Xray crystallography an overview based on Bernie Browns

X-ray crystallography – an overview (based on Bernie Brown’s talk, Dept. of Chemistry, WFU) • Protein is crystallized (sometimes low-gravity atmosphere is helpful e. g. NASA) • X-Rays are scattered by electrons in molecule • Diffraction produces a pattern of spots on a film that must be mathematically deconstructed • Result is electron density (contour map) – need to know protein sequence and match it to density • Hydrogen atoms not typically visible (except at very high resolution)
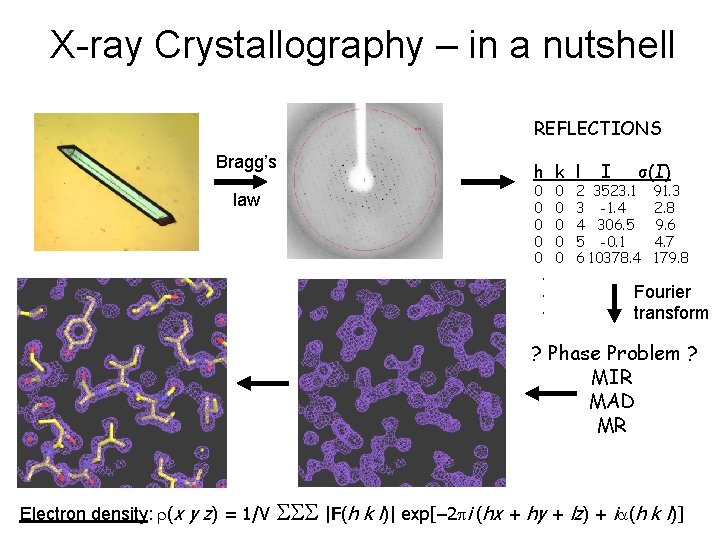
X-ray Crystallography – in a nutshell REFLECTIONS Bragg’s law h k l 0 0 0. . . 0 0 0 I σ(I) 2 3523. 1 3 -1. 4 4 306. 5 5 -0. 1 6 10378. 4 91. 3 2. 8 9. 6 4. 7 179. 8 Fourier transform ? Phase Problem ? MIR MAD MR Electron density: r(x y z) = 1/V SSS |F(h k l)| exp[– 2 pi (hx + hy + lz) + ia(h k l)]

Crystal formation • Start with supersaturated solution of protein • Slowly eliminate water from the protein • Add molecules that compete with the protein for water (3 types: salts, organic solvents, PEGs) • Trial and error • Most crystals ~50% solvent • Crystals may be very fragile

Visible light vs. X-rays Why don’t we just use a microscope to look at proteins? • Size of objects imaged limited by wavelength. Resolution ~ l/2 – Visible light – 4000 -7000 Å (400 -700 nm) – X-rays – 0. 7 -1. 5 Å (0. 07 -0. 15 nm) • It is very difficult to focus X-rays (Fresnel lenses) • Getting around the problem – Defined beam – Regular structure of object (crystal) • Result – diffraction pattern (not a focused image).
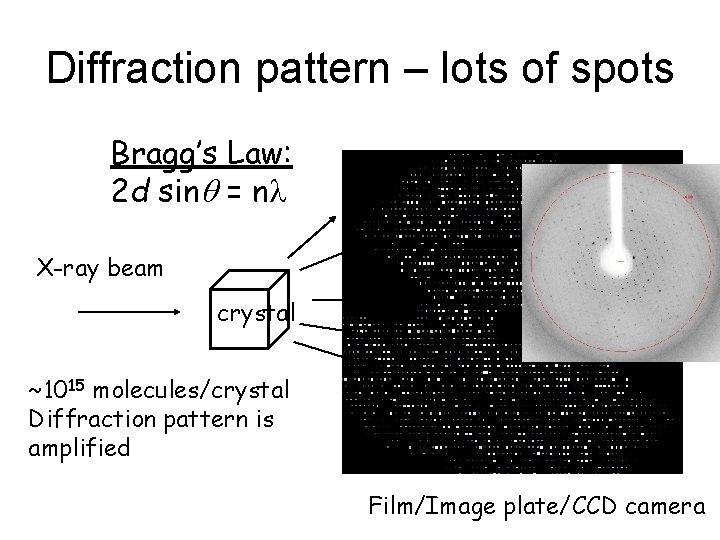
Diffraction pattern – lots of spots Bragg’s Law: 2 d sinq = nl X-ray beam crystal ~1015 molecules/crystal Diffraction pattern is amplified Film/Image plate/CCD camera
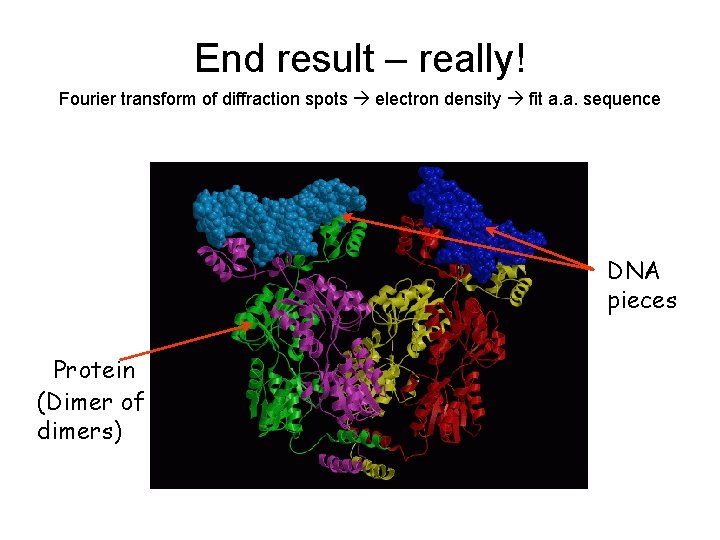
End result – really! Fourier transform of diffraction spots electron density fit a. a. sequence DNA pieces Protein (Dimer of dimers)
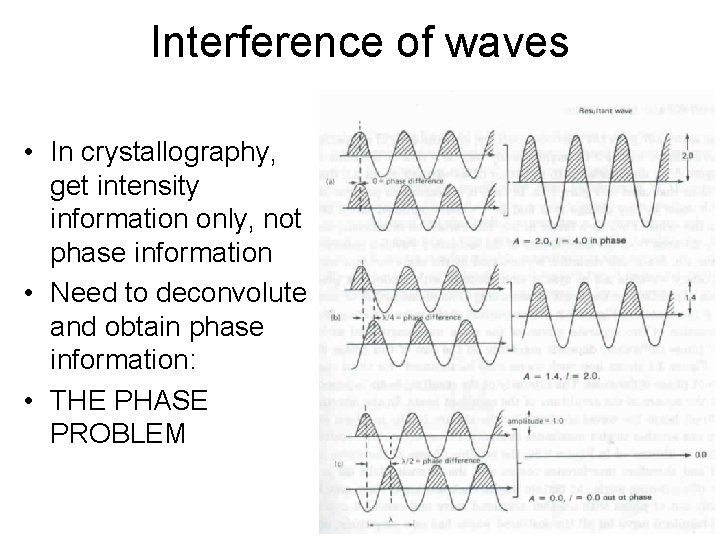
Interference of waves • In crystallography, get intensity information only, not phase information • Need to deconvolute and obtain phase information: • THE PHASE PROBLEM

How to get from spots to structure? • Fourier synthesis • Getting around phase problem – Trial and error – Previous structures – Heavy atom replacement – make a landmark – Ex: Selenomethionine • Plenty of computer algorithms now
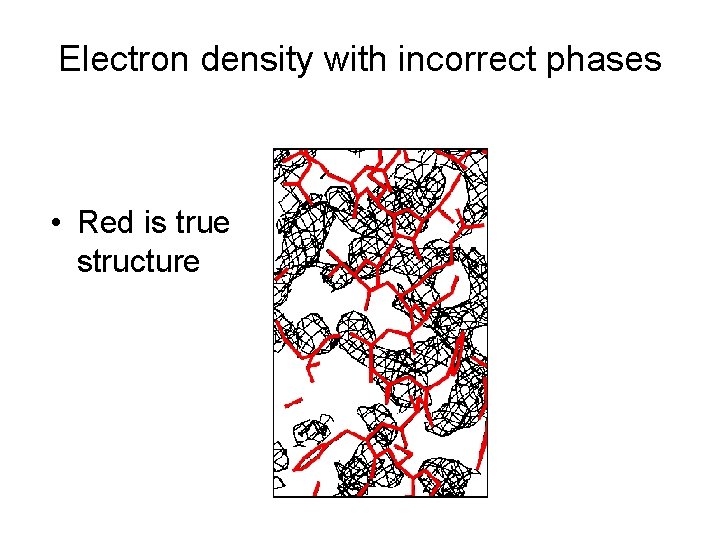
Electron density with incorrect phases • Red is true structure

The effect of resolution More extensive diffraction pattern gives more structural information = higher resolution • 6. 0 -4. 5 Å – secondary structure elements • 3. 0 Å – trace polypeptide chain • 2. 0 Å – side chain, bound water identification • 1. 8 Å – alternate side chain orientations • 1. 2 Å – hydrogen atoms

With computational tools, spots become density Flexible regions give smeared density, often 2 -3 conformations visible, more than that invisible
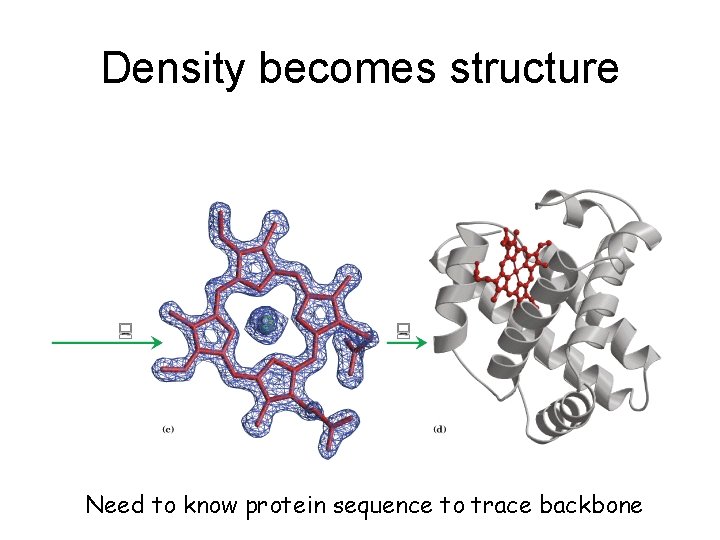
Density becomes structure Need to know protein sequence to trace backbone

Co-crystal structures • Because of relatively high solvent content, can often “soak in” substrate • Then can solve structure of protein with substrate bound • If crystal cracks, good sign that substrate binding or enzyme catalysis results in conformational change in protein • No longer has same crystal arrangement
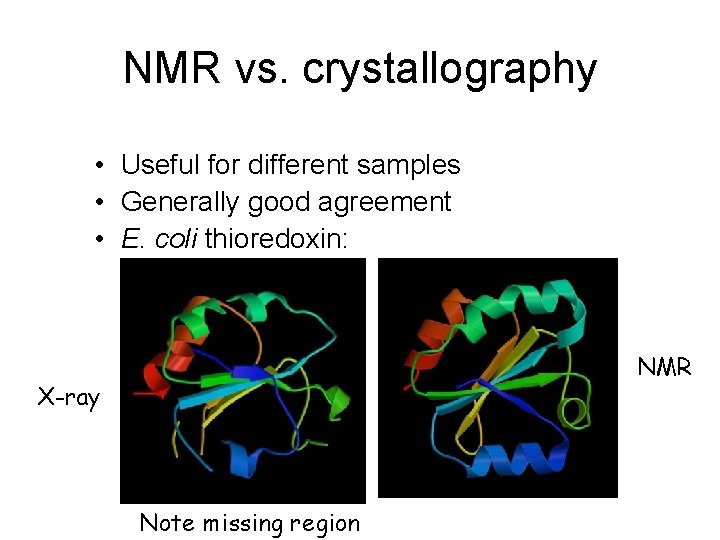
NMR vs. crystallography • Useful for different samples • Generally good agreement • E. coli thioredoxin: NMR X-ray Note missing region

Known protein structures • ~17, 000 protein structures since 1958 • Common depository of x, y, z coordinates: Protein data bank (http: //www. rcsb. org) • Coordinates can be extracted and viewed • Comparisons of structures allows identification of structural motifs • Proteins with similar functions and sequences = homologs
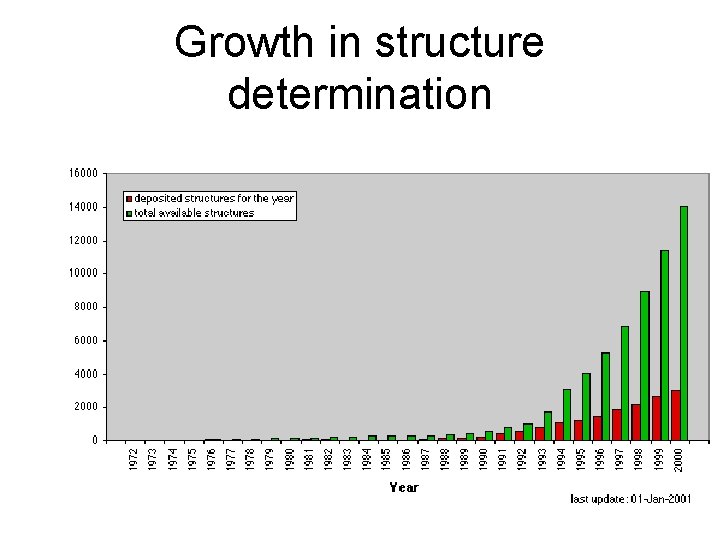
Growth in structure determination
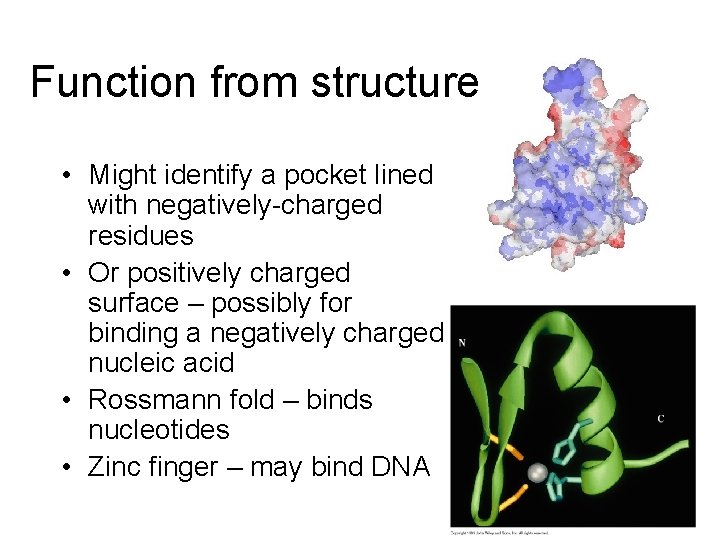
Function from structure • Might identify a pocket lined with negatively-charged residues • Or positively charged surface – possibly for binding a negatively charged nucleic acid • Rossmann fold – binds nucleotides • Zinc finger – may bind DNA

Domain organization • Large proteins have polypeptide regions that fold in isolation • May have distinct functional roles – Example: glyceraldehyde-3 phosphate dehydrogenase

Protein families • Similar function and overall structure • But amino acid sequence may or may not be highly conserved • Limited number of protein domains • Homologs versus structural motifs
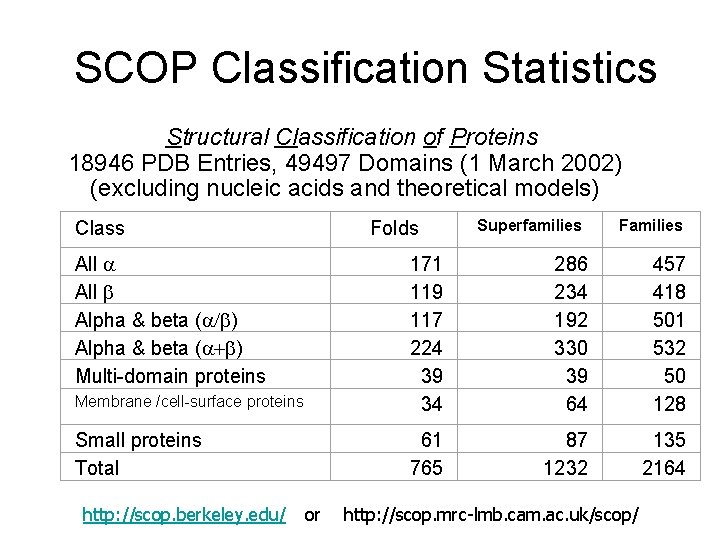
SCOP Classification Statistics Structural Classification of Proteins 18946 PDB Entries, 49497 Domains (1 March 2002) (excluding nucleic acids and theoretical models) Class Folds All a All b Alpha & beta (a/b) Alpha & beta (a+b) Multi-domain proteins Superfamilies Families Membrane /cell-surface proteins 171 119 117 224 39 34 286 234 192 330 39 64 457 418 501 532 50 128 Small proteins Total 61 765 87 1232 135 2164 http: //scop. berkeley. edu/ or http: //scop. mrc-lmb. cam. ac. uk/scop/
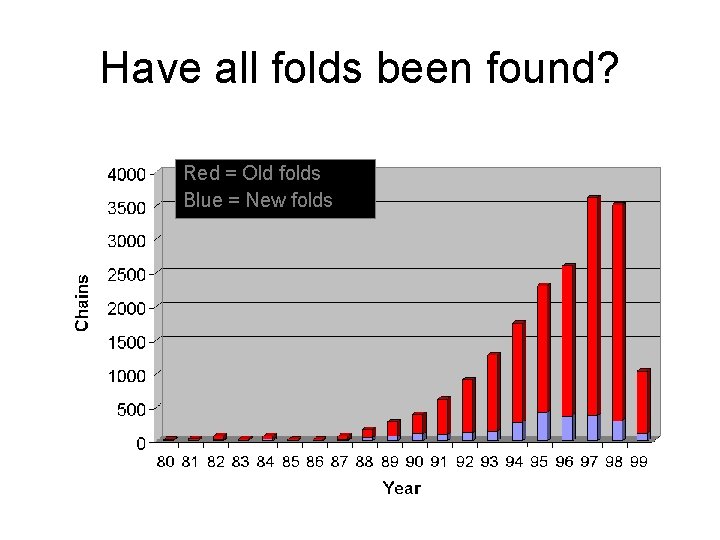
Have all folds been found? Red = Old folds Blue = New folds
- Slides: 21