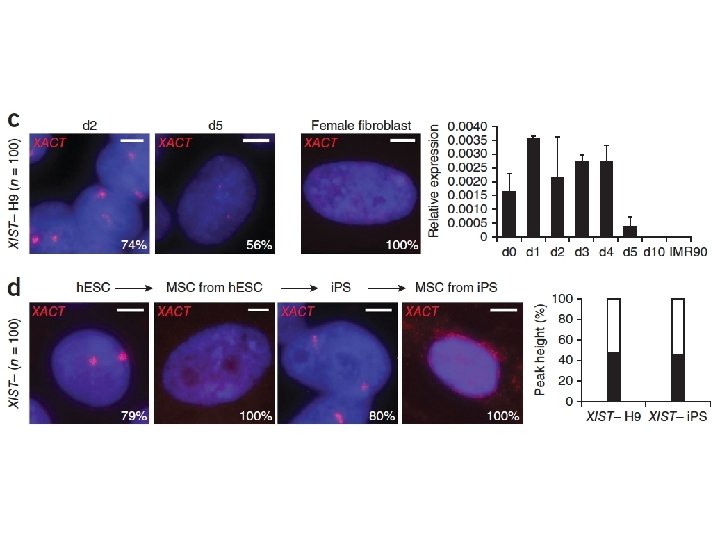
X Chromosome Inactivation Peters et al Nature Genetics






















- Slides: 58

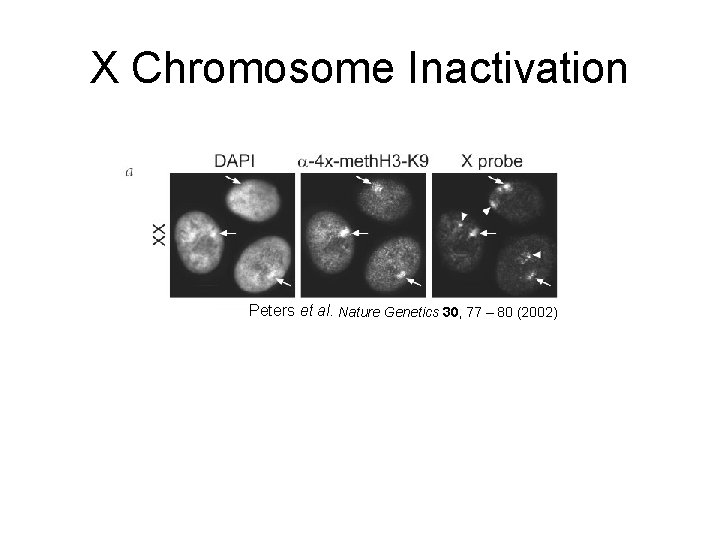
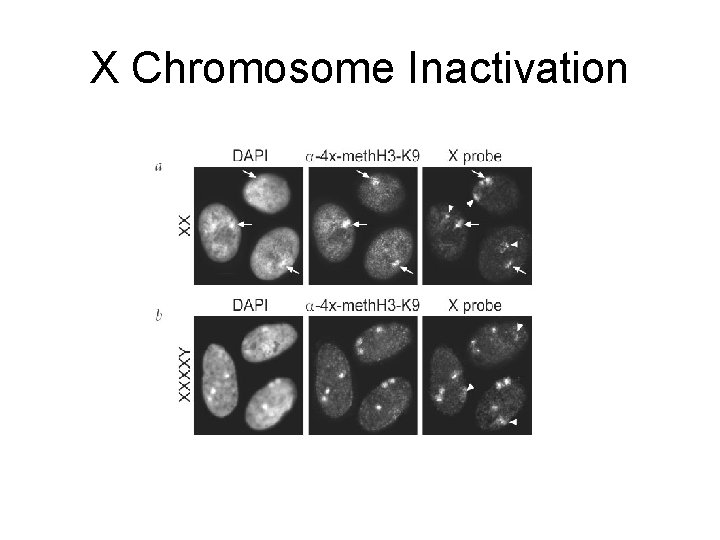
X Chromosome Inactivation Peters et al. Nature Genetics 30, 77 – 80 (2002)

X Chromosome Inactivation • X chromosome inactivation occurs early during development – around 24 cell • Thus, females embryos have two active X chromosomes until one is inactivated

X Chromosome Inactivation

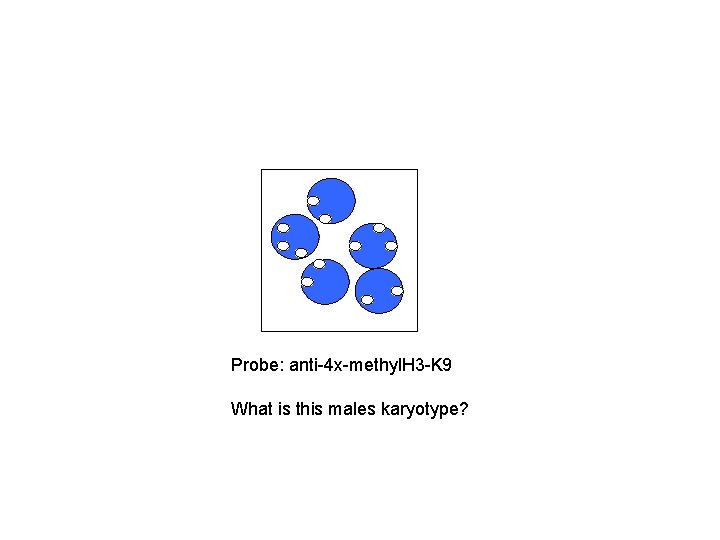
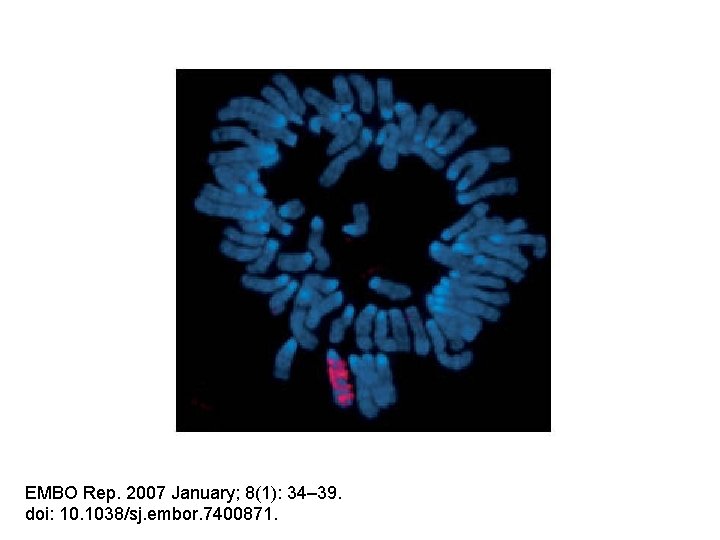
Probe: anti-4 x-methyl. H 3 -K 9 What is this males karyotype?

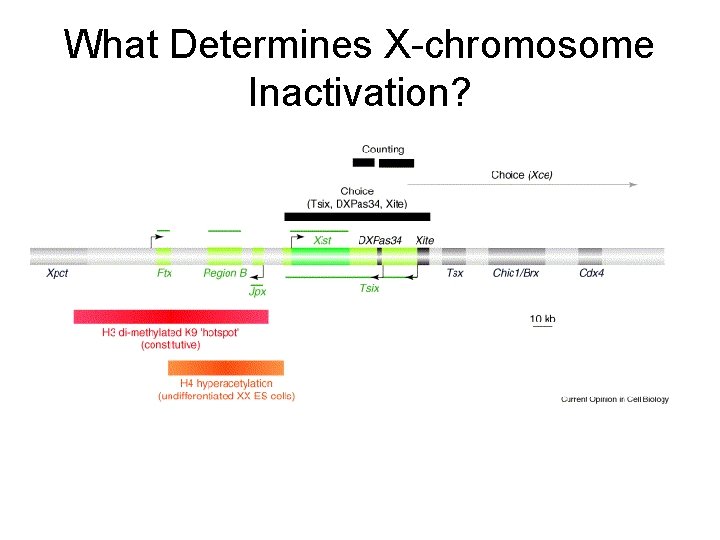
What Determines X-chromosome Inactivation?

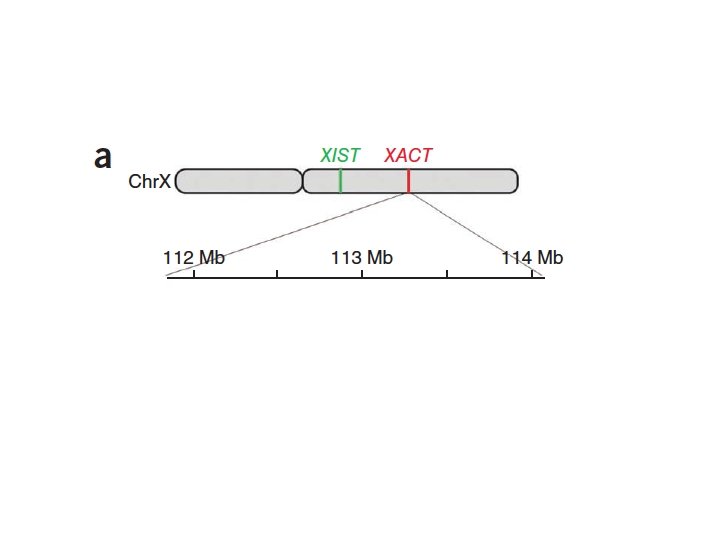
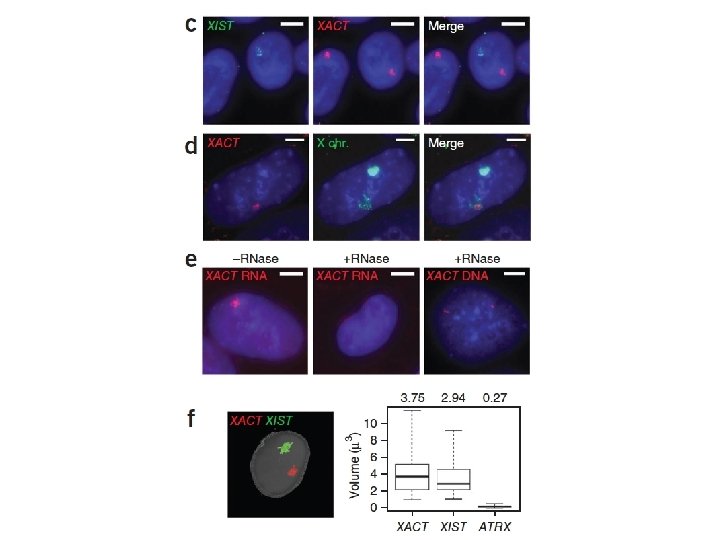
X Chromosome Inactivation • Mechanism of X Chromosome inactivation • XIC – X chromosome Inactivation Center • XIC controls expression of the XIST gene • XIST: X-inactive-specific transcript • XIST produces a non-coding 17 kb RNA molecule • “Coats” the entire local X-chromosome – cis-acting

EMBO Rep. 2007 January; 8(1): 34– 39. doi: 10. 1038/sj. embor. 7400871.

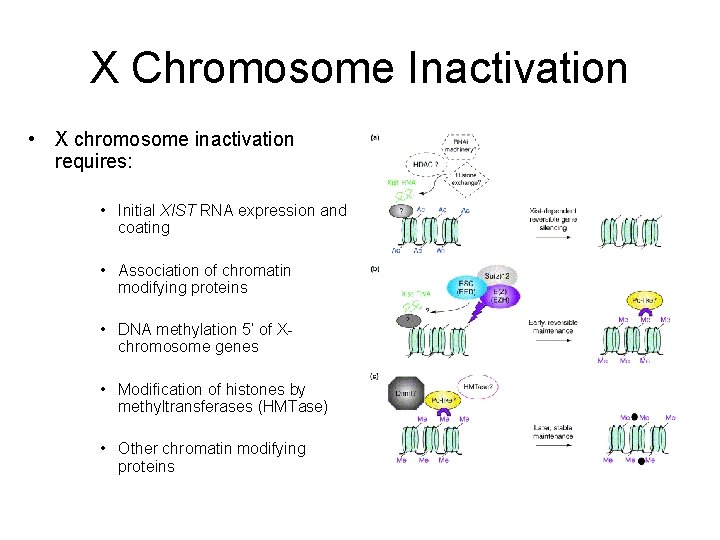
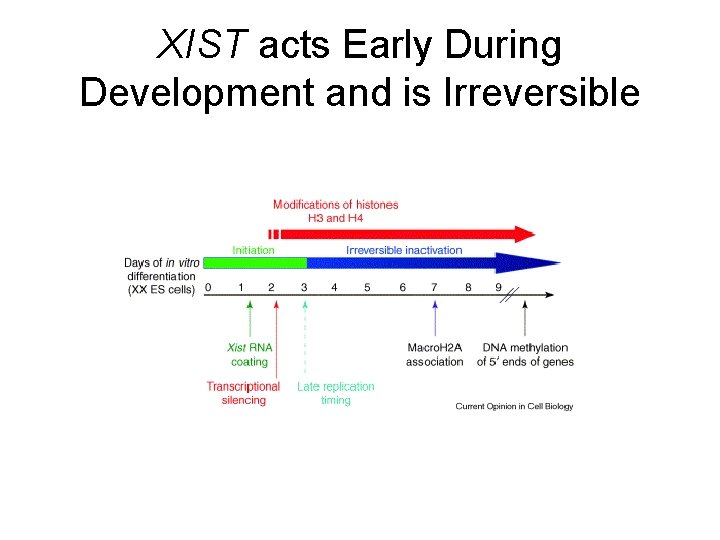
X Chromosome Inactivation • X chromosome inactivation requires: • Initial XIST RNA expression and coating • Association of chromatin modifying proteins • DNA methylation 5’ of Xchromosome genes • Modification of histones by methyltransferases (HMTase) • Other chromatin modifying proteins

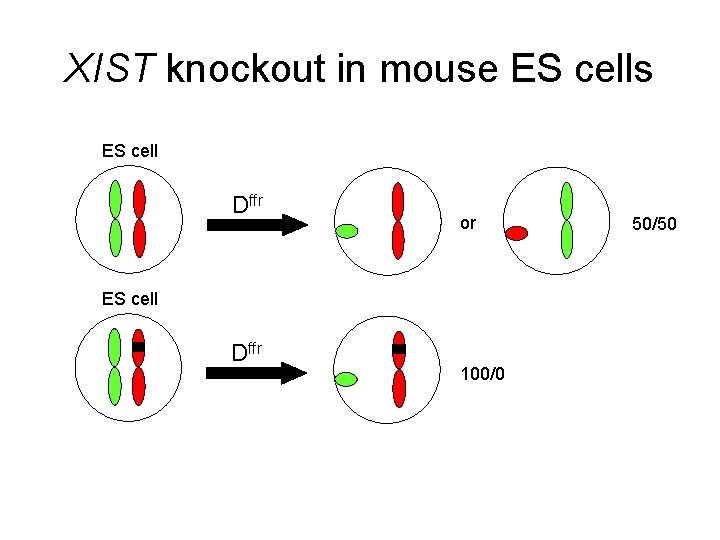
X Chromosome Inactivation • Approaches for examining XIST biology 1) Knock it out! Nature, January 1996

XIST knockout in mouse ES cells ES cell Dffr or ES cell Dffr 100/0 50/50

X Chromosome Inactivation • Approaches for examining XIST biology 2) Knock it in!

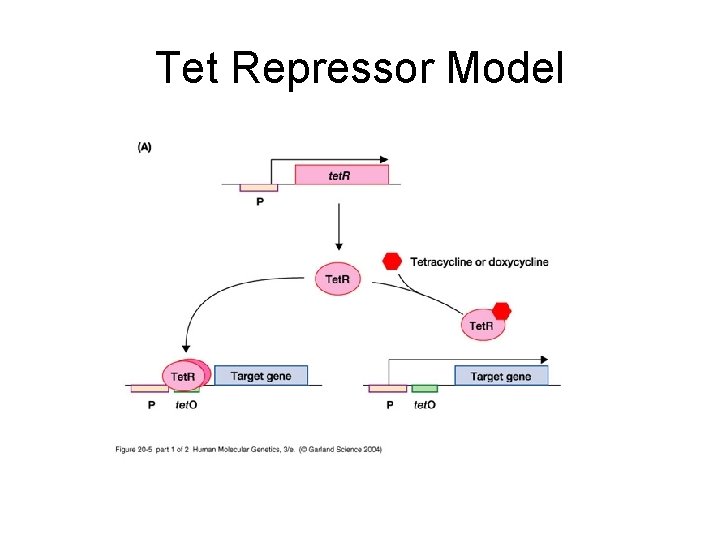
Tet Repressor Model

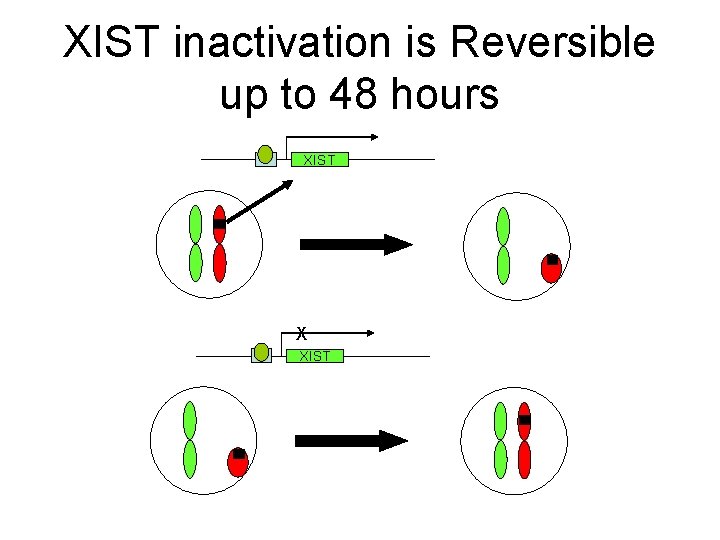
XIST inactivation is Reversible up to 48 hours XIST X XIST

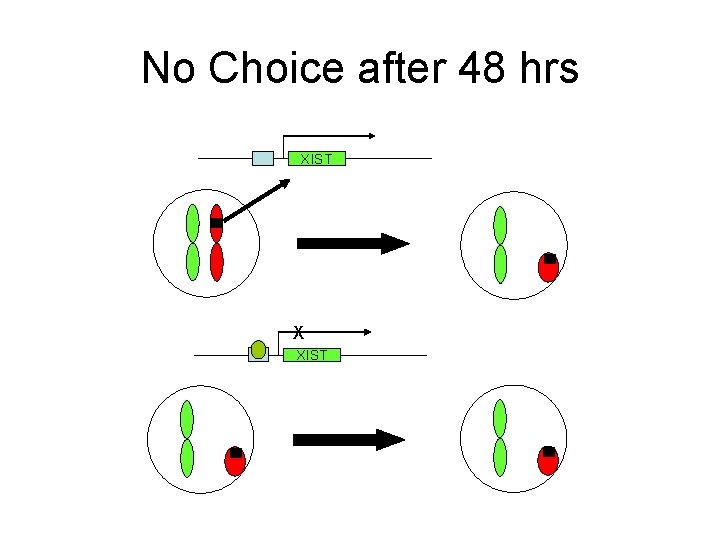
No Choice after 48 hrs XIST X XIST

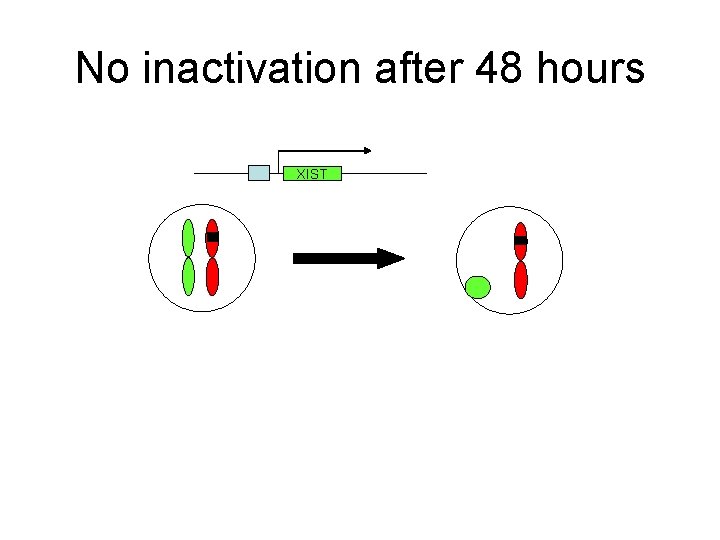
No inactivation after 48 hours XIST

XIST acts Early During Development and is Irreversible

What Controls XIST Expression?

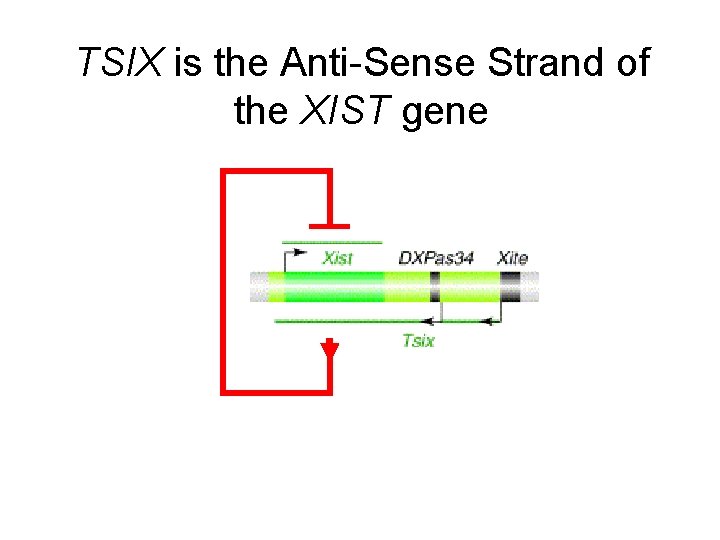
TSIX is the Anti-Sense Strand of the XIST gene

TSIX is the Anti-Sense Stand of the XIST gene

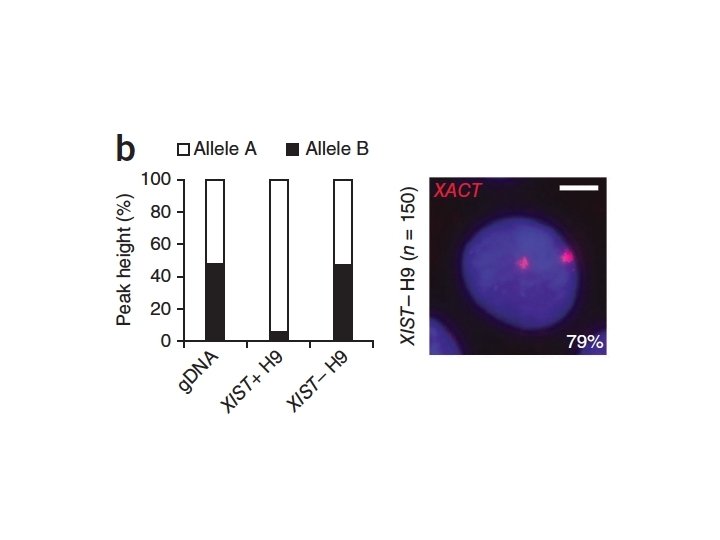
Knock-down of TSIX Causes Skewed X -Chromosome Inactivation X

TSIX Asymmetry Governs Choice • TSIX must be downregulated for XIST expression on the (future) inactivated X Chromosome • TSIX expression must remain for XIST downregulation on the (future) activated X Chromosome

Human Pathology • Without XIST, Human X Chromosome aneuploidy is Severe Molecular cytogenetic characterisation of a small ring X chromosome in a Turner patient and in a male patient with congenital abnormalities: role of X inactivation. Callen DF, Eyre HJ, Dolman G, Garry-Battersby MB, Mc. Creanor JR, Valeba A, Mc. Gill JJ. J Med Genet. 1995 Feb; 32(2): 113 -6.






Ubiquitin – Amino Acid Conservation

Ubiquitin – Nucleotide Conservation * * * * *

Amino Acid Conservation in Critical Domains

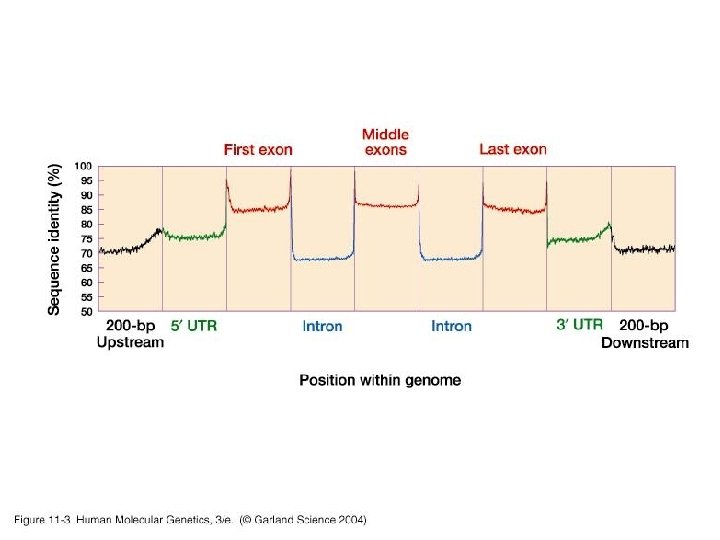
11_03. jpg

Errors in Protein Function • Eg. Cystic Fibrosis • Mutation causes loss-of-function • High occurrence of error may be a result of a heterozygote advantage Nature 393, 79 - 82 (07 May 1998) Salmonella typhi uses CFTR to enter intestinal epithelial cells GERALD B. PIER*, MARTHA GROUT*, TANWEER ZAIDI*, GLORIA MELULENI*, SIMONE S. MUESCHENBORN*, GEORGE BANTING†, ROSEMARY RATCLIFF‡, MARTIN J. EVANS§ & WILLIAM H. COLLEDGE‡

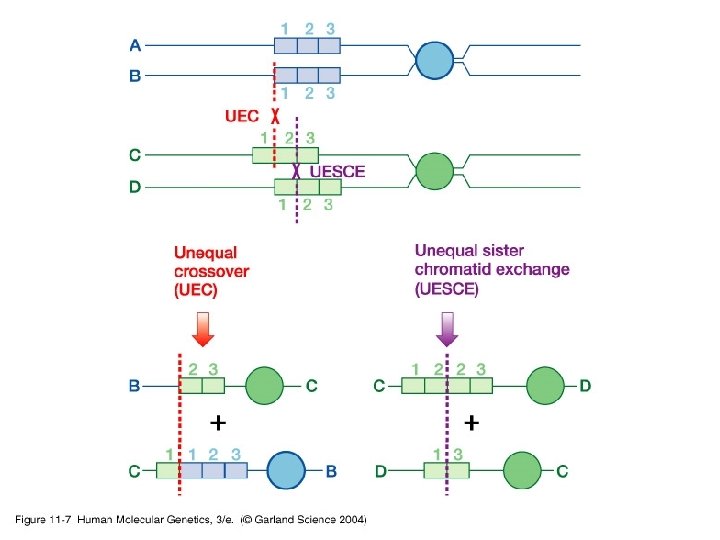
11_07. jpg

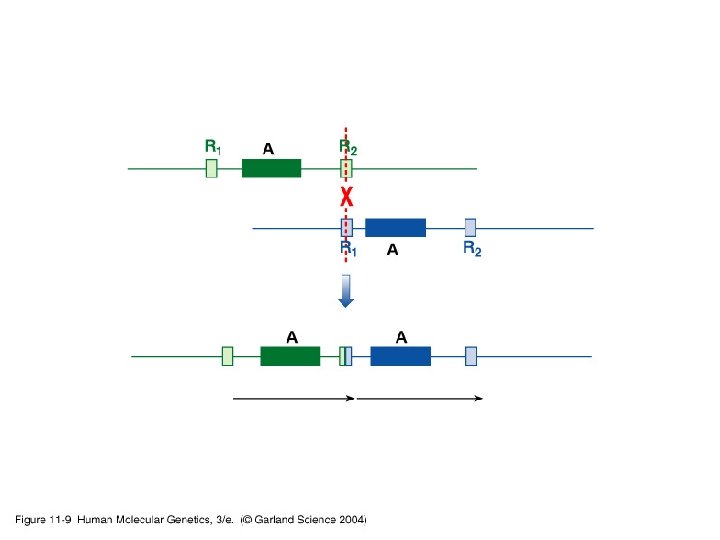
11_09. jpg

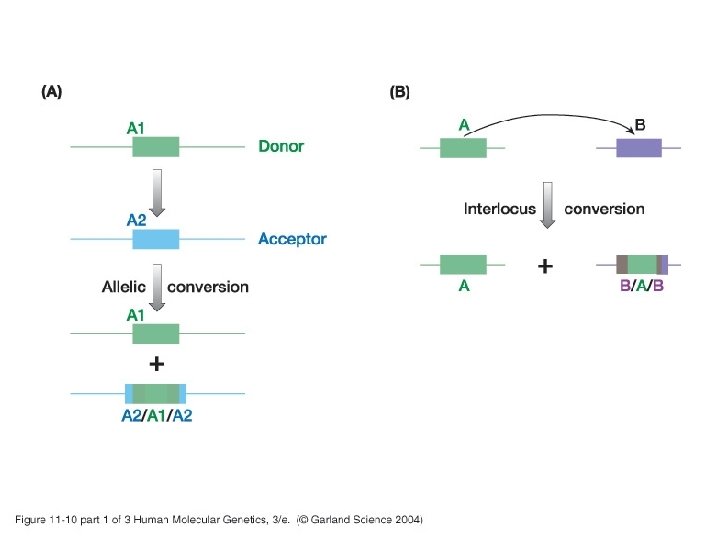
11_10. jpg

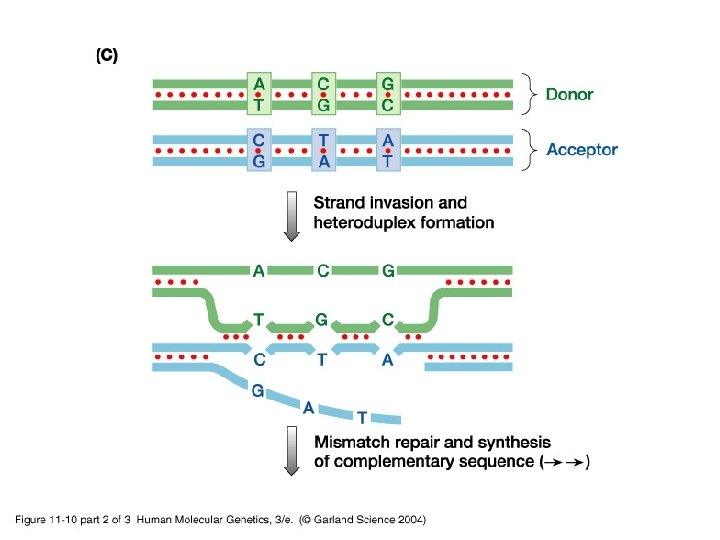
11_10_2. jpg

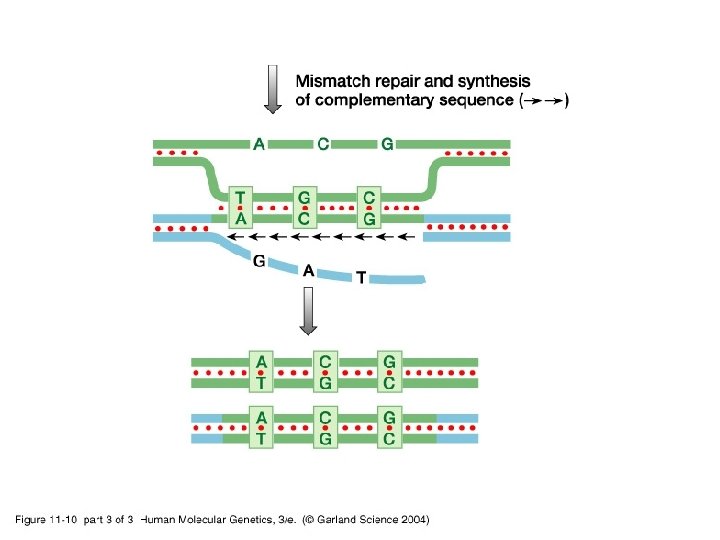
11_10_3. jpg

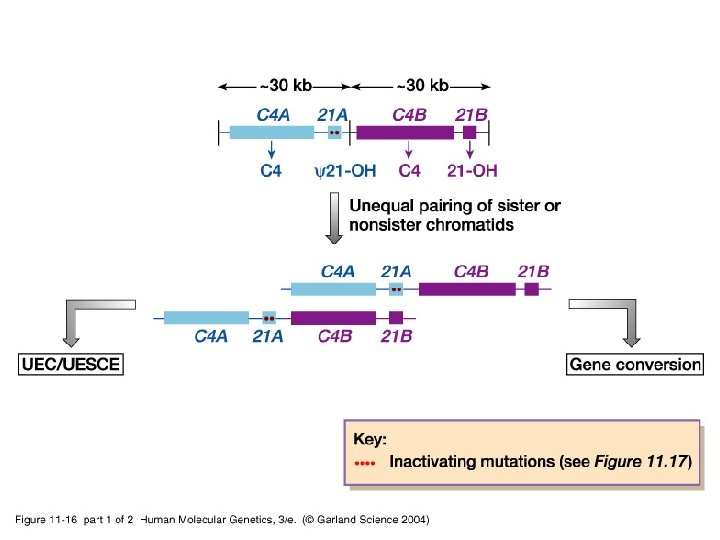
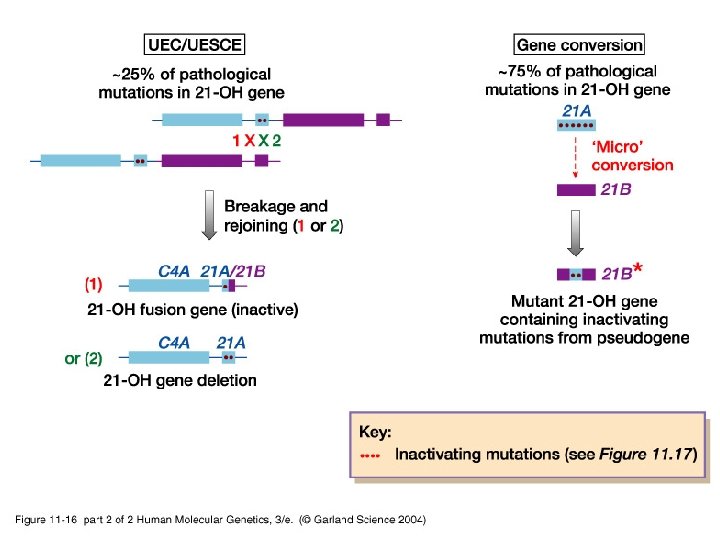
11_16. jpg

11_16_2. jpg

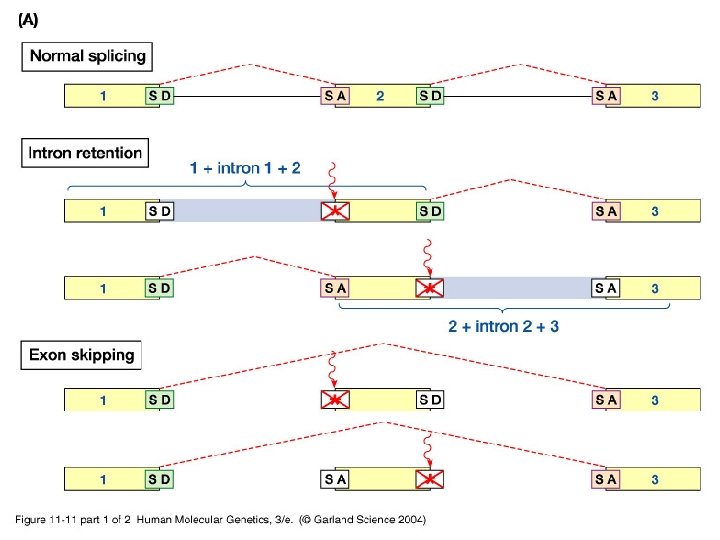
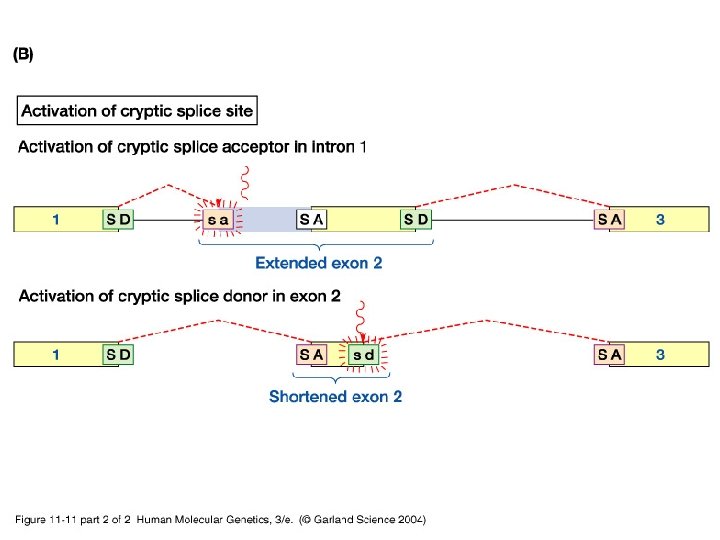
11_11. jpg

11_11_2. jpg

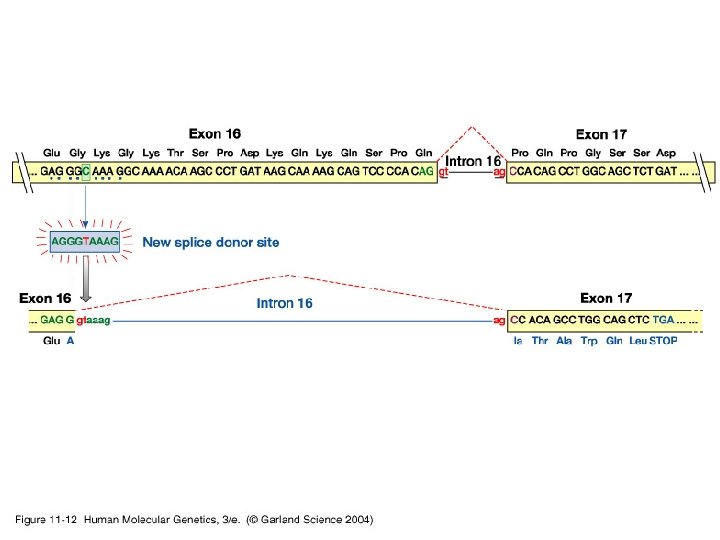
11_12. jpg

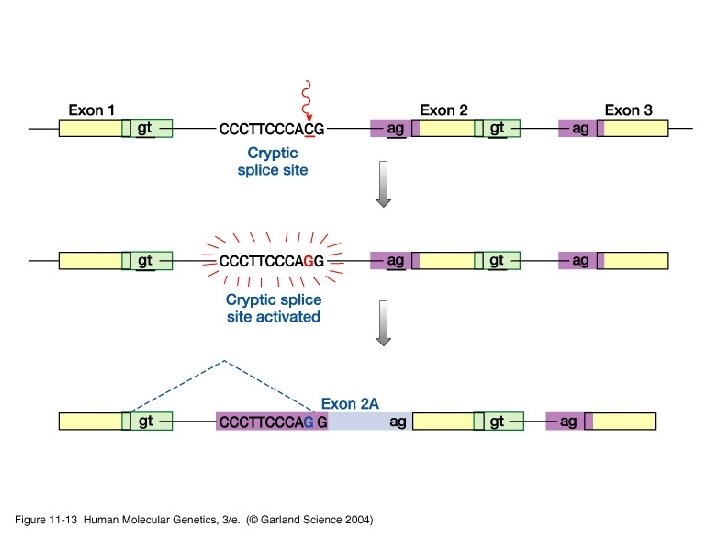
11_13. jpg

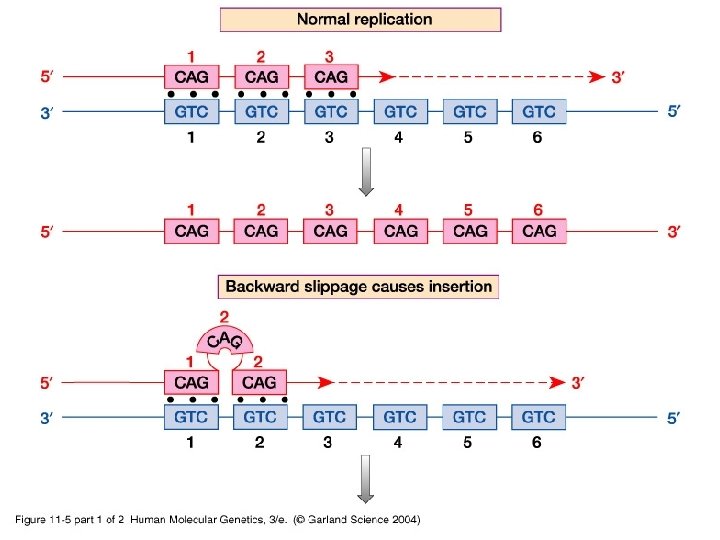
11_05. jpg

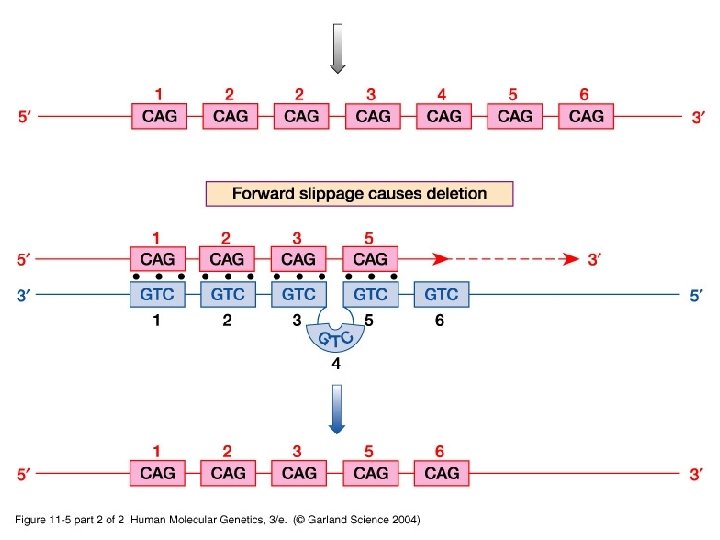
11_05_2. jpg


Huntington’s Disease • CAG repeat codes for glutamine (Q) • poly. Q located near the N-terminus of Huntingtin protein • Expansion in the coding region of the gene (unlike, for eg. FMR 1 – Fragile X syndrome - expansion is in 5’ UTR )

Huntington’s Disease MATLEKLMKA QPLLPQPQPP PEFQKLLGIA APRSLRAALW FGNFANDNEI LVPVEDEHST ELTLHHTQHQ SIVELIAGGG AASSGVSTPG FESLKSFQQQ QQQQQ PPPPGP AVAEEPLHRP MELFLLCSDD AESDVRMVAD RFAELAHLVR PQKCRPYLVN KVLLKAFIAN LKSSSPTIRR LLILGVLLTL RYLVPLLQQQ DHNVVTGALE LLQQLFRTPP SSCSPVLSRK QKGKVLLGEE SAGHDIITE……… QQQQQ KKELSATKKD ECLNKVIKAL LLPCLTRTSK TAAGSAVSIC VKDTSLKGSF PELLQTLTAV EALEDDSESR PPPPP RVNHCLTICE MDSNLPRLQL RPEESVQETL QHSRRTQYFY GVTRKEMEVS GGIGQLTAAK SDVSSSALTA PQLPQPPPQA NIVAQSVRNS ELYKEIKKNG AAAVPKIMAS SWLLNVLLGL PSAEQLVQVY EESGGRSRSG SVKDEISGEL MATLEKLMKA QQQQQ AVAEEPLHRP AESDVRMVAD PQKCRPYLVN LKSSSPTIRR RYLVPLLQQQ LLQQLFRTPP QKGKVLLGEE FESLKSFQQQ QQQQQ KKELSATKKD ECLNKVIKAL LLPCLTRTSK TAAGSAVSIC VKDTSLKGSF PELLQTLTAV EALEDDSESR QQQQQ PQLPQPPPQA NIVAQSVRNS ELYKEIKKNG AAAVPKIMAS SWLLNVLLGL PSAEQLVQVY EESGGRSRSG SVKDEISGEL QQQQQ QPLLPQPQPP PEFQKLLGIA APRSLRAALW FGNFANDNEI LVPVEDEHST ELTLHHTQHQ SIVELIAGGG AASSGVSTPG QQQQQ PPPPGP MELFLLCSDD RFAELAHLVR KVLLKAFIAN LLILGVLLTL DHNVVTGALE SSCSPVLSRK SAGHDIITE… QQQQQ PPPPP RVNHCLTICE MDSNLPRLQL RPEESVQETL QHSRRTQYFY GVTRKEMEVS GGIGQLTAAK SDVSSSALTA

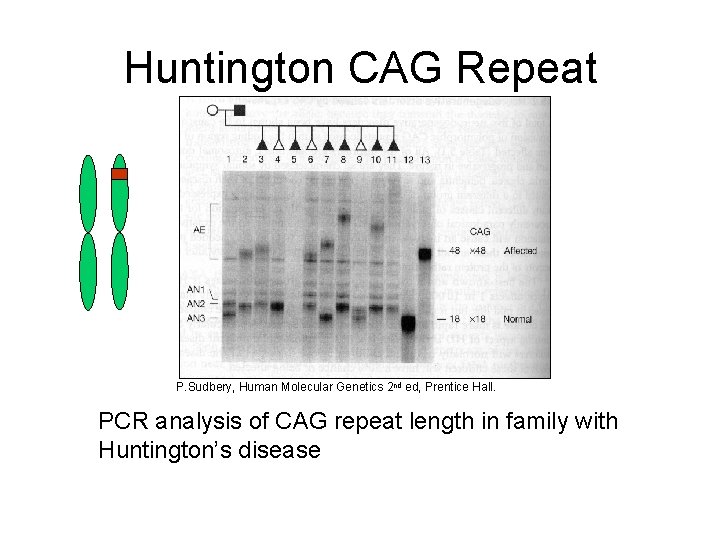
Huntington CAG Repeat P. Sudbery, Human Molecular Genetics 2 nd ed, Prentice Hall. PCR analysis of CAG repeat length in family with Huntington’s disease

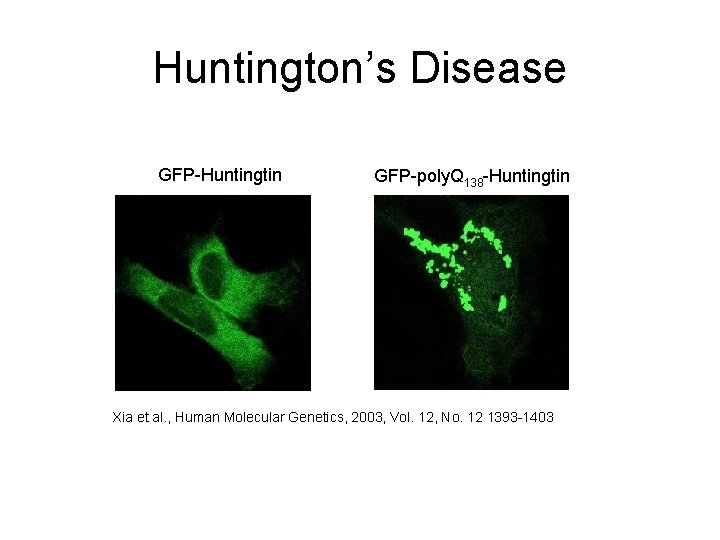
Huntington’s Disease GFP-Huntingtin GFP-poly. Q 138 -Huntingtin Xia et al. , Human Molecular Genetics, 2003, Vol. 12, No. 12 1393 -1403

Heterozygous knockouts are normal!

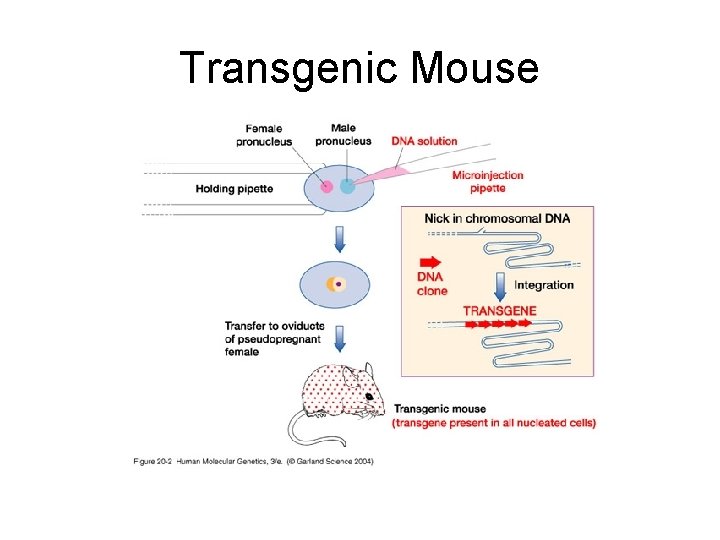
Transgenic Mouse






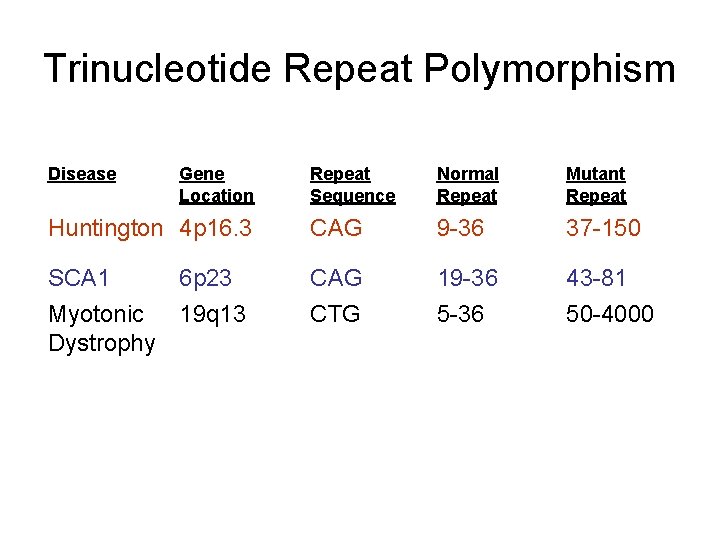
Trinucleotide Repeat Polymorphism Disease Gene Location Repeat Sequence Normal Repeat Mutant Repeat Huntington 4 p 16. 3 CAG 9 -36 37 -150 SCA 1 CAG 19 -36 43 -81 CTG 5 -36 50 -4000 6 p 23 Myotonic 19 q 13 Dystrophy