XIP InVivo Imaging Workspace Software SIG February 7

XIP In-Vivo Imaging Workspace Software SIG February 7, 2007

What is • • • ? The e. Xtensible Imaging Platform (XIP) is an open source environment for rapidly developing medical imaging applications from an extensible set of modular elements. Researchers will be able to easily develop and evaluate new approaches to medical imaging problems, and use them in a translational research setting. ca. Grid makes it possible to develop an XIP architecture that allows users to choose between remotely hosted grid-based components and data sources as well as locally available components and sources. • Components may include analytic services, e. g. • CAD algorithms, • algorithms for quantifying changes in consecutive imaging studies, • algorithms associated with a 3 -D visualization pipeline etc) • Available data sources include • NCIA • DICOM data repositories • Local databases

Why • • • ? One of the goals of the In Vivo Imaging Workspace is to “focus on identifying the ways in which the wealth of information provided by … imaging, performed at academic and other research centers across the country, can be shared, optimized, and most effectively integrated into the ongoing effort to relieve suffering and death from cancer. ” To facilitate the increasing use of imaging based end points in clinical trials, the Workspace has identified the need for an easily extensible open source platform to support image analysis and visualization. This platform will make it easier and less expensive to access specific post-processing applications at multiple sites, simplifying clinical trials, and most importantly, increasing the uniformity of imaging and analysis. Imaging applications developed by research groups will more easily be accessible within the clinical operating environment, simplifying workflows and speeding data processing and analysis. Once validated, the software should be readily transitioned into products through streamlined Federal Drug Administration, (FDA), approval processes due to the re-use of already approved libraries and open source development processes.

XIP Use Cases Clinical Translation • Four Use Cases defined by the ca. BIG IVI Workspace, Software SIG (F. Prior, B. Erickson), in the XIP Requirements Document • Imaging as a bio-marker for drug therapy trials with centralized data analysis within a Cooperative Group: NCCTG • Imaging as a bio-marker for research and drug therapy trials using distributed analysis within a Cooperative Group: COG, NANT • Standardizing and translating emerging imaging methods in a Translational Research Network: NTROI • Annotation of tumors as part of curation process for an image archive to be used for algorithm development: NCIA • These Use Cases illustrate the spectrum of uses of XIP – the Software SIG will guide the prioritization and roll-out of features to meet the needs of such representative users

Imaging as a Bio-marker, Classical Phase 2/3 (NCCTG) • Use Case: • Trial to assess effect on brain tumor of drug with & without radiation • Brain MRI at 2 mo. intervals, collected in central review facility • Functionality needed in XIP Imaging Application: • Query DICOM worklist created for each rater • Visualize baseline (pre-gad, post-gad, T 2, FLAIR) and current scan • Image registration (rigid body) to improve accuracy and reliability • Presentation with linked cursors and multi-planar reformats • Measurement of tumor size based on margin outlines (T 2 and/or FLAIR), both RECIST and volume • Controlled order of case presentation to reduce bias • Quickly find cases with significant inter-reader differences for adjudication
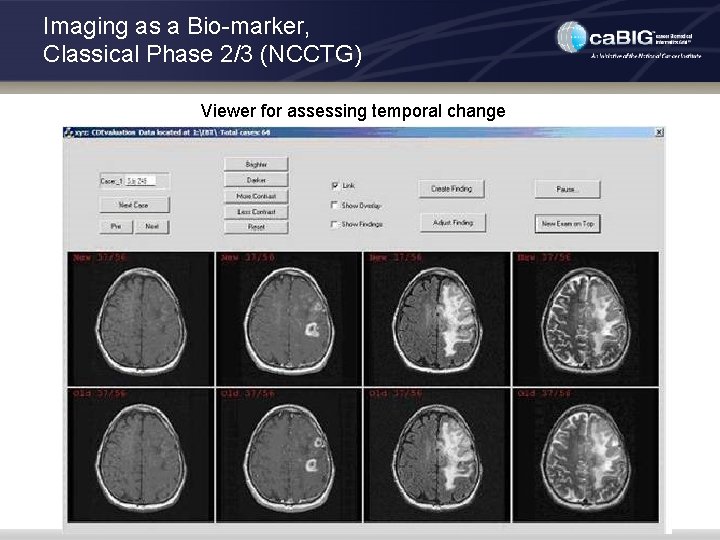
Imaging as a Bio-marker, Classical Phase 2/3 (NCCTG) Viewer for assessing temporal change

Imaging as a Bio-marker in Cooperative Group Trials (COG, NANT) • Use Case: • Evaluation of cases of Peripheral Neuroblastic Tumors (PNTs), integrating radiological and pathological images, patient demographics, … • Emphasis on MR perfusion analysis as bio-marker for tumor growth rate • Functionality needed in XIP Imaging Application: • “Virtual Workbench” for PNT research based on original data, derived results, annotations, and mark-ups of PNT data based on the Cooperative Group Grid resources • Interactive and integrated radiological and pathological image analysis of PNT’s • Publishing of analysis results to Grid Storage systems • Querying of grid-based data systems for discovery of outcome correlations
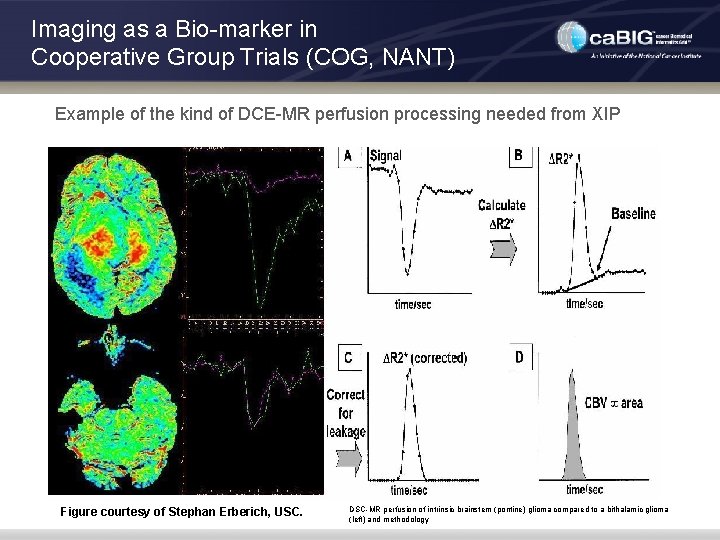
Imaging as a Bio-marker in Cooperative Group Trials (COG, NANT) Example of the kind of DCE-MR perfusion processing needed from XIP Figure courtesy of Stephan Erberich, USC. DSC-MR perfusion of intrinsic brainstem (pontine) glioma compared to a bithalamic glioma (left) and methodology
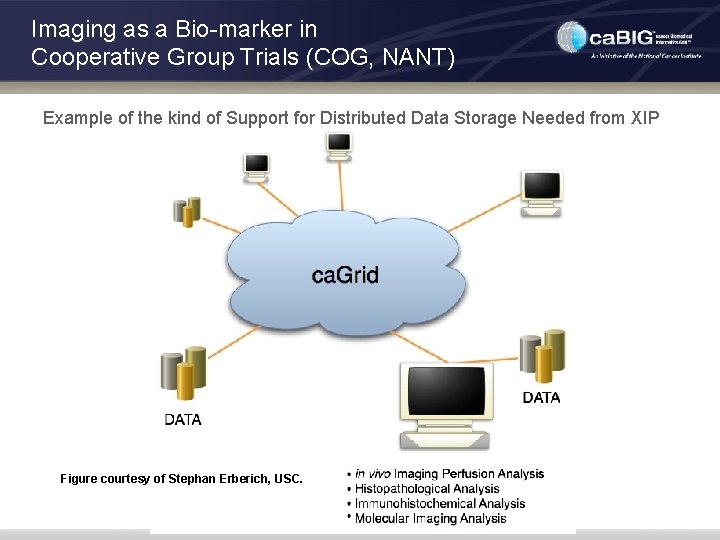
Imaging as a Bio-marker in Cooperative Group Trials (COG, NANT) Example of the kind of Support for Distributed Data Storage Needed from XIP Figure courtesy of Stephan Erberich, USC.

Standardizing Imaging in a Translational Research Network (NTROI) • Use Case • Standardization and validation of optical imaging methods for breast cancer screening and therapy monitoring • Starting point is collaborative validation of physiological measures of unique instruments at each university • End point is trial using common, validated instrument • Functionality Needed in XIP Imaging Application • Multi-modal registration, visual fusion, tumor segmentation • Measurement of tumor volume, optically-derived physiological parameters
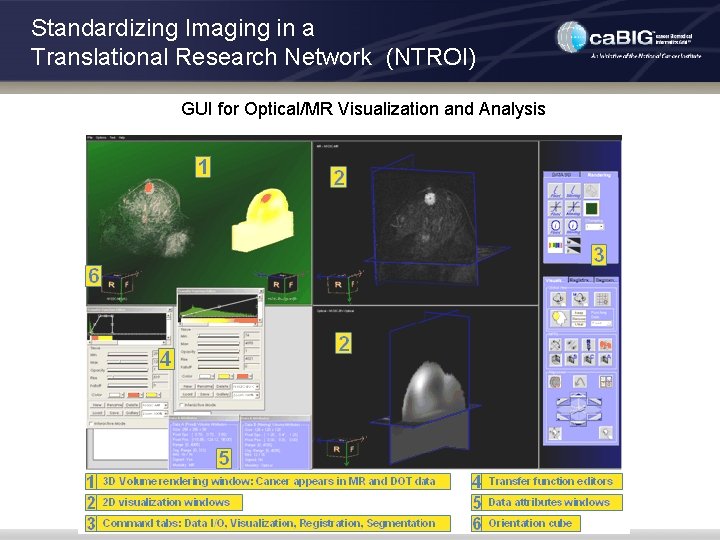
Standardizing Imaging in a Translational Research Network (NTROI) GUI for Optical/MR Visualization and Analysis
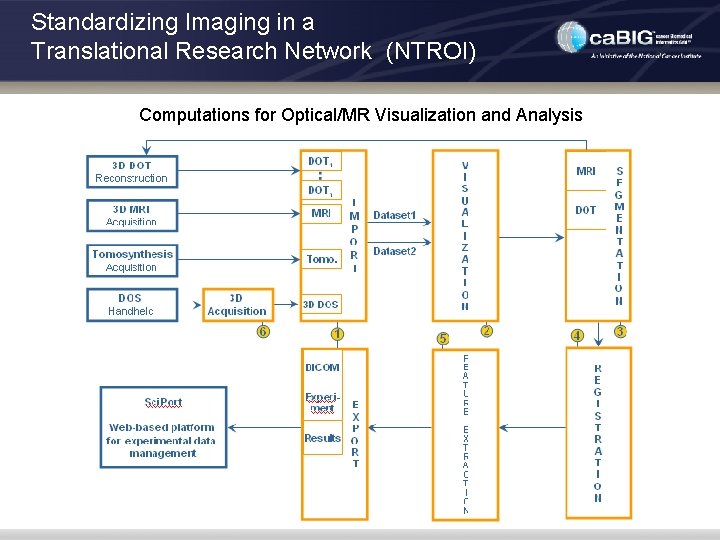
Standardizing Imaging in a Translational Research Network (NTROI) Computations for Optical/MR Visualization and Analysis

Annotation of tumors for an image archive (NCIA) • Use Case • Remote and on-site expert 3 D segmentation and rigorously defined annotation of tumors in National Cancer Image Archive • Annotated images used both to develop and test CAD and other algorithms by academia and industry • Functionality Needed in XIP Imaging Application • Advanced thin-client 3 D visualization and navigation tools for remote curation by domain experts • Mark-up, segmentation, annotation and measurement of tumor volume using a variety of 2 D and 3 D metrics, using rigorously defined vocabularies
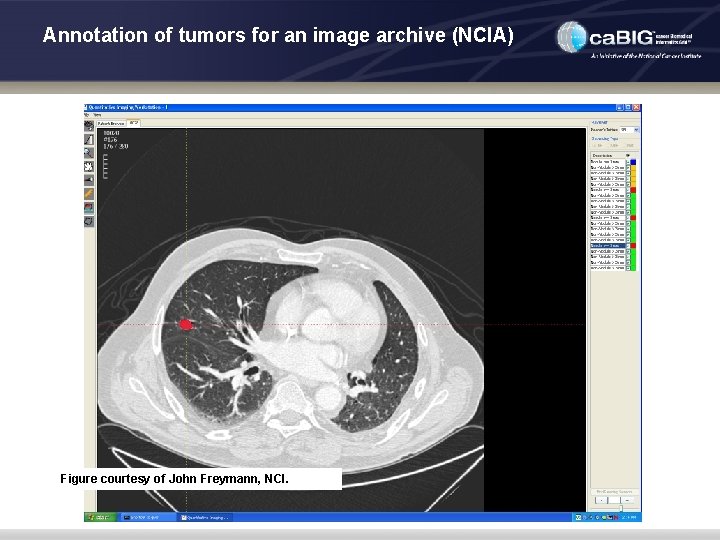
Annotation of tumors for an image archive (NCIA) Figure courtesy of John Freymann, NCI.

What is Included in • • XIP Rapid Application Development Tools and Libraries (RAD) • Development and application build environment • Extensive and extensible set of libraries for imaging and visualization • Uses Open Inventor framework • Includes code generating wizards to create new objects and wrap existing libraries XIP Workstation (WS) • A reference implementation of a medical imaging workstation developed using XIP RAD • Integrated via middleware into ca. GRID • Optimized to support basic cancer research use cases • Includes two key components: • XIP Application – a use case specific “plug-in” application integrated via the DICOM WG-23 Interface • XIP Host – the hosting environment that provides XIP Applications with access to services such as data stores, remote processing, etc.
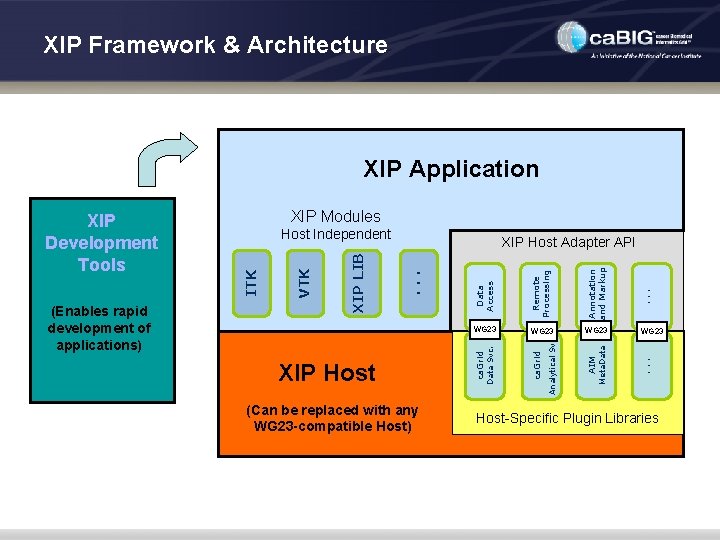
XIP Framework & Architecture XIP Application (Can be replaced with any WG 23 -compatible Host) . . . Annotation and Markup AIM Meta. Data WG 23 Analytical Svc. Remote Processing WG 23 ca. Grid WG 23 Data Svc. XIP Host Data Access XIP Host Adapter API. . . VTK Host Independent XIP LIB (Enables rapid development of applications) XIP Modules ITK XIP Development Tools Host-Specific Plugin Libraries
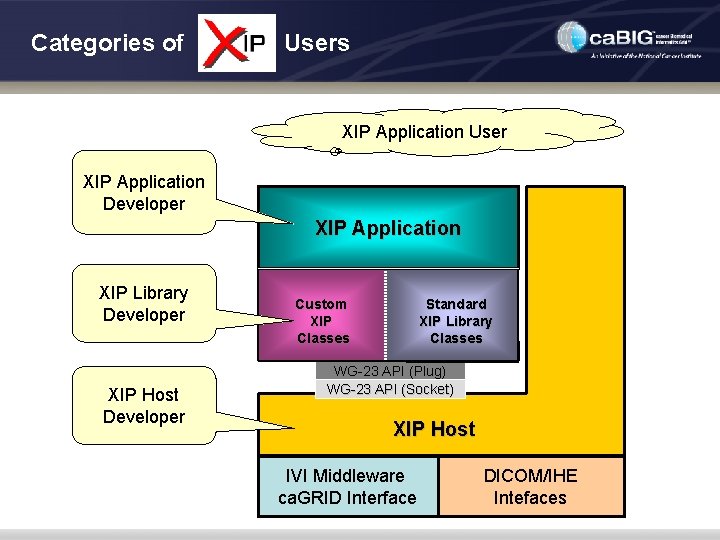
Categories of Users XIP Application User XIP Application Developer XIP Application XIP Library Developer XIP Host Developer Custom XIP Classes Standard XIP Library Classes WG-23 API (Plug) WG-23 API (Socket) XIP Host IVI Middleware ca. GRID Interface DICOM/IHE Intefaces
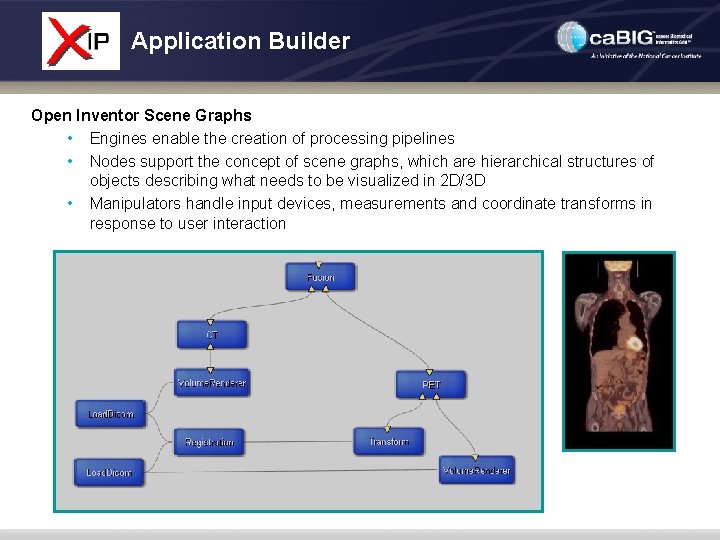
Application Builder Open Inventor Scene Graphs • Engines enable the creation of processing pipelines • Nodes support the concept of scene graphs, which are hierarchical structures of objects describing what needs to be visualized in 2 D/3 D • Manipulators handle input devices, measurements and coordinate transforms in response to user interaction
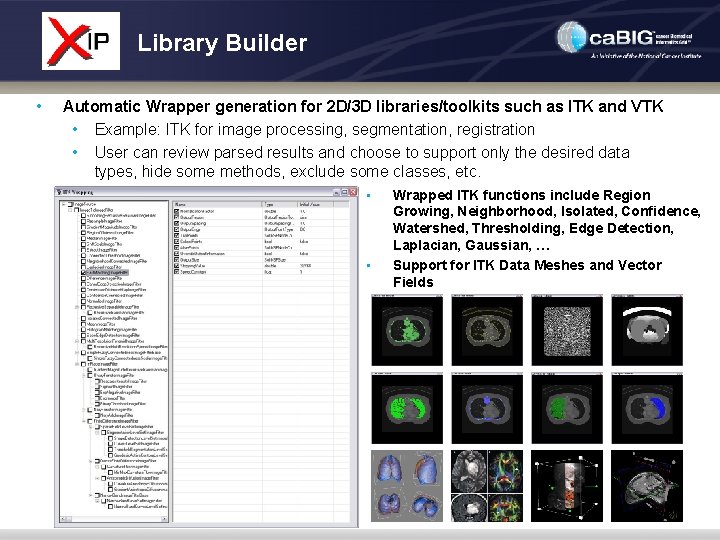
Library Builder • Automatic Wrapper generation for 2 D/3 D libraries/toolkits such as ITK and VTK • Example: ITK for image processing, segmentation, registration • User can review parsed results and choose to support only the desired data types, hide some methods, exclude some classes, etc. • • Wrapped ITK functions include Region Growing, Neighborhood, Isolated, Confidence, Watershed, Thresholding, Edge Detection, Laplacian, Gaussian, … Support for ITK Data Meshes and Vector Fields

Host Functions • • • Provides the infrastructure in which an XIP Application runs • Authenticates user • Manages installation, launching, and termination of XIP Applications • Provides data and services to XIP Applications • Accepts status information and results back from XIP Applications • Deals with auditing and controls access to services and data Isolates the XIP application from the nature of databases, archives, networks, and possibly image data formats • Manages ca. GRID interactions and security • Manages access to DICOM networks, objects, and services • Maps images and associated meta-data from various sources between their native form and a common form useable by the XIP application Handles workflow issues • General Purpose Worklist support, following IHE pattern
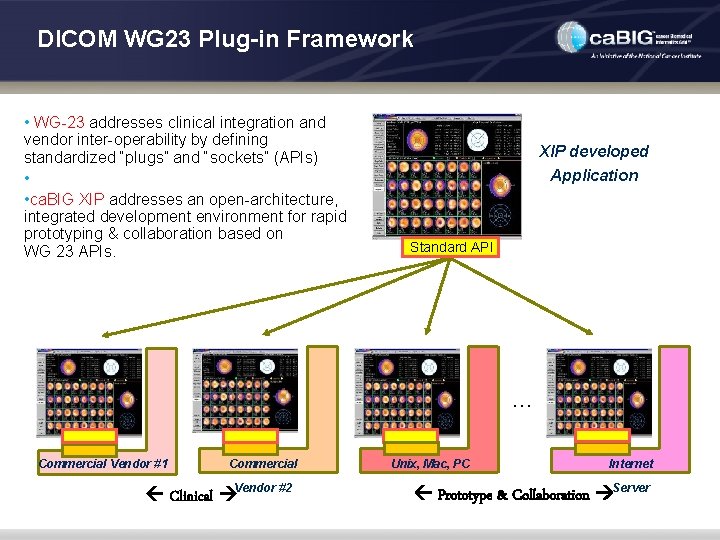
DICOM WG 23 Plug-in Framework • WG-23 addresses clinical integration and vendor inter-operability by defining standardized “plugs” and “sockets” (APIs) • • ca. BIG XIP addresses an open-architecture, integrated development environment for rapid prototyping & collaboration based on WG 23 APIs. XIP developed Application Standard API … Commercial Vendor #1 Commercial Clinical Vendor #2 Unix, Mac, PC Internet Prototype & Collaboration Server

Who is Contributing to • • • ? The ca. BIG In Vivo Imaging Workspace, Software SIG • Released the XIP RFP • Provides primary feedback to the XIP development team Washington University in St. Louis, Electronic Radiology Lab • Main coordinating site • Actively involved in ca. BIG, DICOM, IHE, and serves as imaging core for several clinical trials Siemens Corporate Research (SCR) • Contributing a suite of tools – iv. RAD – that will form the basis for XIP • Experienced in moving ideas from prototypes to commercial reality DICOM WG-23 • Standardizing the interfaces between a hosting system (e. g. workstation) and hosted post-processing applications (a. k. a. “Plug-ins”) • Representation from both vendors and user communities ITK/VTK community • Providing image processing and visualization libraries with the assistance of Kitware

What does iv. RAD bring to the table? • Application development framework following the Open Inventor API • • • Easy to extend • • • Import of medical image data sets (e. g. DICOM, raw) Manipulation of medical image data (e. g. registration, fusion, segmentation) Multi-dimensional visualization (e. g. cine, MPR, MIP, Shaded-surface display, Volume Rendering) Tools for wrapping 3 rd party class libraries into Open Inventor objects • • create custom Open Inventor-style objects create Open Inventor-style wrappers around existing libraries Extensions to support medical visualization and image processing • • Scene graphs coupled to processing pipelines and manipulators Well established, well documented, open source, free Used to incorporate the open source ITK (Insight Tool. Kit) into iv. RAD Hides peculiarities of the underlying host system from the application • • Allows the same application to run stand-alone on MS Windows, or within various versions of Siemens’ medical workstations Since the interfaces that the application sees remain constant, the application is unaware of the differences in the underlying system

iv. RAD to 1. Strip out Siemens-proprietary classes • Siemens-proprietary visualization and processing functions • Functions for Siemens-specific data sources • Siemens ‘look-and-feel’ 2. Change copyright notices to support an open source license 3. Replace Siemens-proprietary classes with open-source classes • Continued use of ITK (Insight Tool. Kit) • Continued use of open source DICOM toolkits (e. g. DCMTK or DCM 4 CHE) • Add visualization via VTK and other open-source classes 4. Modify the host – application interaction per the DICOM WG-23 APIs 5. Add support for new functionality • ca. GRID via the IVI Middleware software • Annotations and markup via the AIM project • Additional platforms (e. g. Linux, MAC) • Other data formats and functions requested in the RFP • XIP ‘look-and-feel’

Open Inventor • • • Open Inventor ® is an object-oriented 3 D toolkit offering a comprehensive solution to interactive graphics programming problems. URL: http: //oss. sgi. com/projects/inventor/ It presents a programming model based on the Model/View/Controller design pattern and the concept of Pipelines. Open Inventor: • is built on top of Open. GL® • defines a standard file format for 3 D data interchange • introduces a simple event model for 3 D interaction • provides animation objects called Engines • provides high performance object picking • is window system and platform independent • is a cross-platform 3 D graphics development system • encourages programmers to create new customized objects
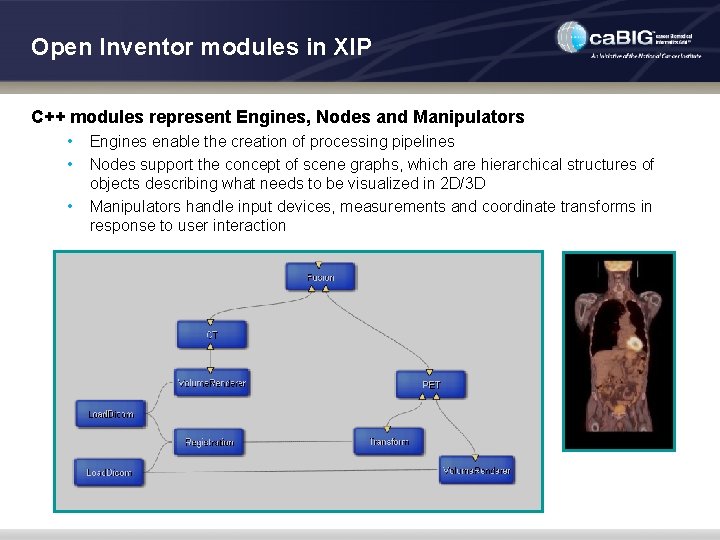
Open Inventor modules in XIP C++ modules represent Engines, Nodes and Manipulators • • • Engines enable the creation of processing pipelines Nodes support the concept of scene graphs, which are hierarchical structures of objects describing what needs to be visualized in 2 D/3 D Manipulators handle input devices, measurements and coordinate transforms in response to user interaction
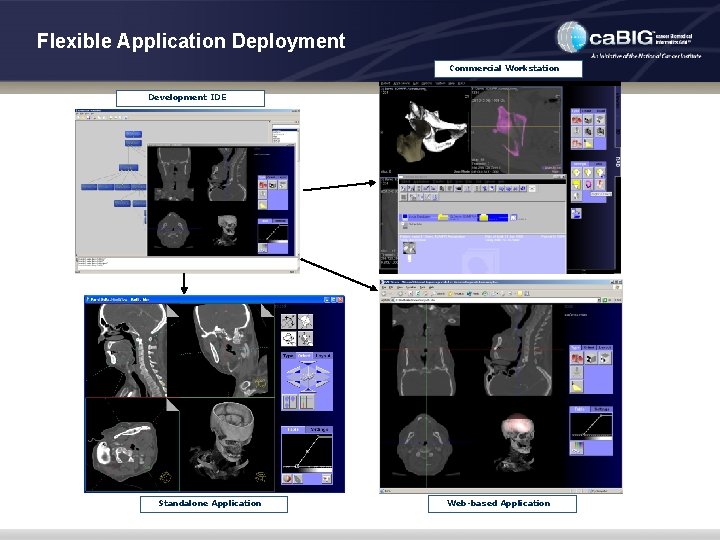
Flexible Application Deployment Commercial Workstation Development IDE Standalone Application Web-based Application
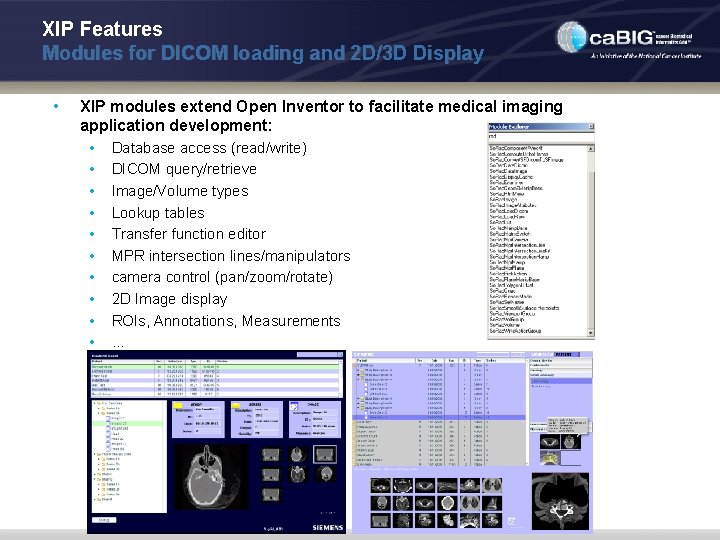
XIP Features Modules for DICOM loading and 2 D/3 D Display • XIP modules extend Open Inventor to facilitate medical imaging application development: • Database access (read/write) • DICOM query/retrieve • Image/Volume types • Lookup tables • Transfer function editor • MPR intersection lines/manipulators • camera control (pan/zoom/rotate) • 2 D Image display • ROIs, Annotations, Measurements • …

XIP Features Modules for Fused Volume Rendering • Features: • Volumes are stored separately and fused at rendering time, not as a preprocessing step • Support for unlimited number of fused large volumes • Each volume has independent control of: • Transformation (rot, trans, scale, shear…) • Color/opacity Transfer function • Crop box • Cut-planes • Rendering mode (VRT, MIP, Min. IP, DRR, SSD) • Voxel Resolution • Sampling rate • Performance • 20 frames/sec during interactivity • 1 sec for final diagnostic-quality update

XIP Features Modules for Client/Server Remote Visualization • • • Visual creation and configuration of distributed services Thin Client and Smart Client configurations are supported Support for ca. GRID remote grid computing services XIP modules for DICOM Query, Sorting, remote data transmission XIP can serialize the entire state to a file, thereby facilitating support for client/server state management and recovery as well as workflow management.
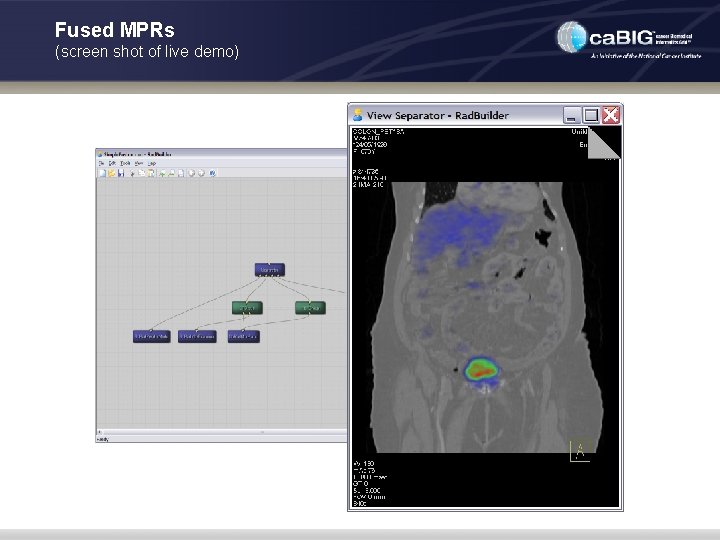
Fused MPRs (screen shot of live demo)
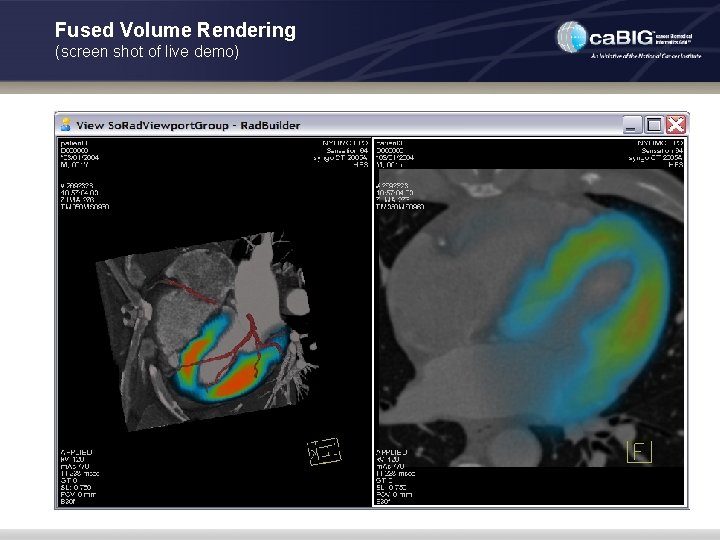
Fused Volume Rendering (screen shot of live demo)
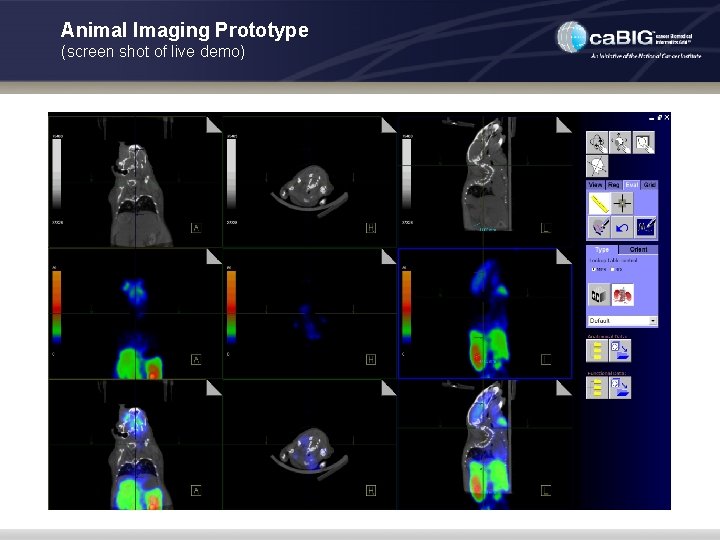
Animal Imaging Prototype (screen shot of live demo)
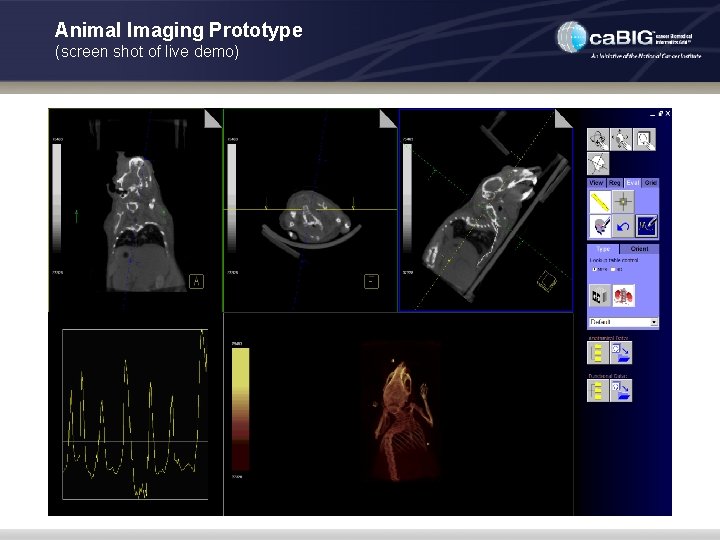
Animal Imaging Prototype (screen shot of live demo)
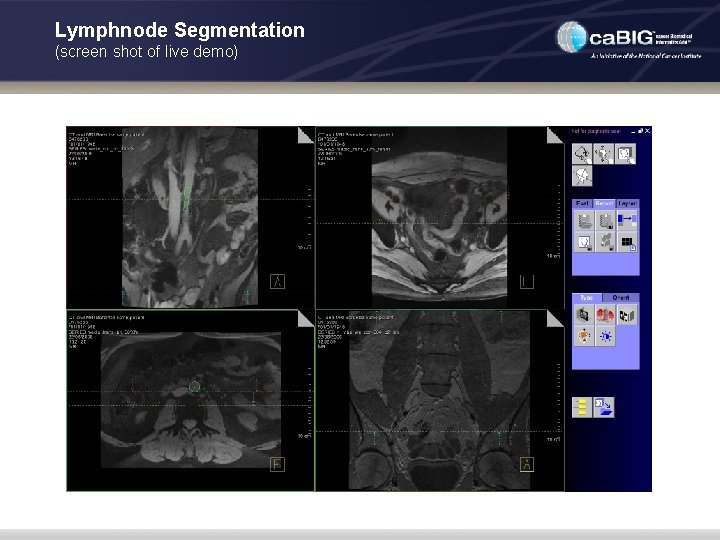
Lymphnode Segmentation (screen shot of live demo)
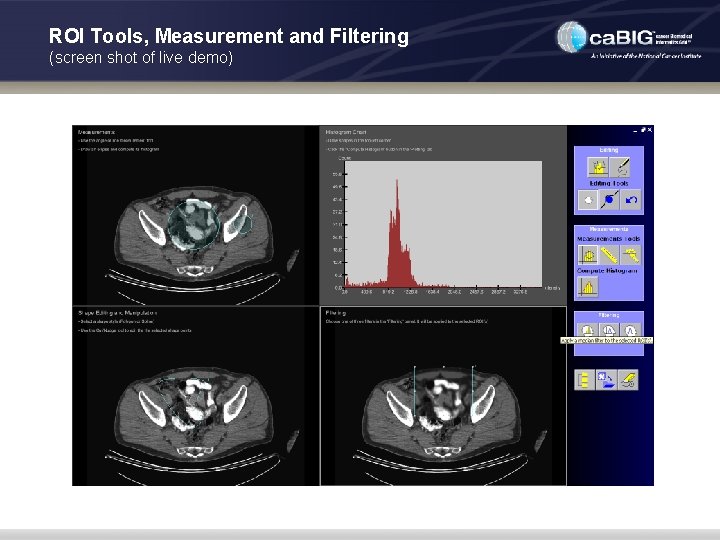
ROI Tools, Measurement and Filtering (screen shot of live demo)
4 D Beating Heart (screen shot of live demo)
ITK Demos: Level Set Networks (screen shot of live demo)

XIP Development Plan Phase 1 – Planning and RSNA/ca. BIG demonstrations (3 months) Planning done in parallel with RSNA/ca. BIG demonstration creation RSNA/ca. BIG demo done utilizing Siemens-internal proprietary development tools Source not released until phase 2. RSNA/ca. BIG demo serves as discussion point for generating use cases and requirements for ongoing development. Phase 2 – Conversion of base Siemens SW to open source (4 months) Largely utilizes the existing code pool, stripping out proprietary references and preparing the code for open source distribution Final output will be similar to, though not exactly the same as the RSNA/ca. BIG demonstration. May include some new additions, but may not fulfill all of the requirements listed in the SOW. Intermediate releases of documentation and code with partial feature sets during the course of conversion. Becomes the first prototype implementation Phase 3 – Addition of new features to XIP, creation of sample reference applications (3 months) Drafts of all documents available with all implemented features available at the end of this phase Intermediate releases available throughout the phase to foster discussion, review, and use. Prototype code with all implemented features at the end of this phase Phase 4 – Finalization of the year’s work products (2 months) No new features added after the start of this phase Bug fixes and documentation corrections as needed Final testing, review, and approvals

First Prototype Release – Early May Focused on the needs of the AIM Project Targeted toward continued internal development, not necessarily end users Will be available open source on the NCI’s g. Forge site for developers and the curious What is included: • Loading and display of stacks of 2 D DICOM images • From the ca. GRID via the IVI Middleware • From DICOM image archives • From the local file system • • • Processing by Open Inventor-wrapped ITK objects Graphics provided by stock Open Inventor objects Graphical medical image markup Simple measurement functions What is not included: • The GUI-based XIP Application Builder (not till summer) • The wrapper-generating toolkit for existing class libraries • 3 D and 4 D Rendering and VTK integration • Non-DICOM image formats • Workflow, Security, Audit, etc.

XIP Development Team • Washington University in St. Louis • • • Lawrence Tarbox Jaroslaw Krych David Maffitt Steve Moore Fred Prior Others • Other Consultants • • Siteman Cancer Center Kitware • Collaborative Projects • • IVI Middleware Annotation and Image Markup NCIA MIRC • Siemens Corporate Research • • • Gianluca Paladini Thomas Moeller Daphne Yu Klaus Engel John Pearson Others • NCI ca. BIG IVI • David Kupferschmid • Booz Allen Hamilton • Paul Mulhern
An Open Platform for Cancer Research
- Slides: 42