yvind Halskau Dept of Mol Biol University of

Øyvind Halskau, Dept. of Mol. Biol, University of Bergen Photo credit: Svein-Magne Tunli
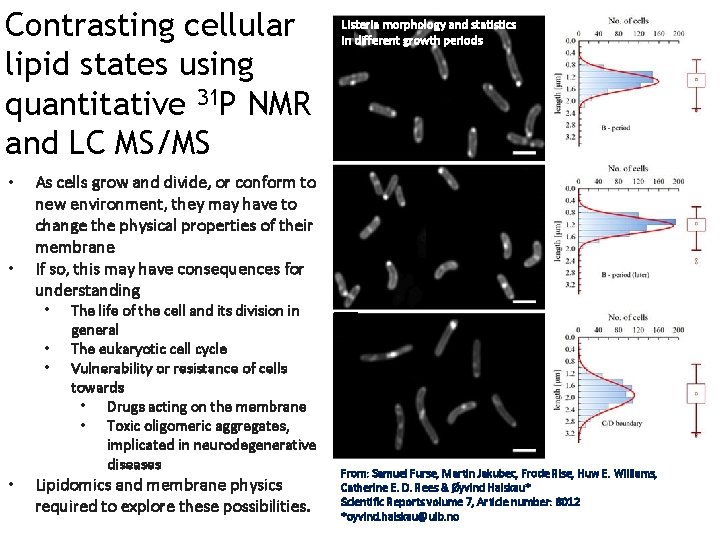
Contrasting cellular lipid states using quantitative 31 P NMR and LC MS/MS • • As cells grow and divide, or conform to new environment, they may have to change the physical properties of their membrane If so, this may have consequences for understanding • • Listeria morphology and statistics in different growth periods The life of the cell and its division in general The eukaryotic cell cycle Vulnerability or resistance of cells towards • Drugs acting on the membrane • Toxic oligomeric aggregates, implicated in neurodegenerative diseases Lipidomics and membrane physics required to explore these possibilities. From: Samuel Furse, Martin Jakubec, Frode Rise, Huw E. Williams, Catherine E. D. Rees & Øyvind Halskau* Scientific Reports volume 7, Article number: 8012 *oyvind. halskau@uib. no
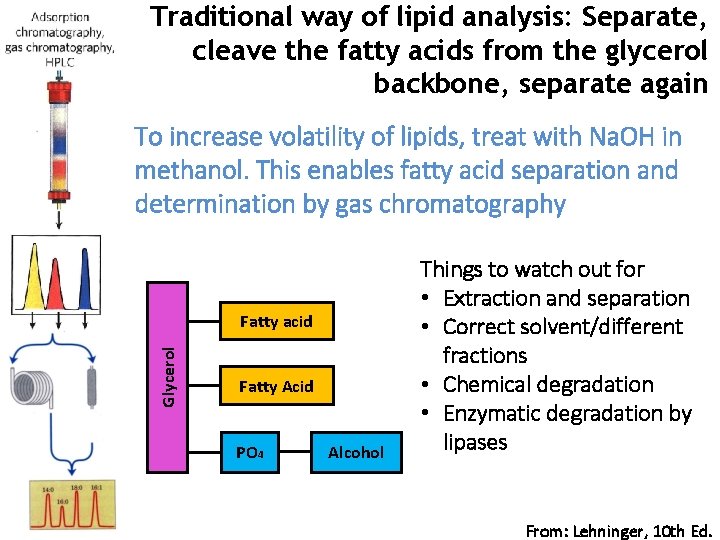
Traditional way of lipid analysis: Separate, cleave the fatty acids from the glycerol backbone, separate again To increase volatility of lipids, treat with Na. OH in methanol. This enables fatty acid separation and determination by gas chromatography Glycerol Fatty acid Fatty Acid PO 4 Alcohol Things to watch out for • Extraction and separation • Correct solvent/different fractions • Chemical degradation • Enzymatic degradation by lipases From: Lehninger, 10 th Ed.

Could 31 P NMR be used to identify and quantify lipids? 31 P is fairly sensitive, natural abundance practically 100% Quantification by NMR requires: • Complete relaxation between scans Relaxation delays of R 1 x 5 = 5 sec • Complete decoupling Inverse gated decoupling, WALTZ 16 works fine • Equal excitation of all signals Most relevant signals within 6 -7 ppm, plenty margin available • Good baseline and good separation Let’s have a closer look • Minimum of about 3: 1 signal/noise ratio Let’s have a closer look
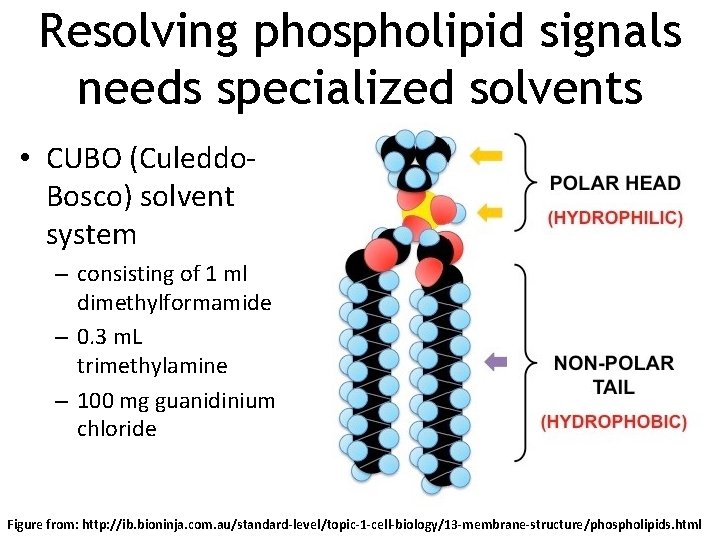
Resolving phospholipid signals needs specialized solvents • CUBO (Culeddo. Bosco) solvent system – consisting of 1 ml dimethylformamide – 0. 3 m. L trimethylamine – 100 mg guanidinium chloride Figure from: http: //ib. bioninja. com. au/standard-level/topic-1 -cell-biology/13 -membrane-structure/phospholipids. html
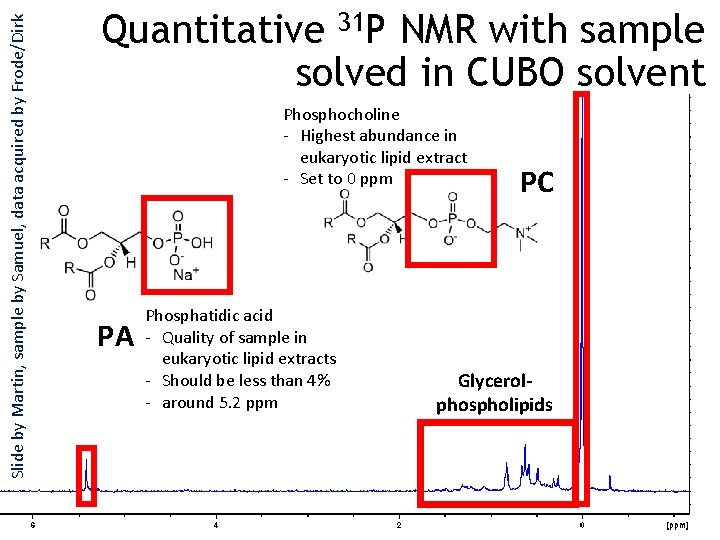
Slide by Martin, sample by Samuel, data acquired by Frode/Dirk Quantitative 31 P NMR with sample solved in CUBO solvent Phosphocholine - Highest abundance in eukaryotic lipid extract - Set to 0 ppm PA Phosphatidic acid - Quality of sample in eukaryotic lipid extracts - Should be less than 4% - around 5. 2 ppm PC Glycerolphospholipids
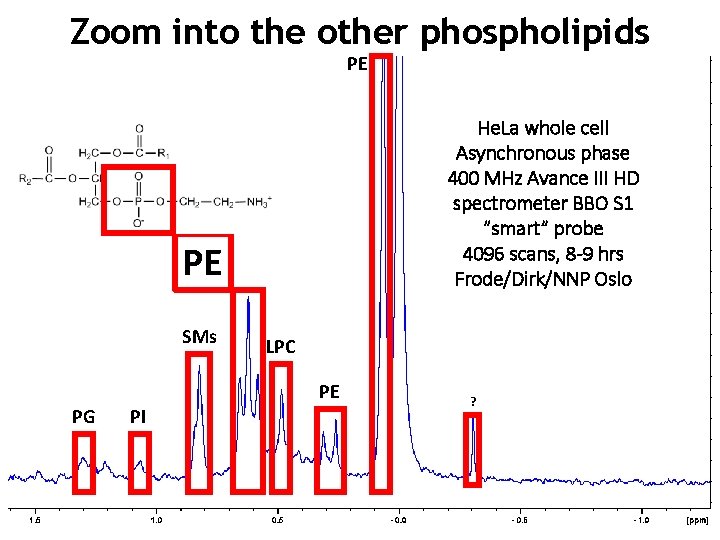
Zoom into the other phospholipids PE He. La whole cell Asynchronous phase 400 MHz Avance III HD spectrometer BBO S 1 “smart” probe 4096 scans, 8 -9 hrs Frode/Dirk/NNP Oslo PE SMs LPC PE PG PI ?
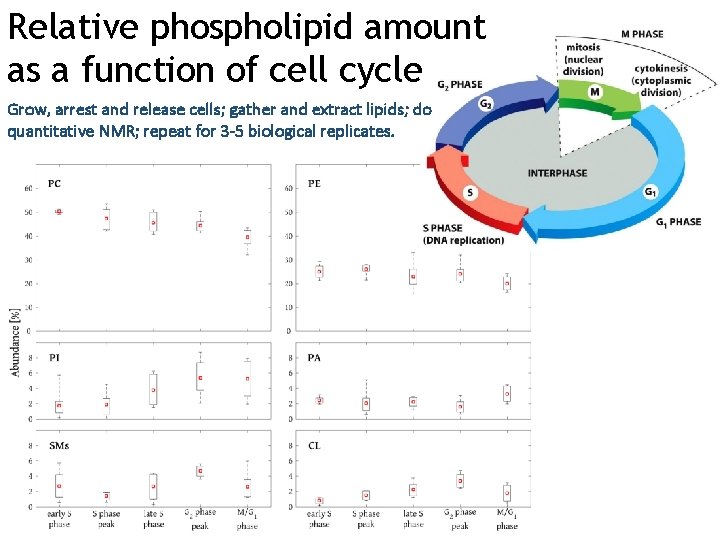
Relative phospholipid amount as a function of cell cycle Grow, arrest and release cells; gather and extract lipids; do quantitative NMR; repeat for 3 -5 biological replicates.
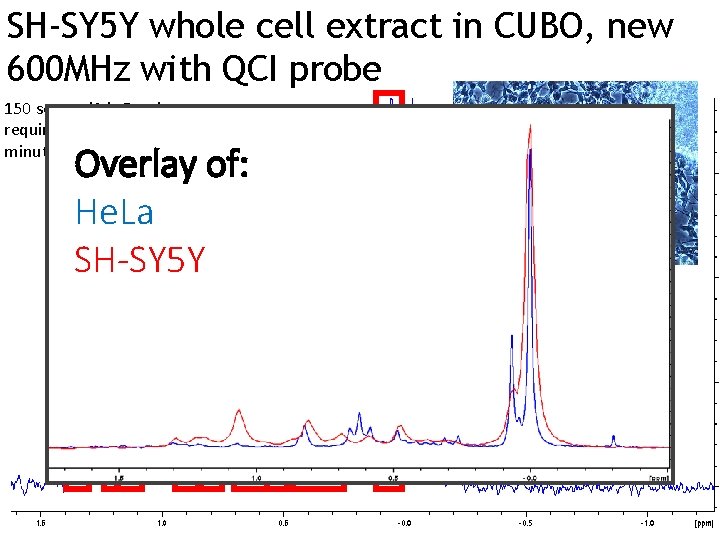
SH-SY 5 Y whole cell extract in CUBO, new 600 MHz with QCI probe 150 scans, d 1 is 8 s, time SMs required is as little as 23 minutes PI PE Overlay of: He. La SH-SY 5 Y ? PE PS PG ? PE SH-SYS 5 cells derived from neuroblastoma of human patient; can show characteristics of neural cells, and therefore used as a model system
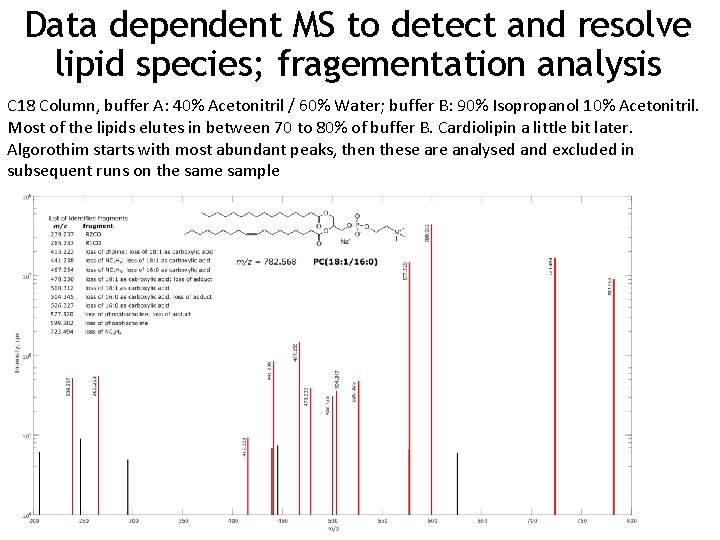
Data dependent MS to detect and resolve lipid species; fragementation analysis C 18 Column, buffer A: 40% Acetonitril / 60% Water; buffer B: 90% Isopropanol 10% Acetonitril. Most of the lipids elutes in between 70 to 80% of buffer B. Cardiolipin a little bit later. Algorothim starts with most abundant peaks, then these are analysed and excluded in subsequent runs on the sample
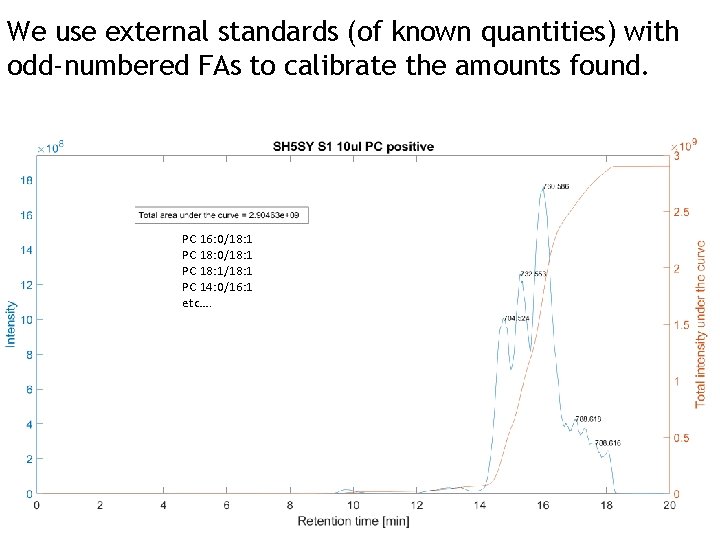
We use external standards (of known quantities) with odd-numbered FAs to calibrate the amounts found. PC 16: 0/18: 1 PC 18: 1/18: 1 PC 14: 0/16: 1 etc….

Summary of pros and cons: 31 P NMR • No separation required • Easier and better quantification than MSbased methods • Large amounts (ca 10 mg per sample) of material, or instrument time • Only phosho-containing lipids detectable i current setup • Little or no acyl-chain info Mass spectrometry • Potentially complete identification of all lipids • Acyl-chains characterized • Small amounts of sample needed • Only around 16 scan per second. With about a 1000 lipids eluting at the same time, we have to do a lot of runs • Extraction AND separation needed • No quantification, unless you have specifically set up methods using external standards

ACKNOWLEDGMENTS Martin Jakubec Samuel Furse Aurelia Lewis Frode Rise For SH-SYS 5 work also Espen Bariås Linda Veka Hjørnevik For Listeria paper also: Hew Willaims Catherine Rees Funding and Resources: University of Bergen RCN grant #240063 NNP Oslo and Bergen Nodes Infrastructure grant 226244 / F 50
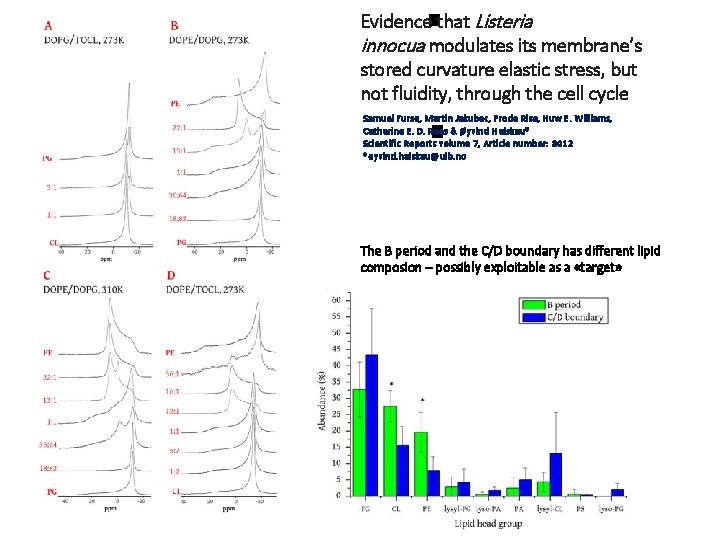
Evidence that Listeria innocua modulates its membrane’s stored curvature elastic stress, but not fluidity, through the cell cycle Samuel Furse, Martin Jakubec, Frode Rise, Huw E. Williams, Catherine E. D. Rees & Øyvind Halskau* Scientific Reports volume 7, Article number: 8012 *oyvind. halskau@uib. no The B period and the C/D boundary has different lipid composion – possibly exploitable as a «target»
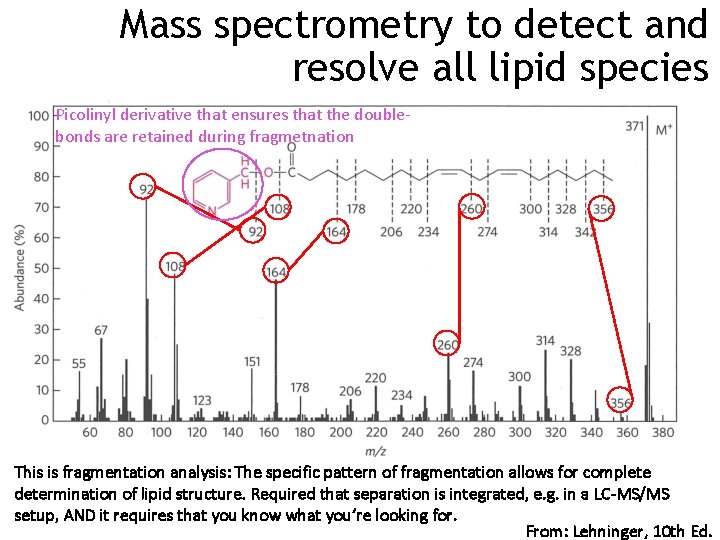
Mass spectrometry to detect and resolve all lipid species Picolinyl derivative that ensures that the doublebonds are retained during fragmetnation This is fragmentation analysis: The specific pattern of fragmentation allows for complete determination of lipid structure. Required that separation is integrated, e. g. in a LC-MS/MS setup, AND it requires that you know what you’re looking for. From: Lehninger, 10 th Ed.
- Slides: 15