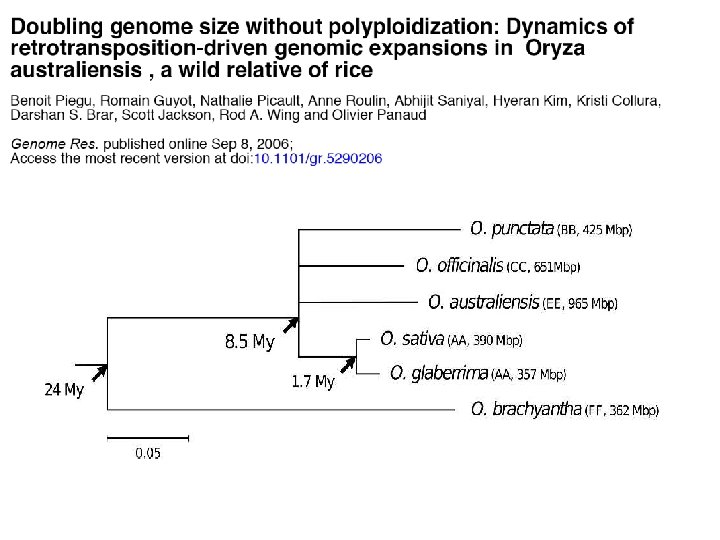
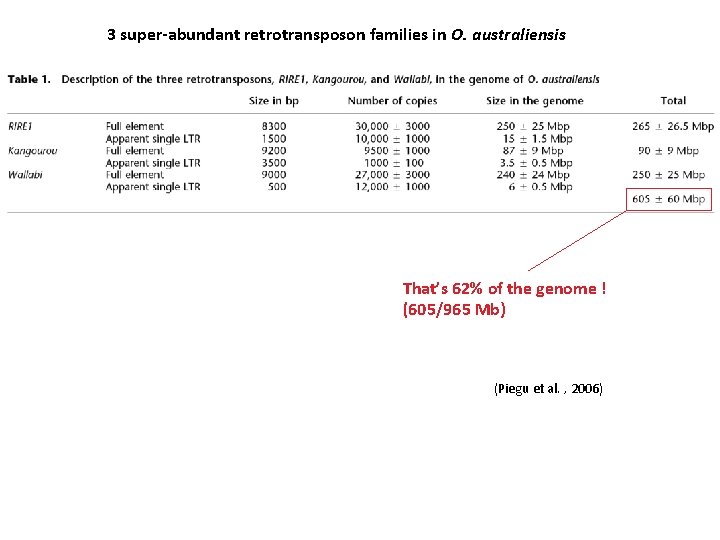
3 super-abundant retrotransposon families in O. australiensis That’s 62% of the genome ! (605/965 Mb) (Piegu et al. , 2006)

The solution to the paradox Most eukaryotic DNA does not code for proteins, so there is no reason to expect a complex organism to have a large genome or a simple organism to have a small one.

“The C-value paradox vanished the moment geneticists abandoned the concept of the genome consisting of the genes, all the genes, and nothing but the genes”

C…. G…. &…. . I values • C -Value : The amount DNA found in haploid genome, measured in million base pairs or in pg. • G- Value: The number of gene found in the haploid genome; the number includes predicted and ORFs. • I- value: The amount of information embedded by the genome.

“We’re pretty good at thinking about how individual genes are turned on and off. We’re not as good at thinking about how the whole genome is coordinated. ” Quote of Jeanne Lawrence in “The Cell Nucleus Shapes up“ Science 1993, Vol 259, pp 1257 -1259

Constitution of eukaryotic genome

Eukaryotic genomes composition 1. 2. 3. 4. Structural genes => Interrupted genes Conserved exons & unique introns Gene numbers Repetitive DNA (e. g. tandem gene clusters, tandem arrays)

1, 2 Eukaryotic genes are often interrupted Interrupted gene= A gene in which the coding sequence is not continuous due to the presence of introns.
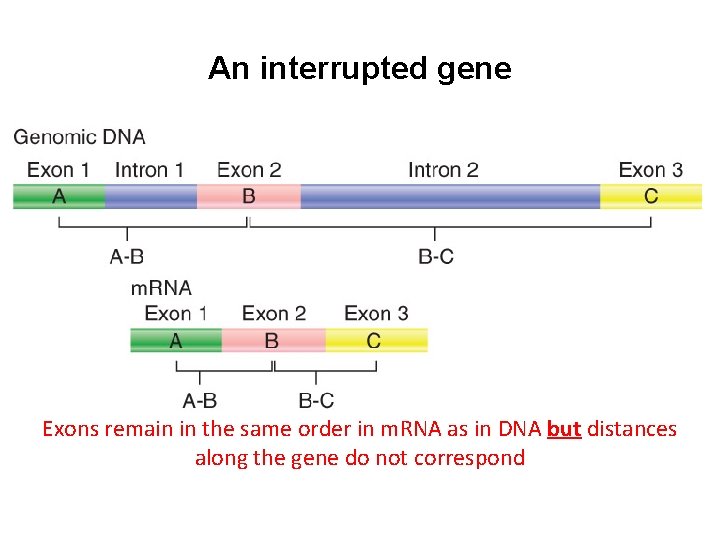
An interrupted gene Exons remain in the same order in m. RNA as in DNA but distances along the gene do not correspond

Interrupted genes • An interrupted gene consists of exons + introns • Introns usually do not encode proteins. • Introns are removed by the process of RNA splicing, which occurs only in cis "on the same side" on an individual RNA molecule. Organization may be conserved
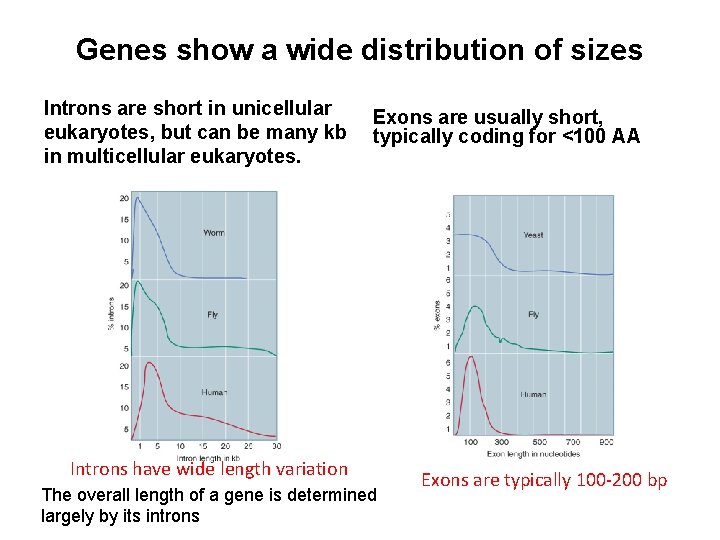
Genes show a wide distribution of sizes Introns are short in unicellular eukaryotes, but can be many kb in multicellular eukaryotes. Exons are usually short, typically coding for <100 AA Introns have wide length variation The overall length of a gene is determined largely by its introns Exons are typically 100 -200 bp
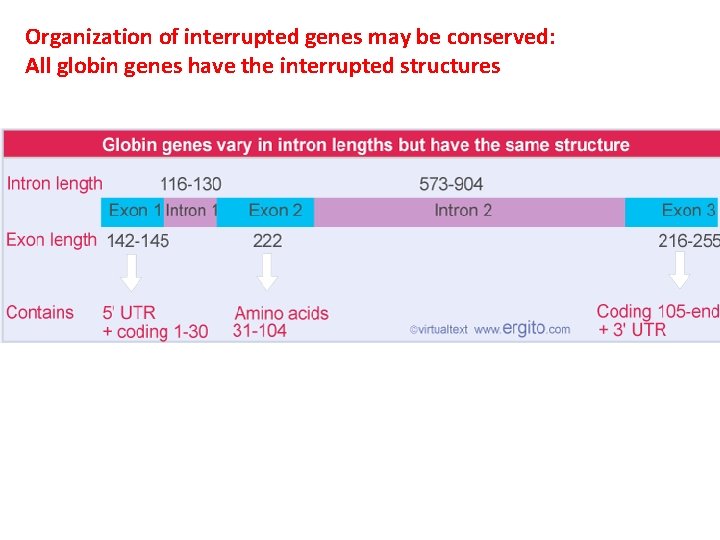
Organization of interrupted genes may be conserved: All globin genes have the interrupted structures
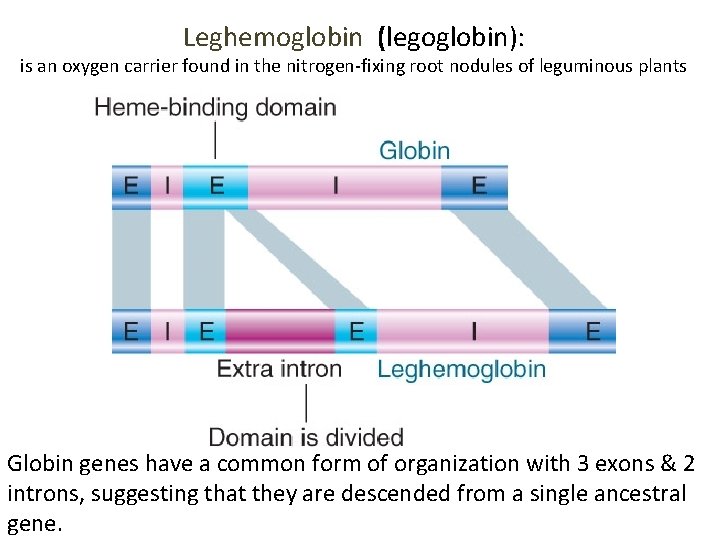
Leghemoglobin (legoglobin): is an oxygen carrier found in the nitrogen-fixing root nodules of leguminous plants Globin genes have a common form of organization with 3 exons & 2 introns, suggesting that they are descended from a single ancestral gene.
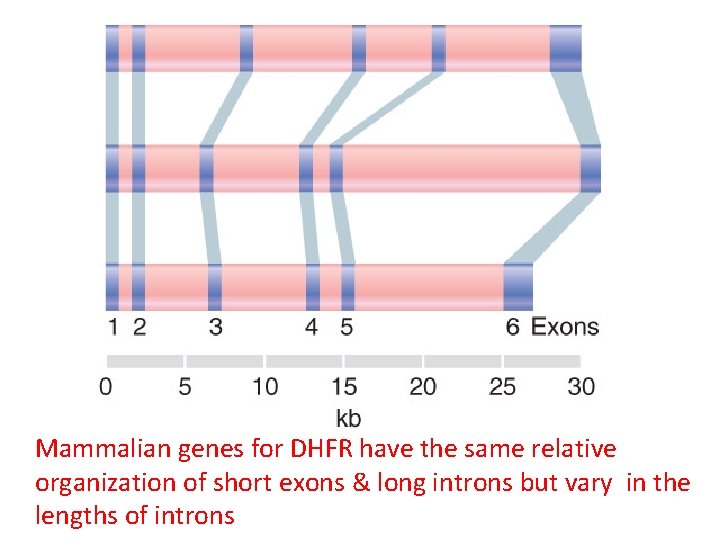
Mammalian genes for DHFR have the same relative organization of short exons & long introns but vary in the lengths of introns
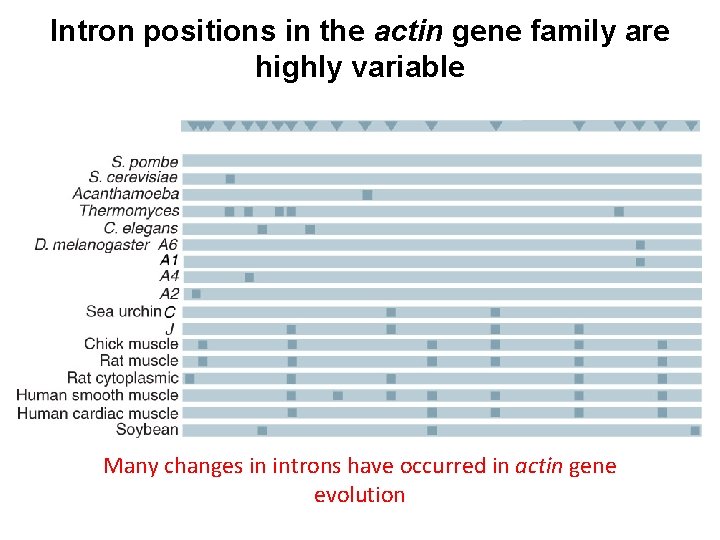
Intron positions in the actin gene family are highly variable Many changes in introns have occurred in actin gene evolution
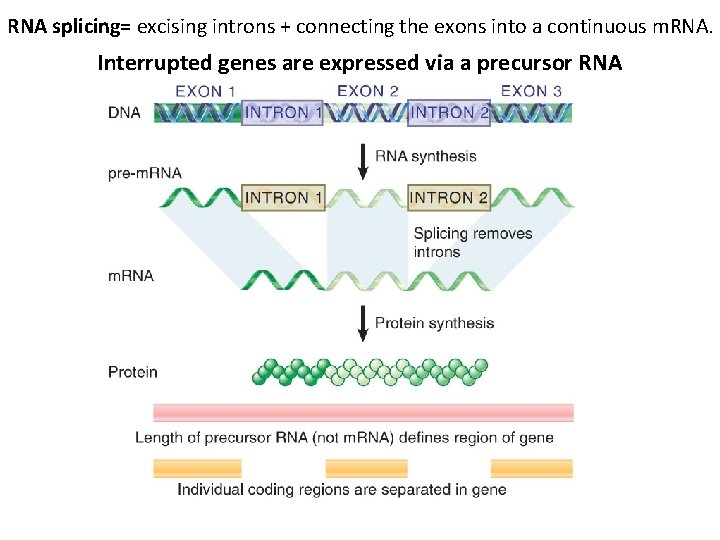
RNA splicing= excising introns + connecting the exons into a continuous m. RNA. Interrupted genes are expressed via a precursor RNA

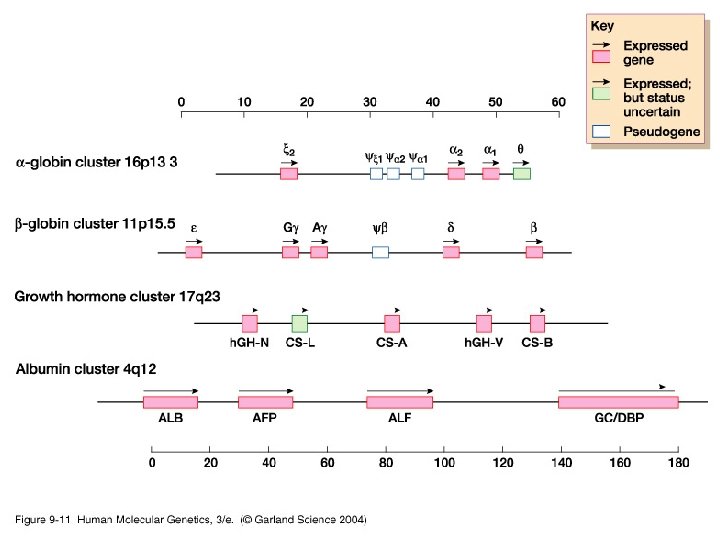
4, 5. tandemly repeated DNA a) coding: Members of a Gene Family Have a Common Organization • gene family= A set of genes within a genome that codes for related or identical proteins or RNAs. – The members were derived by duplication of an ancestral gene followed by accumulation of changes in sequence between the copies. – Most often the members are related but not identical. – Often encoded by recently duplicated genes e. g. αglobin genes. Very occasionally some genes on different chromosomes encode identical polypeptides.

Functionally similar genes are occasionally clustered in the human genome, but are more often dispersed over different chromosomes • Histone genes= 86 genes in 10 chromosomes, with 2 large clusters on short arm of chromosome 6. • Ubiquitin genes: highly conserved 76 AAs ubiquitin involved in protein degradation and cellular stress response. Distributed over several chromosomes
09_11. jpg
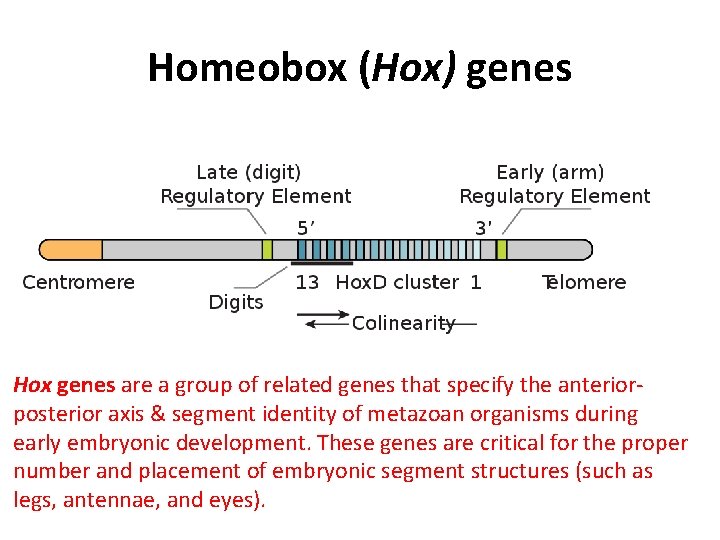
Homeobox (Hox) genes Hox genes are a group of related genes that specify the anteriorposterior axis & segment identity of metazoan organisms during early embryonic development. These genes are critical for the proper number and placement of embryonic segment structures (such as legs, antennae, and eyes).

Fruit fly genes regulated by the Hox genes
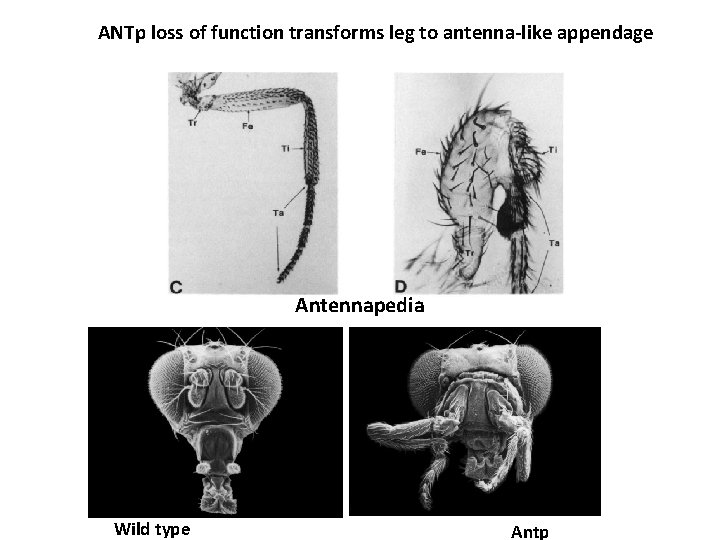
ANTp loss of function transforms leg to antenna-like appendage Antennapedia Wild type Antp

b) non-coding tandem arrays 1. VNTR (variable number tandem repeat)= very short repeated sequences, include: I. Microsatellite DNAs consist of repetitions of extremely short (typically <10 bp) units. II. Minisatellite DNAs consist of ~10 copies of a short repeating sequence. • The number of repeats varies between individual genomes. 2. Transposable elements derived
09_01. jpg
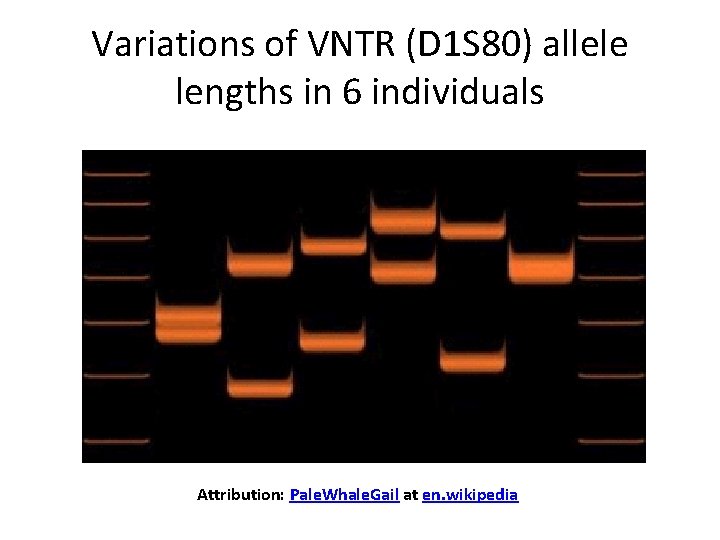
Variations of VNTR (D 1 S 80) allele lengths in 6 individuals Attribution: Pale. Whale. Gail at en. wikipedia
- Slides: 50