WSSP Chapter 9 Determine ORF and BLASTP atttaccgtg
WSSP Chapter 9 Determine ORF and BLASTP atttaccgtg ttggattgaa attatcttgc atgagccagc tgatgagtat gatacagttt tccgtattaa taacgg ccggaaatag gatcccgatc atgattgctt caatatttt acttcaatga ttggttctaa gcattcgaat gcgtacccgt ttgattaata tttccatttc tgtcccagtt tttaattttc atttcttttg gttaaaaaat tcccagtctc ttgaatgctt
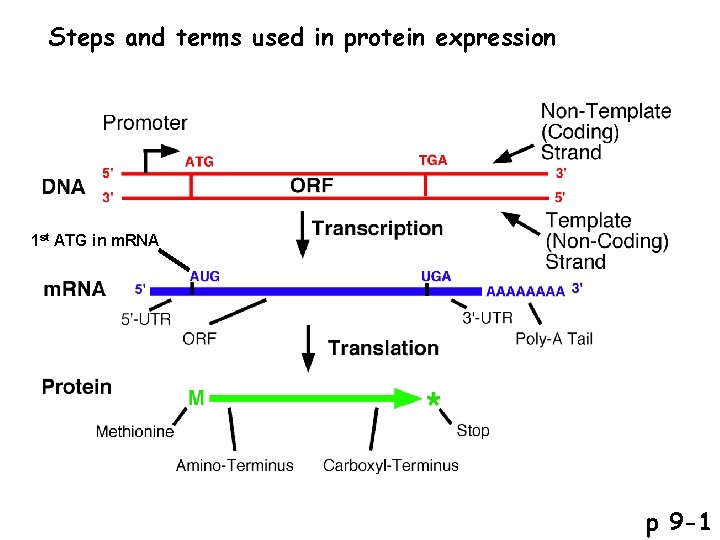
Steps and terms used in protein expression 1 st ATG in m. RNA p 9 -1

Cloning the c. DNA library p 9 -1
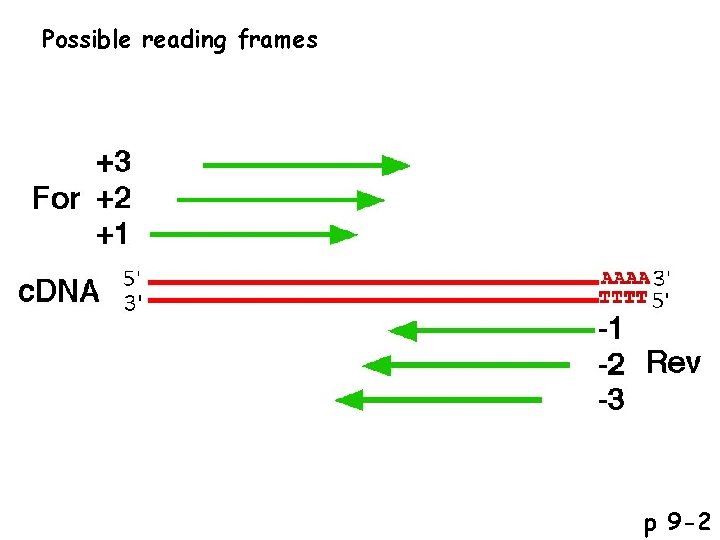
Possible reading frames p 9 -2
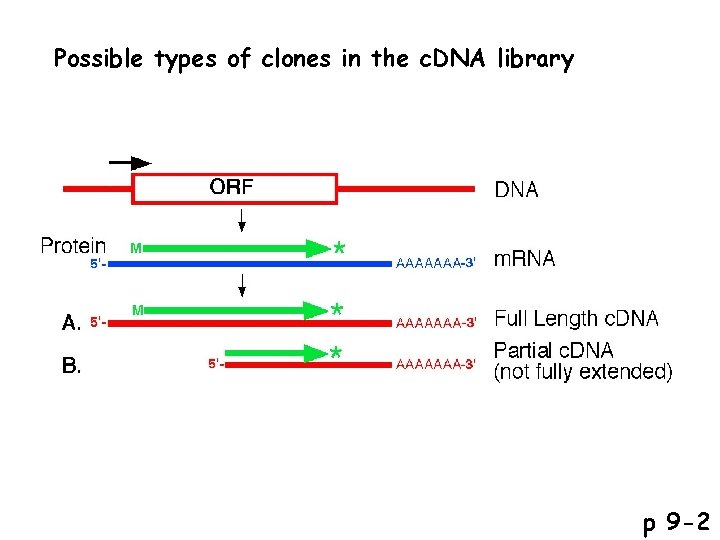
Possible types of clones in the c. DNA library p 9 -2
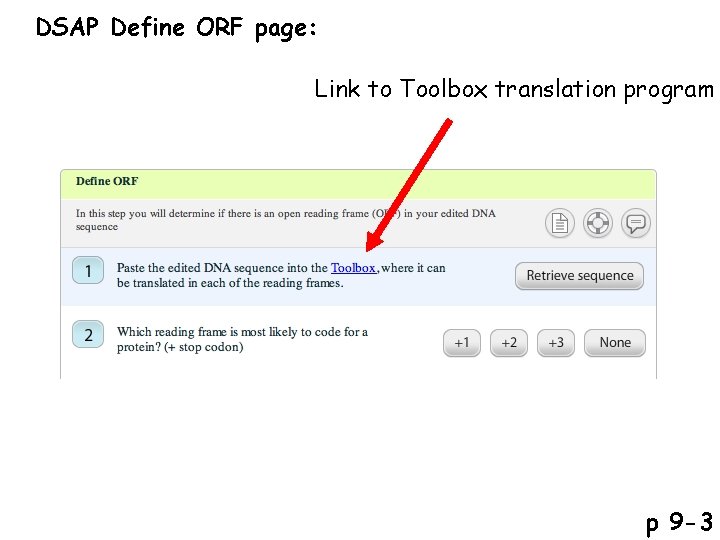
DSAP Define ORF page: Link to Toolbox translation program p 9 -3
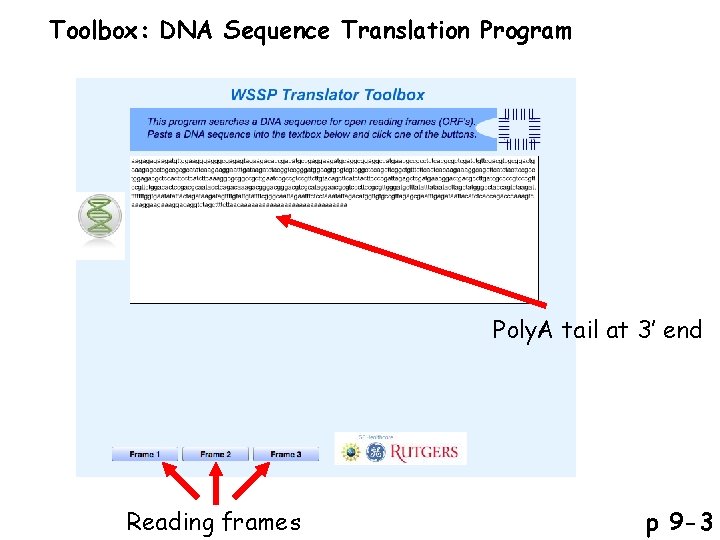
Toolbox: DNA Sequence Translation Program Poly. A tail at 3’ end Reading frames p 9 -3
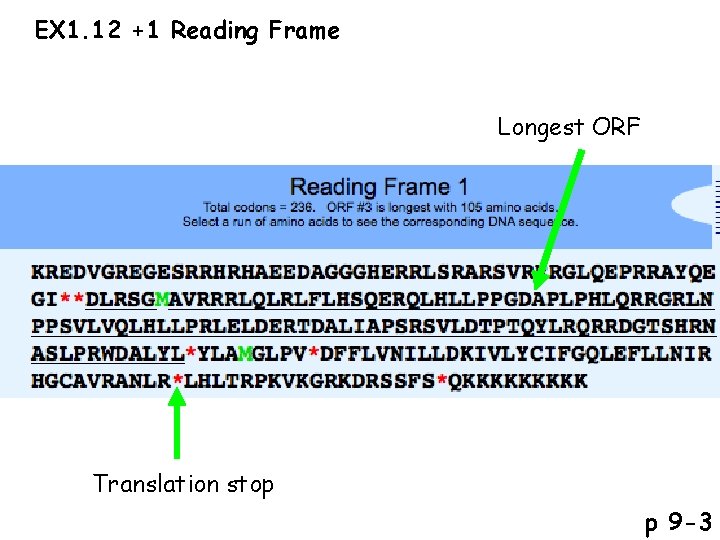
EX 1. 12 +1 Reading Frame Longest ORF Translation stop p 9 -3

Which one of these would be the correct ORF? A) B) Rule #1: If downstream of a stop codon, translation of the protein MUST start with an M (MET) p 9 -3
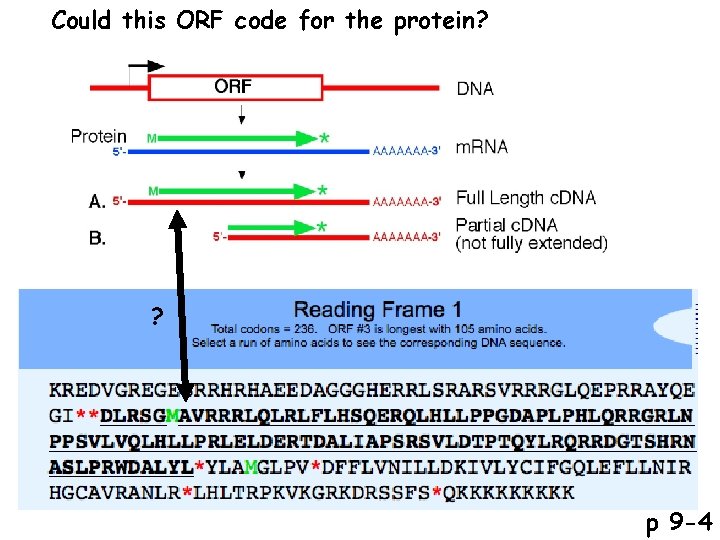
Could this ORF code for the protein? ? p 9 -4
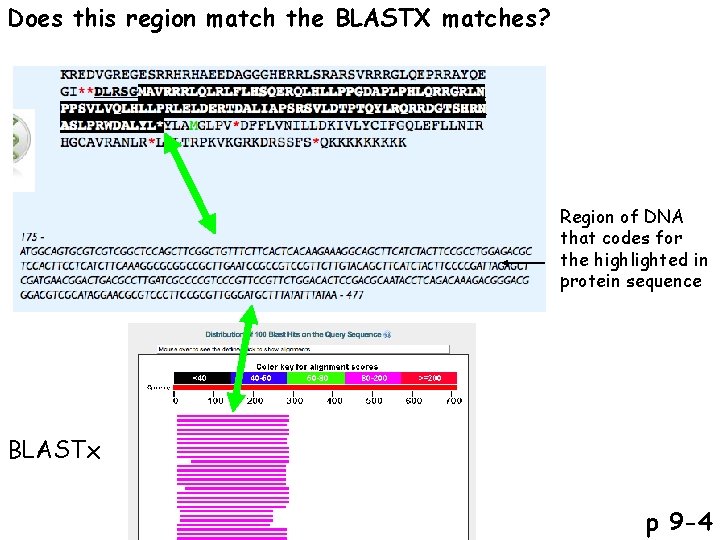
Does this region match the BLASTX matches? Region of DNA that codes for the highlighted in protein sequence BLASTx p 9 -4
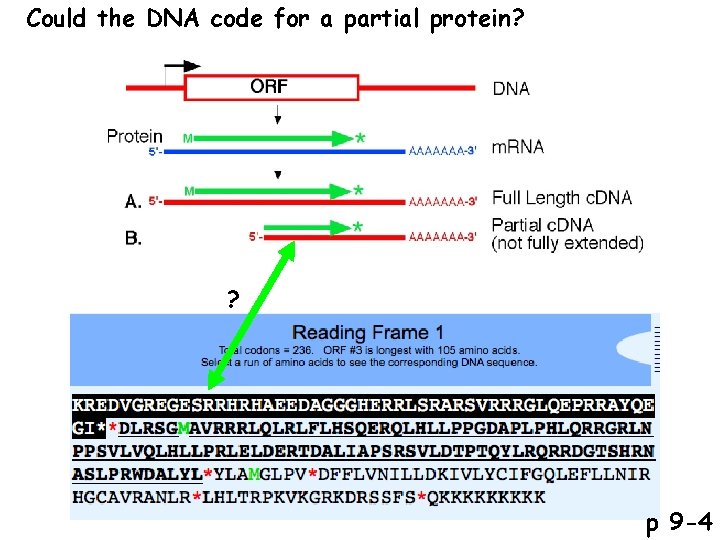
Could the DNA code for a partial protein? ? p 9 -4

Does this region match the BLASTX matches? Region of DNA that codes for the highlighted in protein sequence BLASTx p 9 -4

Does this region match the BLASTX matches? Region of DNA that codes for the highlighted in protein sequence BLASTx p 9 -4

An example of a partial coding sequence Similar Seq.
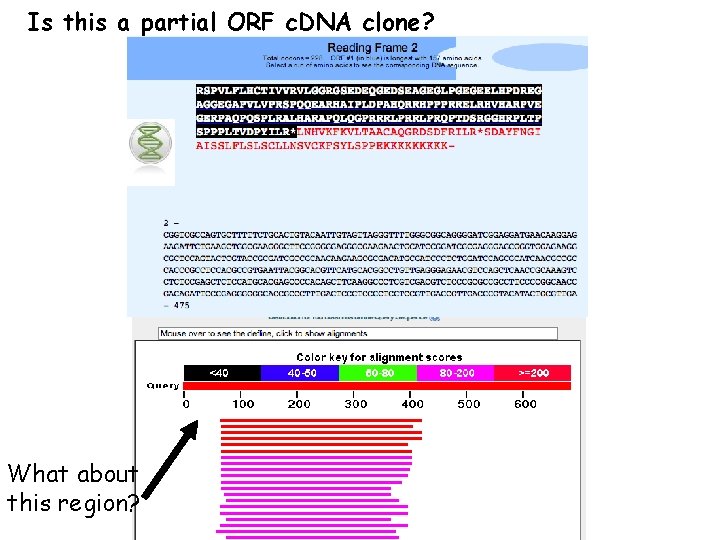
Is this a partial ORF c. DNA clone? What about this region?
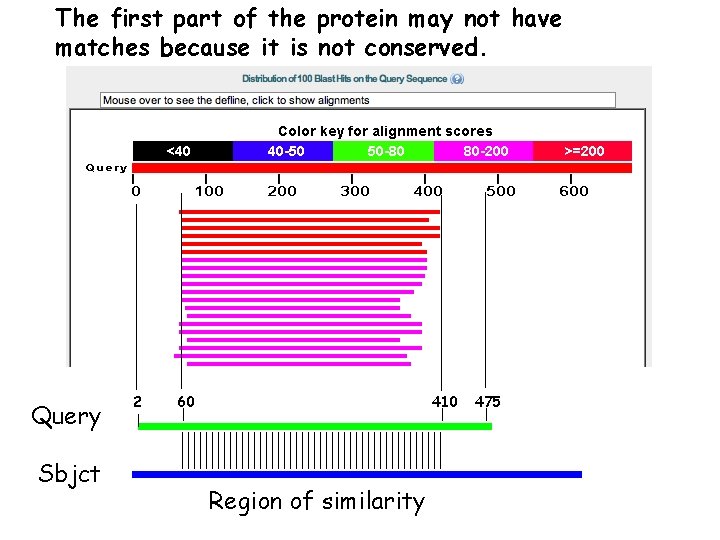
The first part of the protein may not have matches because it is not conserved. Query Sbjct 2 60 410 Region of similarity 475

The BLASTx helps determine which reading frame is correct >ref|NP_001150519. 1| dynein light chain LC 6, flagellar outer arm [Zea mays] Length=93 Score = 158 bits (400), Expect = 5 e-37 Identities = 73/93 (78%), Positives = 83/93 (89%), Gaps = 0/93 (0%) Frame = +2 Query 11 MLEGRARVEDTDMPRKMQAEAMNAASHALDLFDVADCKSLAAHIKKEFDKIYGPGWQCVV 190 MLEG+A VEDTDMP KMQA+AM+AAS ALD FDV DC+S+A+HIKKEFD I+GPGWQCVV Sbjct 1 MLEGKAVVEDTDMPAKMQAQAMSAASRALDRFDVLDCRSIASHIKKEFDAIHGPGWQCVV 60 Query 191 GSSFGCFFTHKKGSFIYFRLETLHFLIFKGAAA 289 GS FGC+ TH KGSFIYFRLE+L FL+FKGAAA Sbjct 61 GSGFGCYITHSKGSFIYFRLESLRFLVFKGAAA 93 It also helps suggest the start point p 9 -6
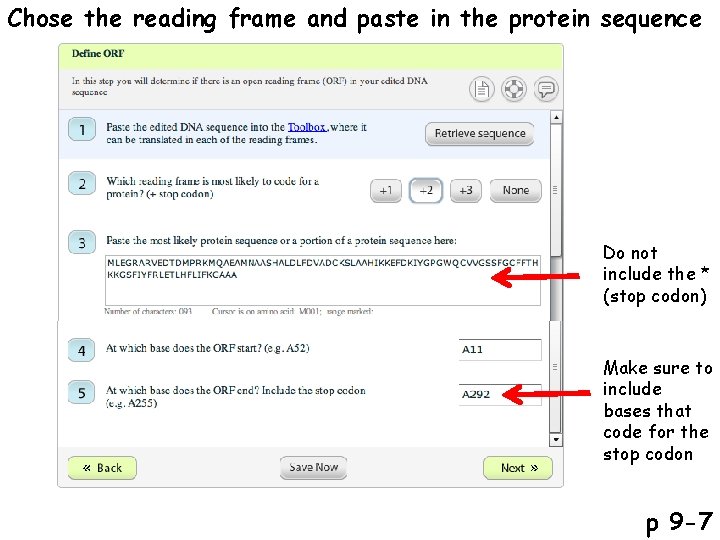
Chose the reading frame and paste in the protein sequence Do not include the * (stop codon) Make sure to include bases that code for the stop codon p 9 -7

The Five Commandments of DSAP I. The stop codon is part of the ORF
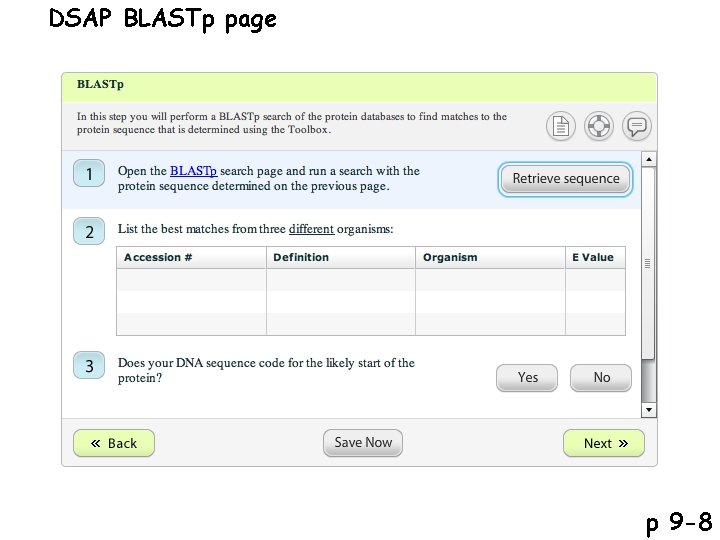
DSAP BLASTp page p 9 -8
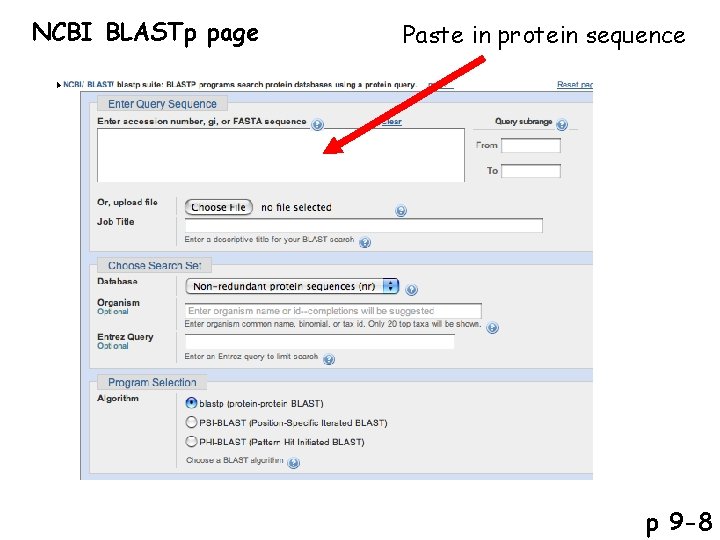
NCBI BLASTp page Paste in protein sequence p 9 -8
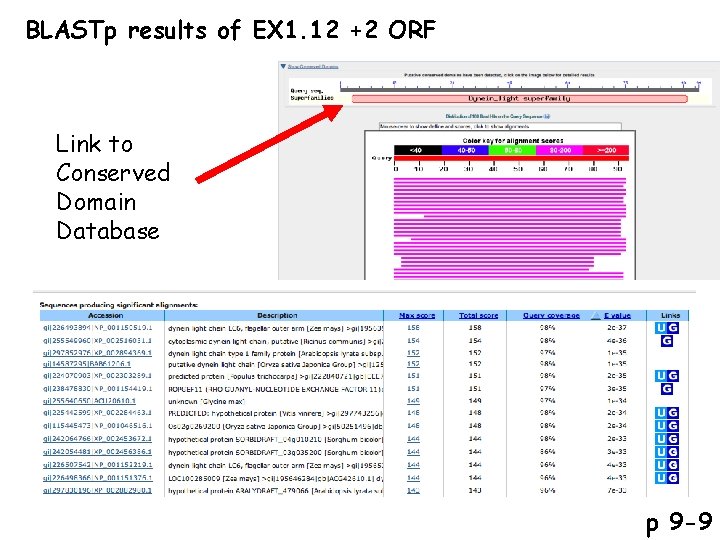
BLASTp results of EX 1. 12 +2 ORF Link to Conserved Domain Database p 9 -9
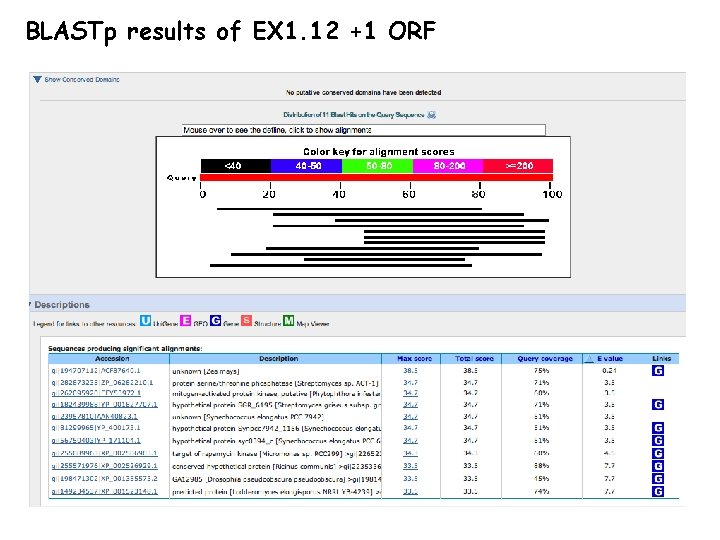
BLASTp results of EX 1. 12 +1 ORF
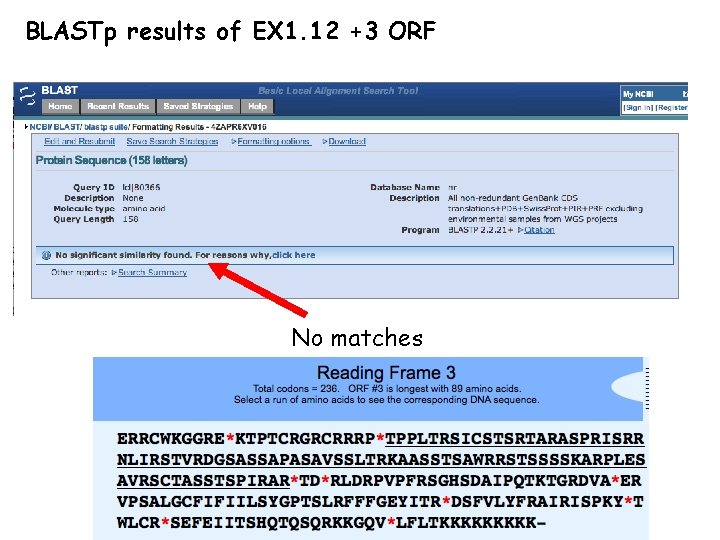
BLASTp results of EX 1. 12 +3 ORF No matches
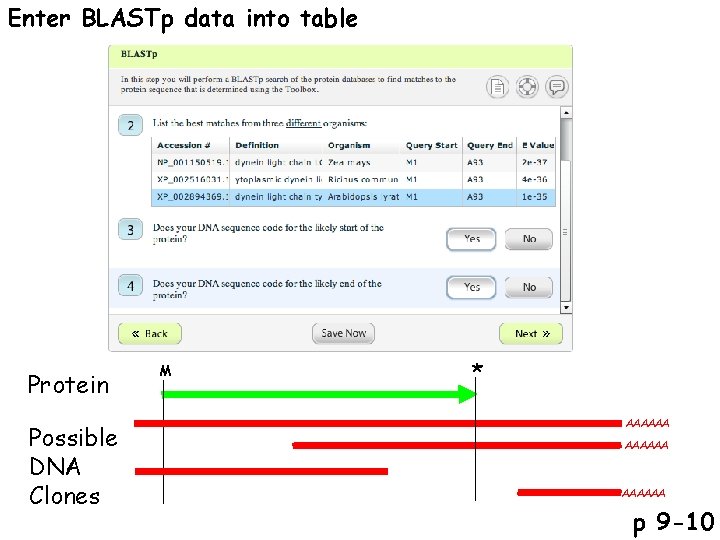
Enter BLASTp data into table Protein Possible DNA Clones M * AAAAAA p 9 -10

Suppose the c. DNA was missing the first 13 bp Does this DNA code for the start of the protein? >gi|226493894|ref|NP_001150519. 1| dynein light chain LC 6, flagellar outer arm [Zea mays] Length=93 Score = 139 bits (351), Expect = 8 e-32 Identities = 63/81 (77%), Positives = Query 1 MPRKMQAEAMNAASHALDLFDVADCKSLAAHIKKEFDKIYGPGWQCVVGSSFGCFFTHKK 60 MP KMQA+AM+AAS ALD FDV DC+S+A+HIKKEFD I+GPGWQCVVGS FGC+ TH K Sbjct 13 MPAKMQAQAMSAASRALDRFDVLDCRSIASHIKKEFDAIHGPGWQCVVGSGFGCYITHSK 72 Query 61 GSFIYFRLETLHFLIFKGAAA 81 GSFIYFRLE+L FL+FKGAAA Sbjct 73 GSFIYFRLESLRFLVFKGAAA 93
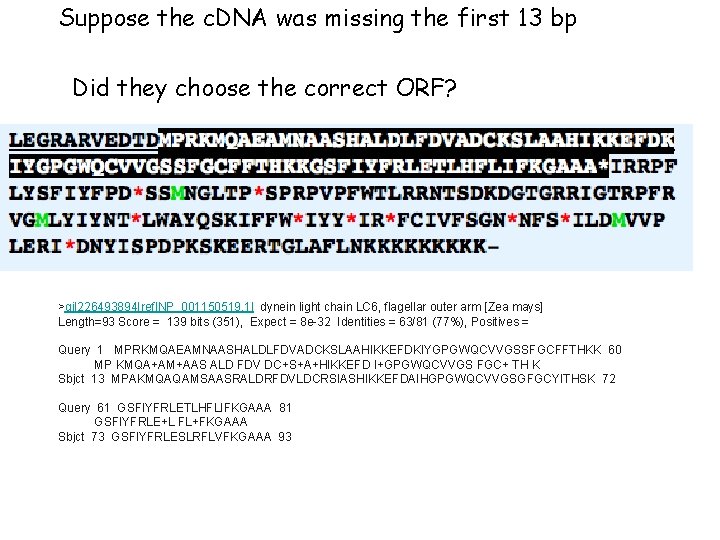
Suppose the c. DNA was missing the first 13 bp Did they choose the correct ORF? >gi|226493894|ref|NP_001150519. 1| dynein light chain LC 6, flagellar outer arm [Zea mays] Length=93 Score = 139 bits (351), Expect = 8 e-32 Identities = 63/81 (77%), Positives = Query 1 MPRKMQAEAMNAASHALDLFDVADCKSLAAHIKKEFDKIYGPGWQCVVGSSFGCFFTHKK 60 MP KMQA+AM+AAS ALD FDV DC+S+A+HIKKEFD I+GPGWQCVVGS FGC+ TH K Sbjct 13 MPAKMQAQAMSAASRALDRFDVLDCRSIASHIKKEFDAIHGPGWQCVVGSGFGCYITHSK 72 Query 61 GSFIYFRLETLHFLIFKGAAA 81 GSFIYFRLE+L FL+FKGAAA Sbjct 73 GSFIYFRLESLRFLVFKGAAA 93

Suppose the c. DNA was missing the first 13 bp Did they choose the correct ORF? BLASTP starting here >gi|226493894|ref|NP_001150519. 1| dynein light chain LC 6, flagellar outer arm [Zea mays] Length=93 Score = 139 bits (351), Expect = 8 e-32 Identities = 63/81 (77%), Positives = Query 1 MPRKMQAEAMNAASHALDLFDVADCKSLAAHIKKEFDKIYGPGWQCVVGSSFGCFFTHKK 60 MP KMQA+AM+AAS ALD FDV DC+S+A+HIKKEFD I+GPGWQCVVGS FGC+ TH K Sbjct 13 MPAKMQAQAMSAASRALDRFDVLDCRSIASHIKKEFDAIHGPGWQCVVGSGFGCYITHSK 72 Query 61 GSFIYFRLETLHFLIFKGAAA 81 GSFIYFRLE+L FL+FKGAAA Sbjct 73 GSFIYFRLESLRFLVFKGAAA 93 BLASTP starting here >gi|226493894|ref|NP_001150519. 1| dynein light chain LC 6, flagellar outer arm [Zea mays] Score = 156 bits (395), Expect = 6 e-37 Identities = 72/92 (78%), Positives = 82/92 (89%), Gaps = 0/92 (0%) Query 1 LEGRARVEDTDMPRKMQAEAMNAASHALDLFDVADCKSLAAHIKKEFDKIYGPGWQCVVG 60 LEG+A VEDTDMP KMQA+AM+AAS ALD FDV DC+S+A+HIKKEFD I+GPGWQCVVG Sbjct 2 LEGKAVVEDTDMPAKMQAQAMSAASRALDRFDVLDCRSIASHIKKEFDAIHGPGWQCVVG 61 Query 61 SSFGCFFTHKKGSFIYFRLETLHFLIFKGAAA 92 S FGC+ TH KGSFIYFRLE+L FL+FKGAAA Sbjct 62 SGFGCYITHSKGSFIYFRLESLRFLVFKGAAA 93
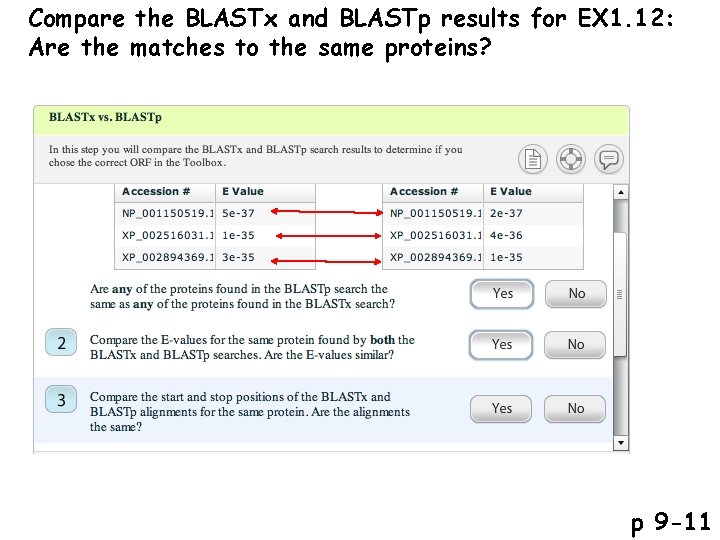
Compare the BLASTx and BLASTp results for EX 1. 12: Are the matches to the same proteins? p 9 -11
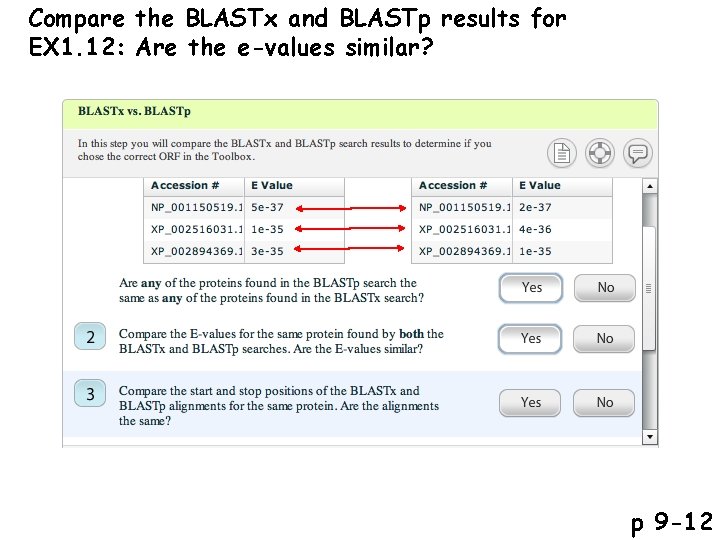
Compare the BLASTx and BLASTp results for EX 1. 12: Are the e-values similar? p 9 -12

Compare the BLASTx and BLASTp results for EX 1. 12: Are the alignments similar? BLASTx >ref|NP_001150519. 1| dynein light chain LC 6, flagellar outer arm [Zea mays] Length=93 Query 11 MLEGRARVEDTDMPRKMQAEAMNAASHALDLFDVADCKSLAAHIKKEFDKIYGPGWQCVV 190 MLEG+A VEDTDMP KMQA+AM+AAS ALD FDV DC+S+A+HIKKEFD I+GPGWQCVV Sbjct 1 MLEGKAVVEDTDMPAKMQAQAMSAASRALDRFDVLDCRSIASHIKKEFDAIHGPGWQCVV 60 Query 191 GSSFGCFFTHKKGSFIYFRLETLHFLIFKGAAA 289 GS FGC+ TH KGSFIYFRLE+L FL+FKGAAA Sbjct 61 GSGFGCYITHSKGSFIYFRLESLRFLVFKGAAA 93 BLASTp >gi|226493894|ref|NP_001150519. 1| dynein light chain LC 6, flagellar outer arm [Zea mays] Length=93 Score = 158 bits (400), Expect = 2 e-37 Query 1 MLEGRARVEDTDMPRKMQAEAMNAASHALDLFDVADCKSLAAHIKKEFDKIYGPGWQCVV 60 MLEG+A VEDTDMP KMQA+AM+AAS ALD FDV DC+S+A+HIKKEFD I+GPGWQCVV Sbjct 1 MLEGKAVVEDTDMPAKMQAQAMSAASRALDRFDVLDCRSIASHIKKEFDAIHGPGWQCVV 60 Query 61 GSSFGCFFTHKKGSFIYFRLETLHFLIFKGAAA 93 GS FGC+ TH KGSFIYFRLE+L FL+FKGAAA Sbjct 61 GSGFGCYITHSKGSFIYFRLESLRFLVFKGAAA 93 p 9 -12
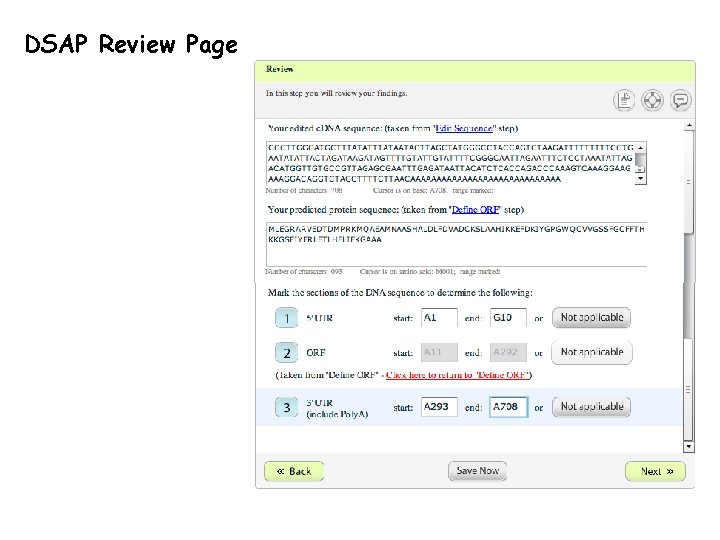
DSAP Review Page
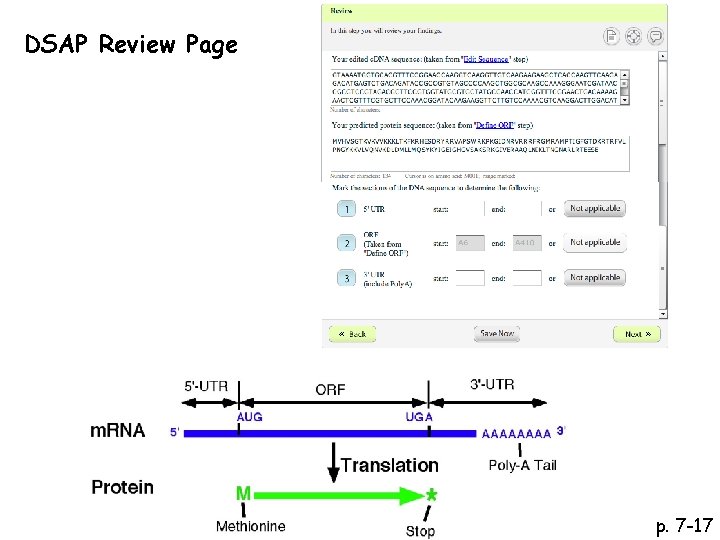
DSAP Review Page p. 7 -17
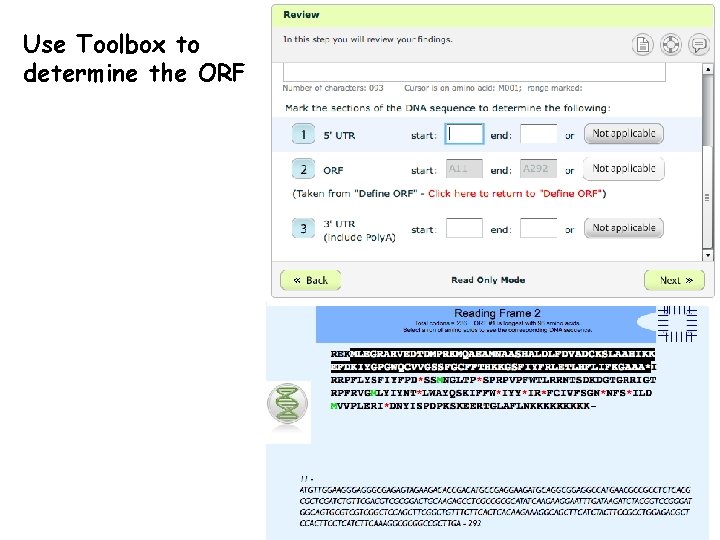
Use Toolbox to determine the ORF
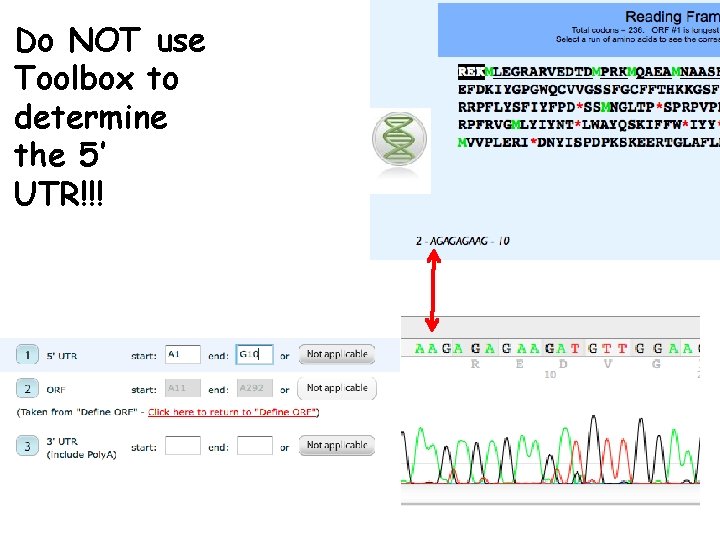
Do NOT use Toolbox to determine the 5’ UTR!!!

The Five Commandments of DSAP I. The stop codon is part of the ORF II. The start of the 5’ UTR is always the first base
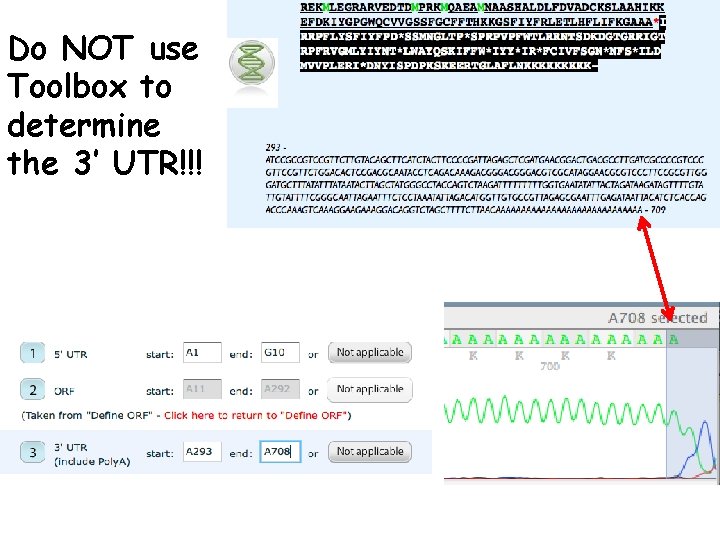
Do NOT use Toolbox to determine the 3’ UTR!!!
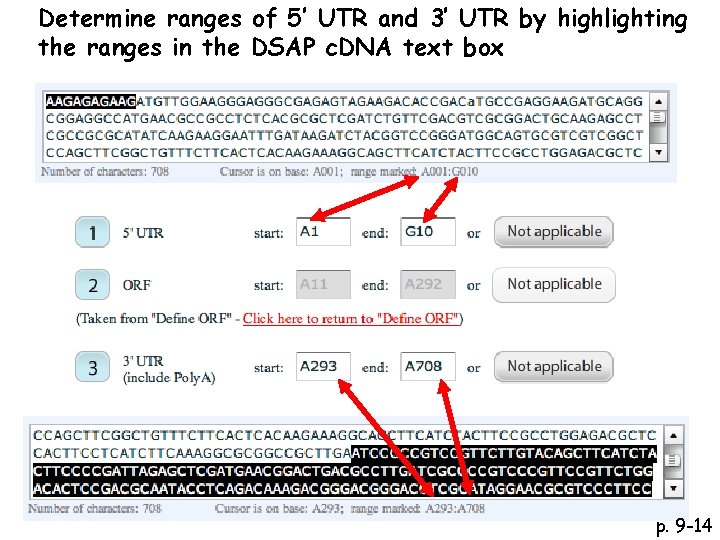
Determine ranges of 5’ UTR and 3’ UTR by highlighting the ranges in the DSAP c. DNA text box p. 9 -14
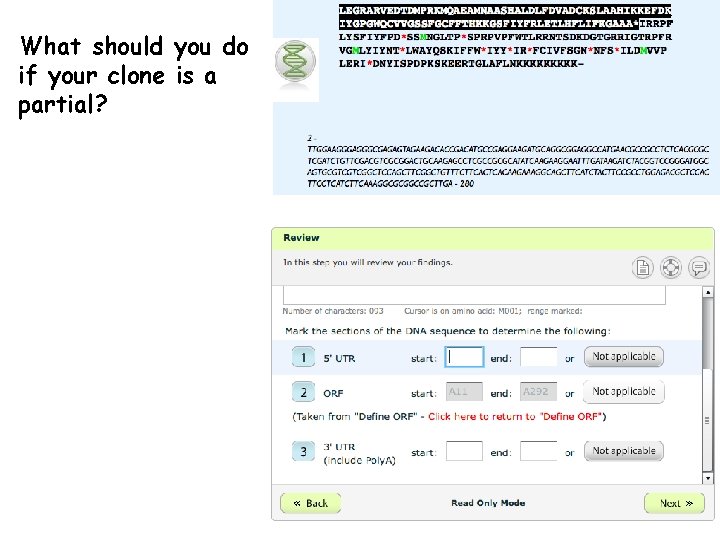
What should you do if your clone is a partial?

An example of a partial coding sequence Similar Seq.

The first bases are part of the reading frame ? S I R XGC TCA ATC CGT

The Five Commandments of DSAP I. The stop codon is part of the ORF II. The start of the 5’ UTR is always the first base III. If the clone is a partial, there is no 5’ UTR IV. If the clone is a partial, the start of the ORF is always the first base
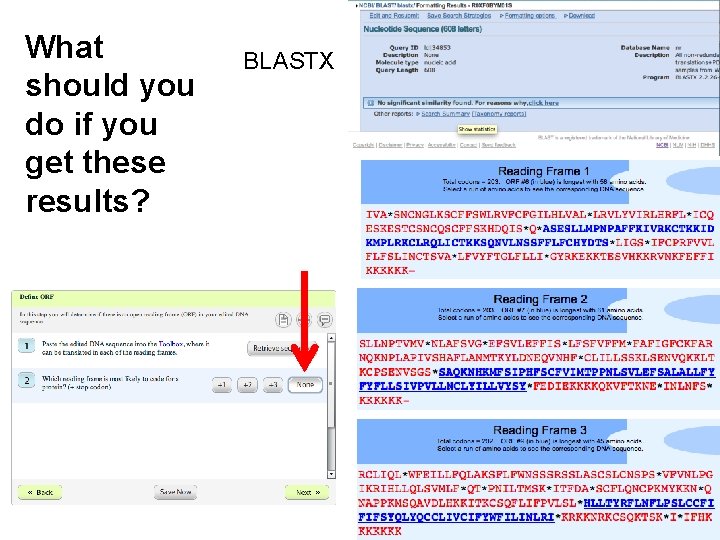
What should you do if you get these results? BLASTX

Why is my c. DNA noncoding? Genomic DNA RNA c. DNA (Partial) ORF AAAAAAA Recent genome wide RNA sequence studies show that more than 10% of poly. A RNAs are non-coding
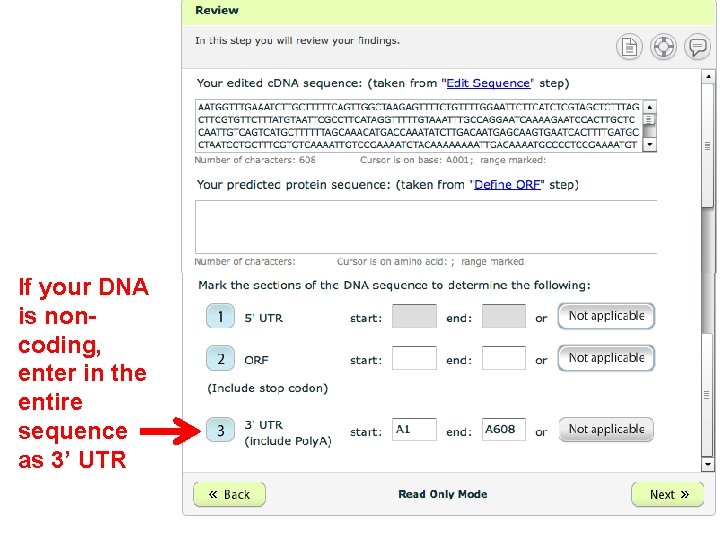
If your DNA is noncoding, enter in the entire sequence as 3’ UTR

The Five Commandments of DSAP I. The stop codon is part of the ORF II. The start of the 5’ UTR is always the first base III. If the clone is a partial, there is no 5’ UTR IV. If the clone is a partial, the start of the ORF is always the first base V. If the clone is non-coding, the entire DNA 3’ UTR

The Five Commandments of DSAP I. The stop codon is part of the ORF II. The start of the 5’ UTR is always the first base III. If the clone is a partial, there is no 5’ UTR. IV. If the clone is a partial, the start of the ORF is always the first base. V. If the clone is non-coding, the entire DNA 3’ UTR
- Slides: 49