Wheat Gene Expression Browser Leveraging 1000 publicly available
Wheat Gene Expression Browser Leveraging >1000 publicly available RNA-Seq samples for the wheat research and breeding community Cristobal Uauy (cristobal. uauy@jic. ac. uk) @Cristobal. Uauy
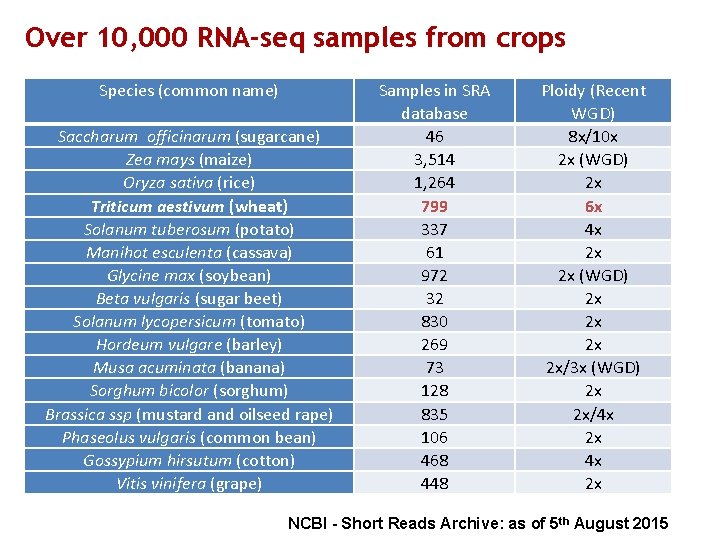
Over 10, 000 RNA-seq samples from crops Species (common name) Saccharum officinarum (sugarcane) Zea mays (maize) Oryza sativa (rice) Triticum aestivum (wheat) Solanum tuberosum (potato) Manihot esculenta (cassava) Glycine max (soybean) Beta vulgaris (sugar beet) Solanum lycopersicum (tomato) Hordeum vulgare (barley) Musa acuminata (banana) Sorghum bicolor (sorghum) Brassica ssp (mustard and oilseed rape) Phaseolus vulgaris (common bean) Gossypium hirsutum (cotton) Vitis vinifera (grape) Samples in SRA database 46 3, 514 1, 264 799 337 61 972 32 830 269 73 128 835 106 468 448 Ploidy (Recent WGD) 8 x/10 x 2 x (WGD) 2 x 6 x 4 x 2 x 2 x (WGD) 2 x 2 x/3 x (WGD) 2 x 2 x/4 x 2 x NCBI - Short Reads Archive: as of 5 th August 2015
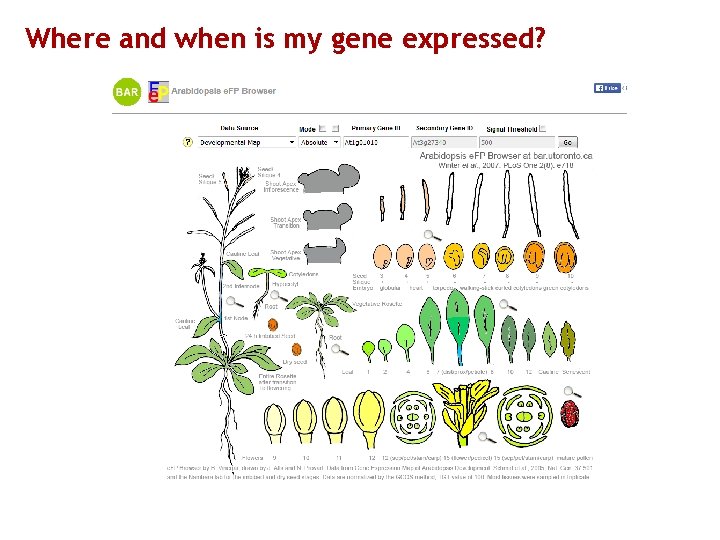
Where and when is my gene expressed?

exp. VIP (expression Visualisation and Integration Platform) • • • User-friendly Intuitive Dynamic/interactive interface (easy to ask questions) Flexible (more data can be added) Simple inputs
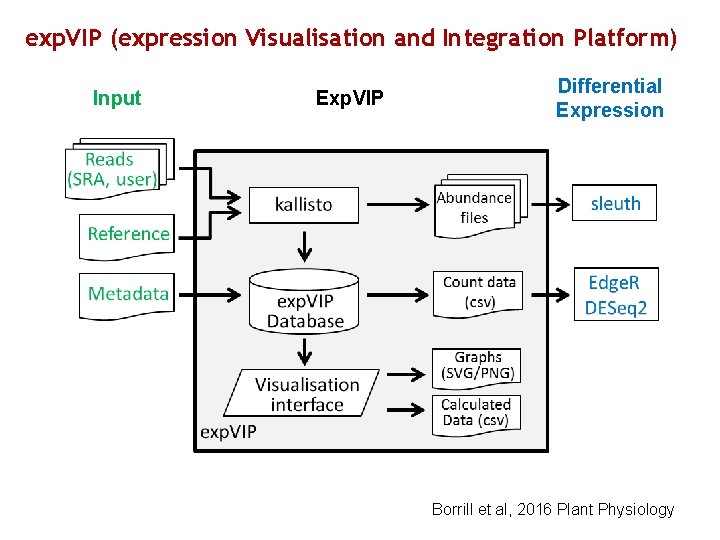
exp. VIP (expression Visualisation and Integration Platform) Input Exp. VIP Differential Expression Borrill et al, 2016 Plant Physiology
Input files • Fastq files from SRA (>850 RNA-seq samples) • Internal/personal files • IWGSC v 2. 26 transcriptome • TGAC transcriptome • Ref. Seqv 1. 0 and v 1. 1 transcriptomes • Information to classify samples (tissue, developmental stage, stress, variety) (Choulet et al, 2014; IWGSC, 2014; Clavijo et al, 2017, IWGSC 2018 )

Input SRA reads 29 studies; 850 RNA-seq samples; >20 billion reads
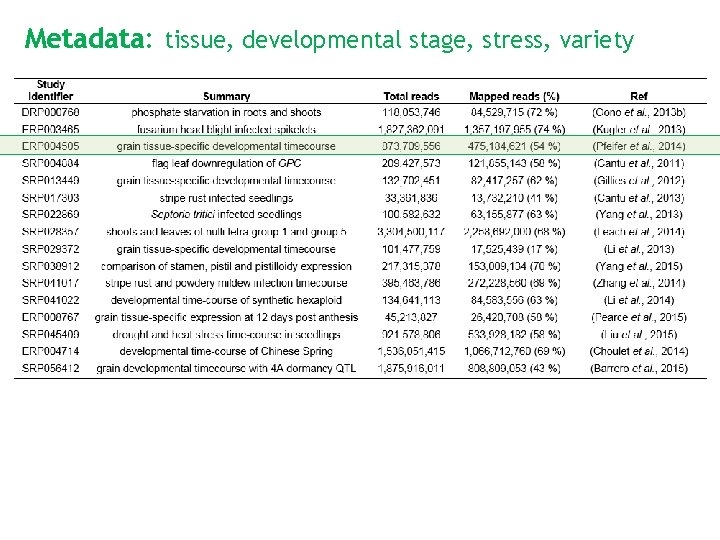
Metadata: tissue, developmental stage, stress, variety
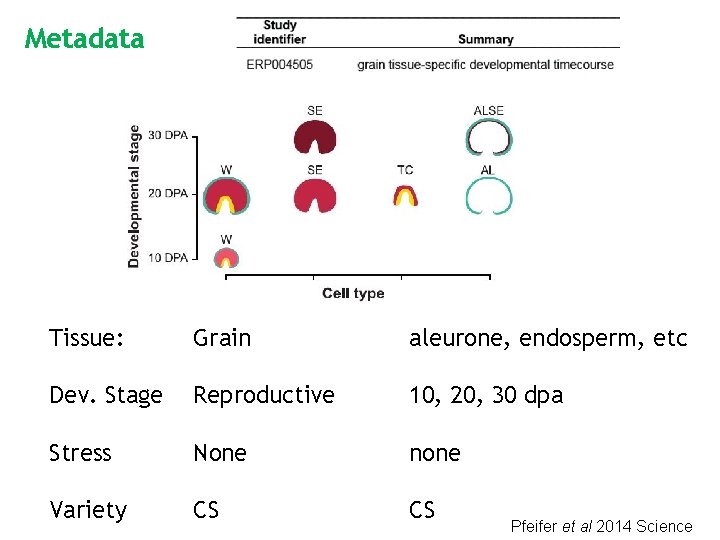
Metadata Tissue: Grain aleurone, endosperm, etc Dev. Stage Reproductive 10, 20, 30 dpa Stress None none Variety CS CS Pfeifer et al 2014 Science
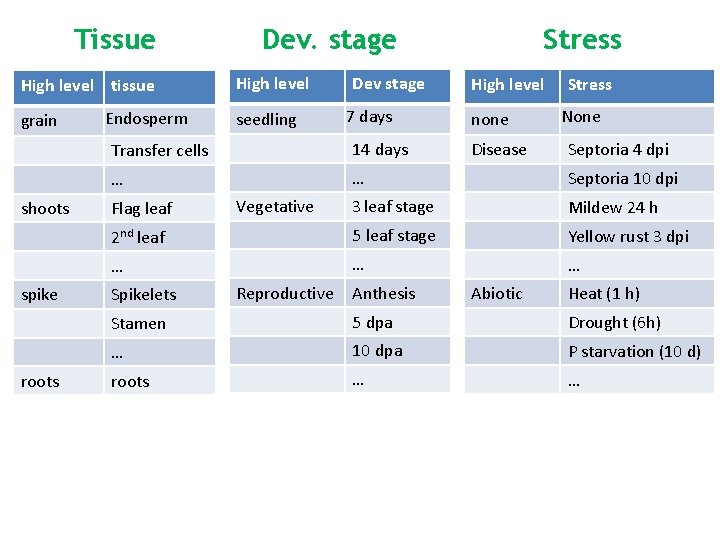
Tissue High level tissue grain shoots spike roots Endosperm Dev. stage High level seedling Dev stage 7 days Stress High level none Stress None Transfer cells 14 days … … Septoria 10 dpi 3 leaf stage Mildew 24 h 2 nd leaf 5 leaf stage Yellow rust 3 dpi … … … Flag leaf Spikelets Vegetative Reproductive Anthesis Disease Abiotic Septoria 4 dpi Heat (1 h) Stamen 5 dpa Drought (6 h) … 10 dpa P starvation (10 d) roots … …
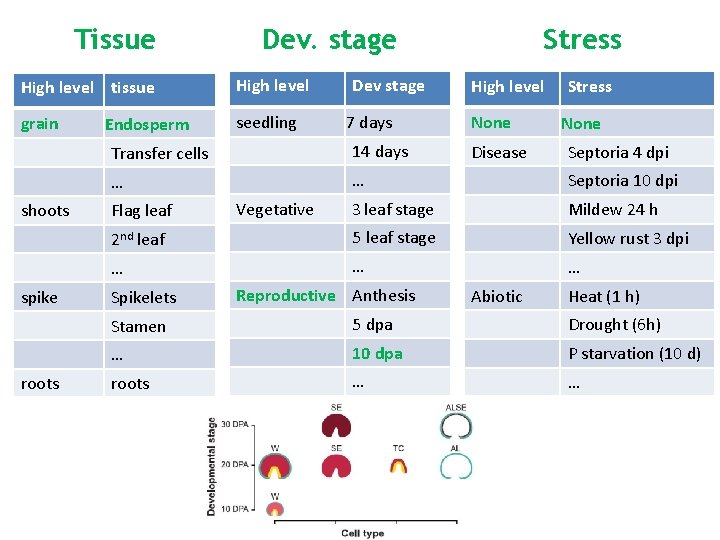
Tissue Dev. stage High level tissue High level grain seedling shoots spike roots Endosperm Dev stage 7 days Stress High level None Stress None Transfer cells 14 days … … Septoria 10 dpi 3 leaf stage Mildew 24 h 2 nd leaf 5 leaf stage Yellow rust 3 dpi … … … Flag leaf Spikelets Vegetative Reproductive Anthesis Disease Abiotic Septoria 4 dpi Heat (1 h) Stamen 5 dpa Drought (6 h) … 10 dpa P starvation (10 d) roots … …
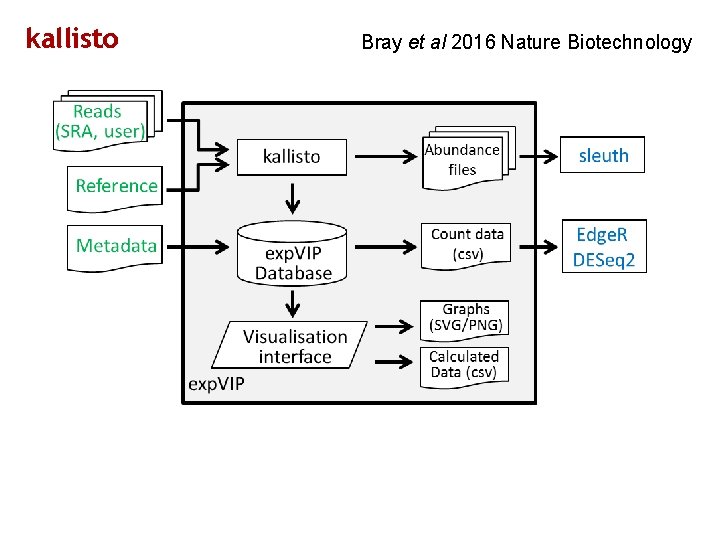
kallisto Bray et al 2016 Nature Biotechnology
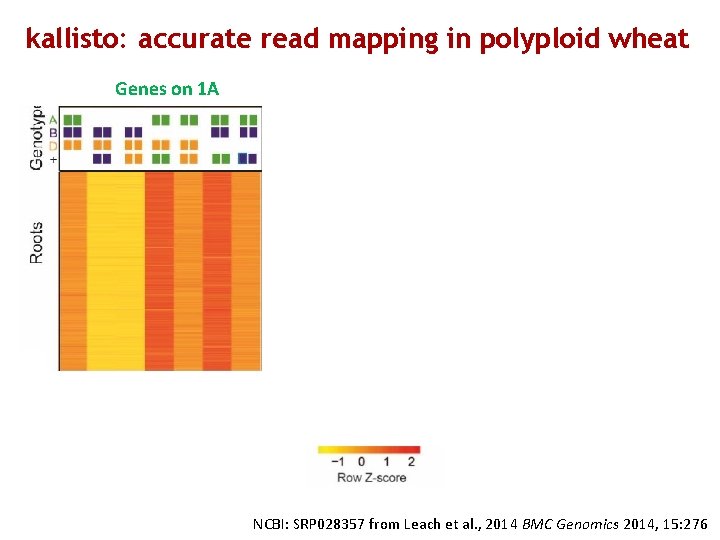
kallisto: accurate read mapping in polyploid wheat Genes on 1 A Genes on 1 B Genes on 1 D NCBI: SRP 028357 from Leach et al. , 2014 BMC Genomics 2014, 15: 276
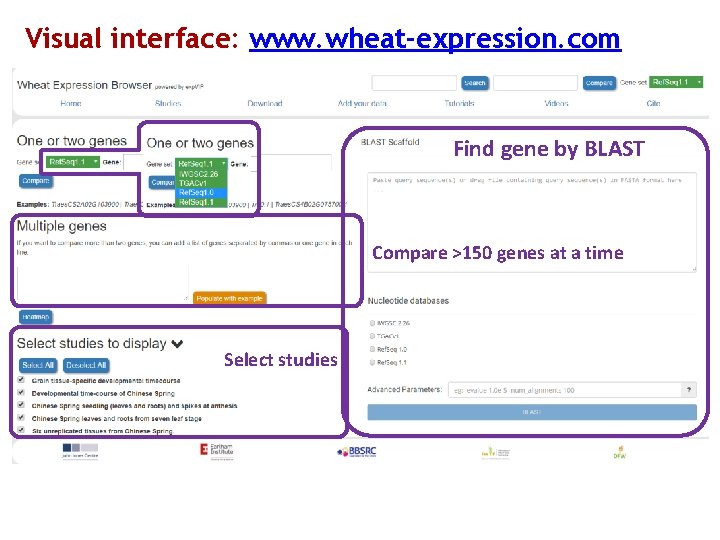
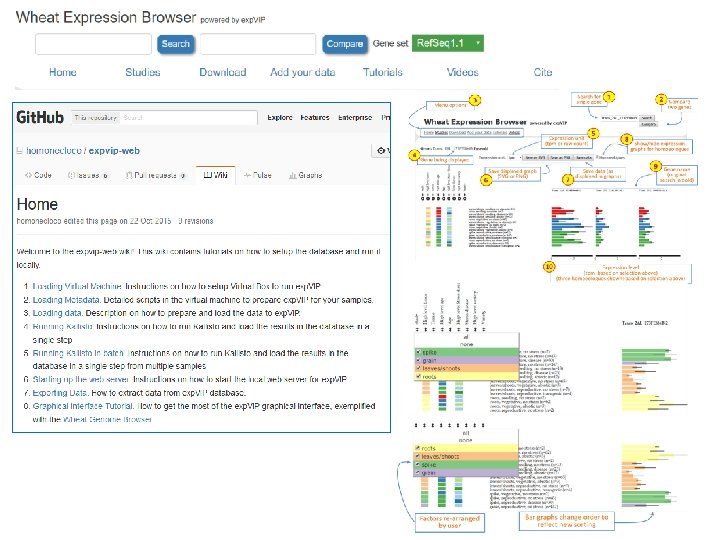
Visual interface: www. wheat-expression. com Find gene by BLAST Compare >150 genes at a time Select studies

Visual interface: www. wheat-expression. com

Visual interface: heatmap
www. wheat-expression. com
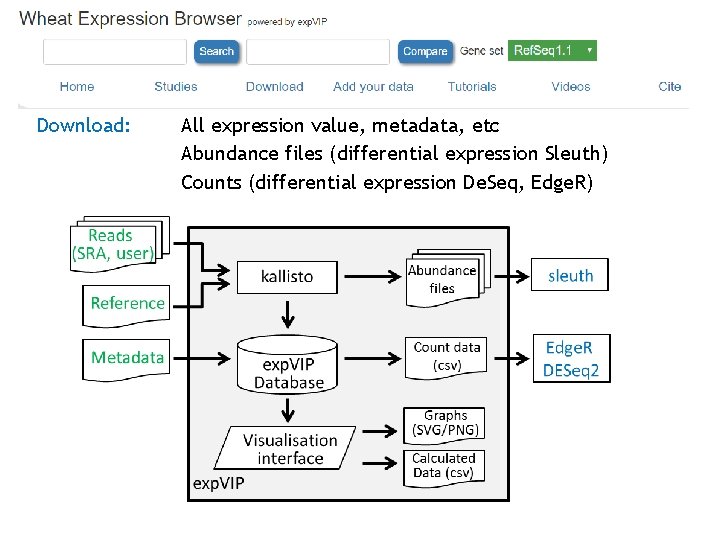
Download: All expression value, metadata, etc Abundance files (differential expression Sleuth) Counts (differential expression De. Seq, Edge. R)
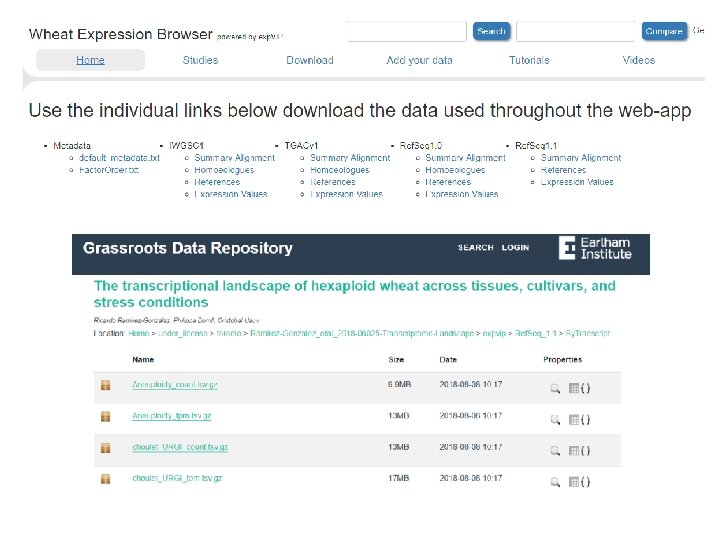
Add your data: download Virtual Machine • With all wheat data • Empty (any transcriptome)
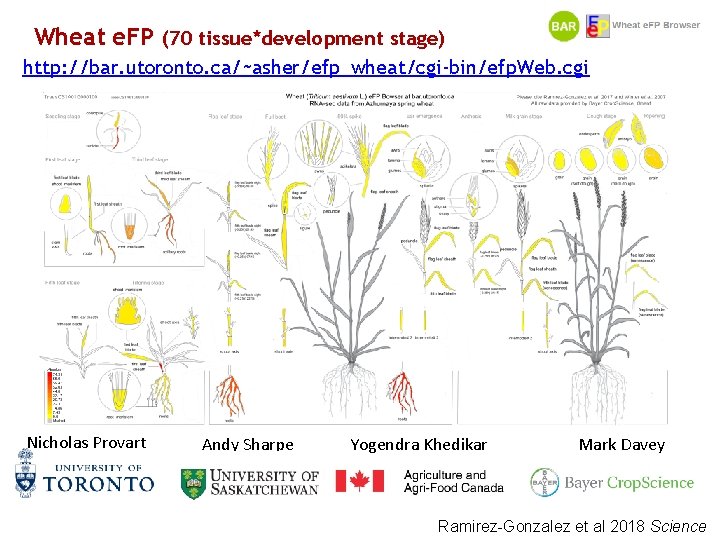
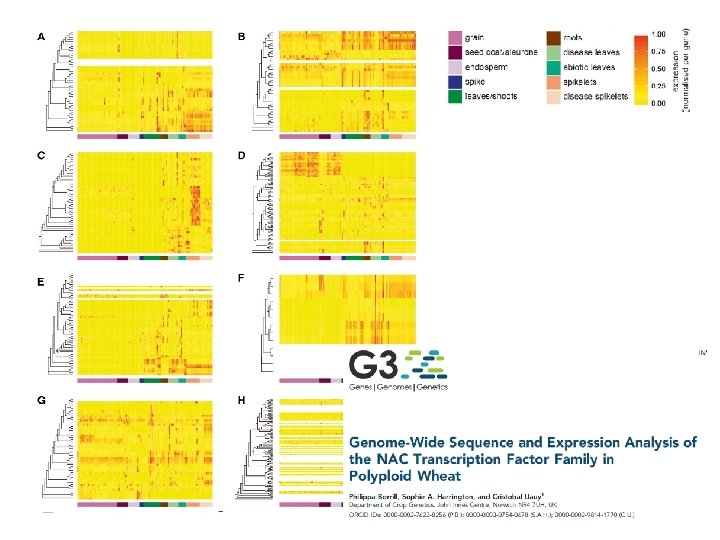
Wheat e. FP (70 tissue*development stage) http: //bar. utoronto. ca/~asher/efp_wheat/cgi-bin/efp. Web. cgi Nicholas Provart Andy Sharpe Yogendra Khedikar Mark Davey Ramirez-Gonzalez et al 2018 Science
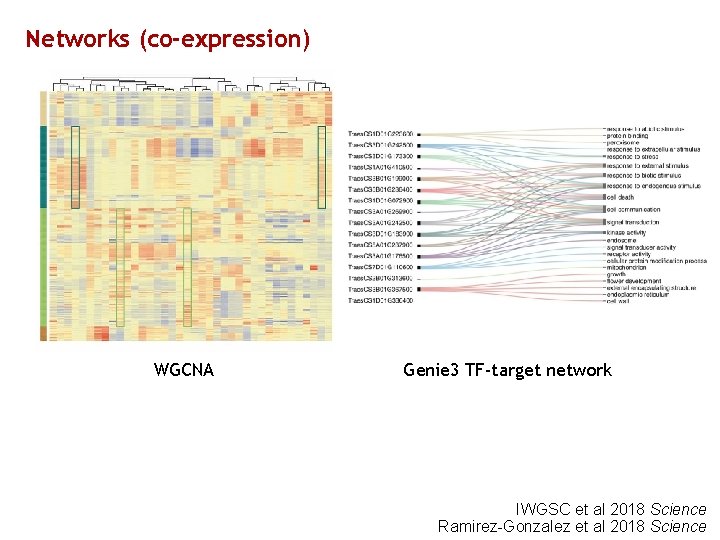
Networks (co-expression) WGCNA Genie 3 TF-target network IWGSC et al 2018 Science Ramirez-Gonzalez et al 2018 Science
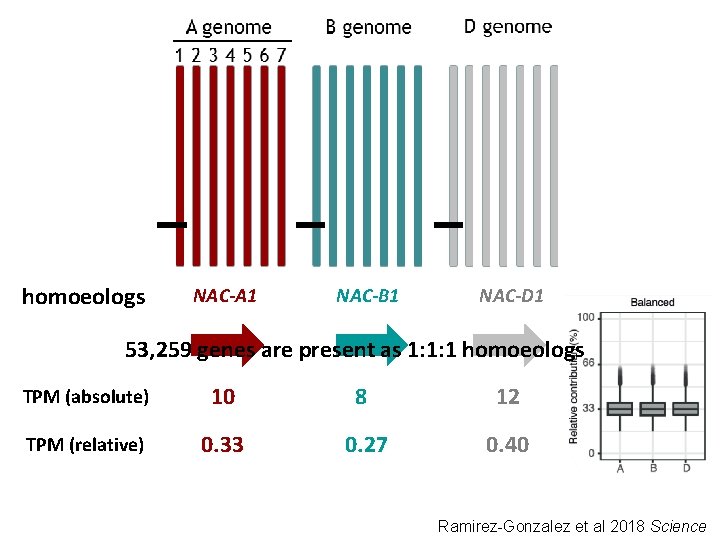
homoeologs NAC-A 1 NAC-B 1 NAC-D 1 53, 259 genes are present as 1: 1: 1 homoeologs TPM (absolute) 10 8 12 TPM (relative) 0. 33 0. 27 0. 40 Ramirez-Gonzalez et al 2018 Science
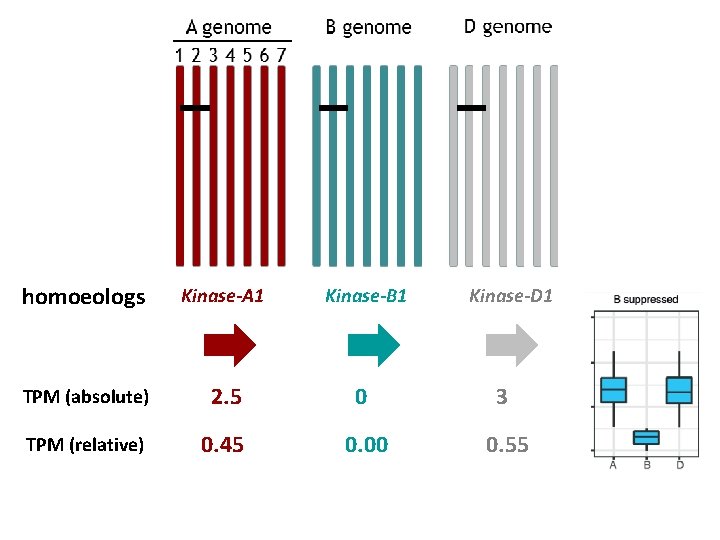
homoeologs Kinase-A 1 Kinase-B 1 TPM (absolute) 2. 5 0 3 TPM (relative) 0. 45 0. 00 0. 55 Kinase-D 1
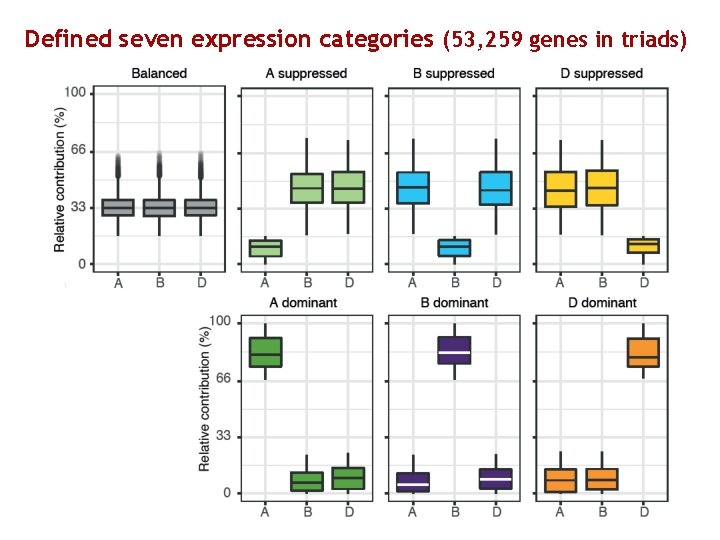
Defined seven expression categories (53, 259 genes in triads) Natural variation
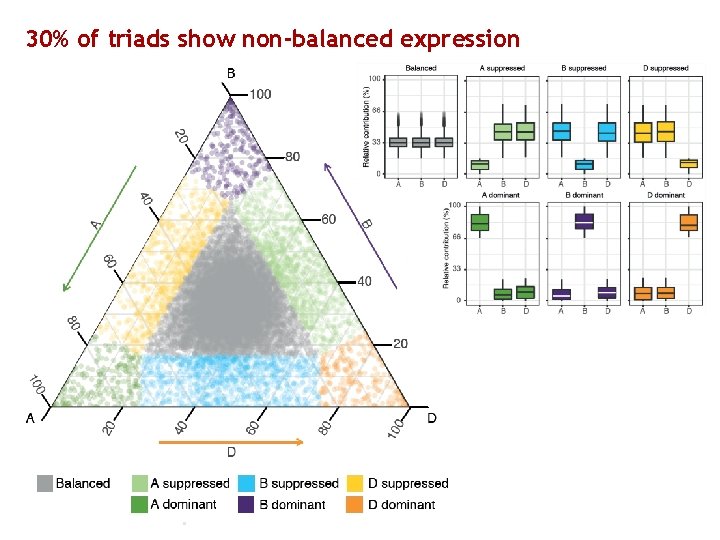
30% of triads show non-balanced expression
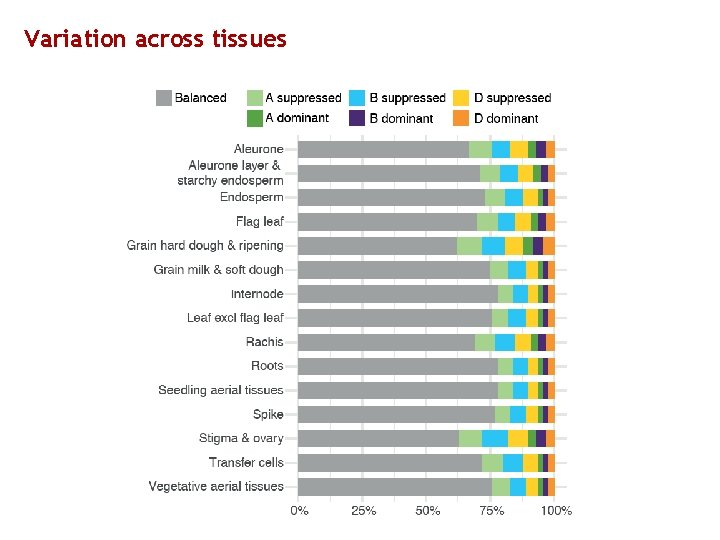
Variation across tissues
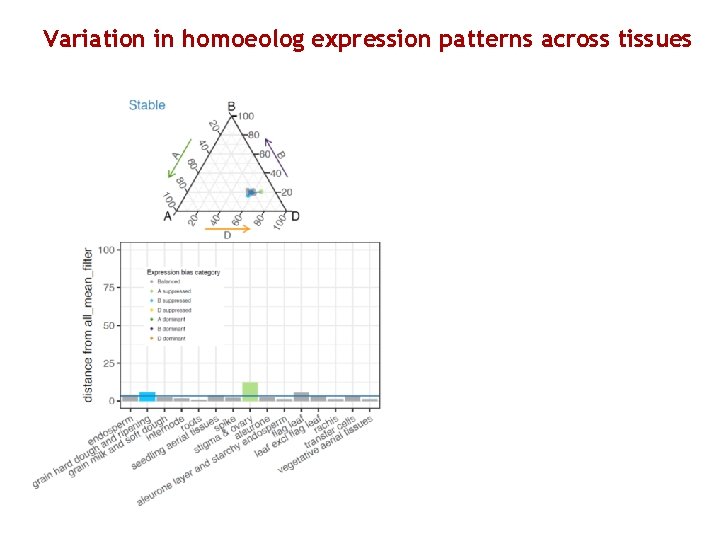
Variation in homoeolog expression patterns across tissues
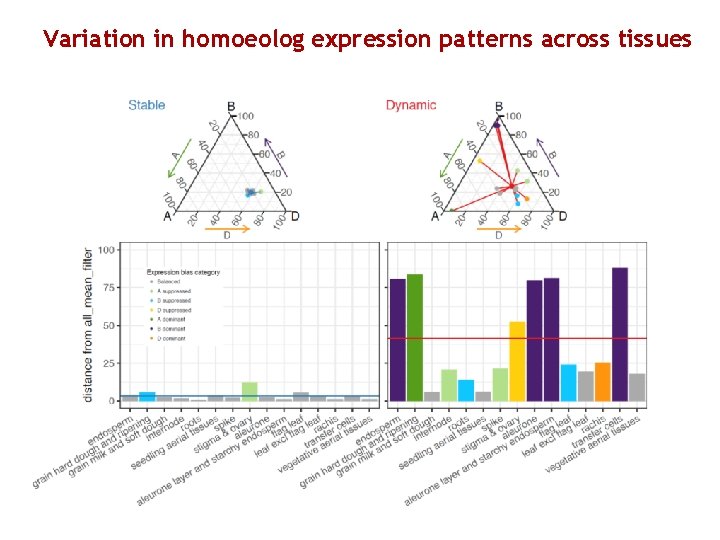
Variation in homoeolog expression patterns across tissues
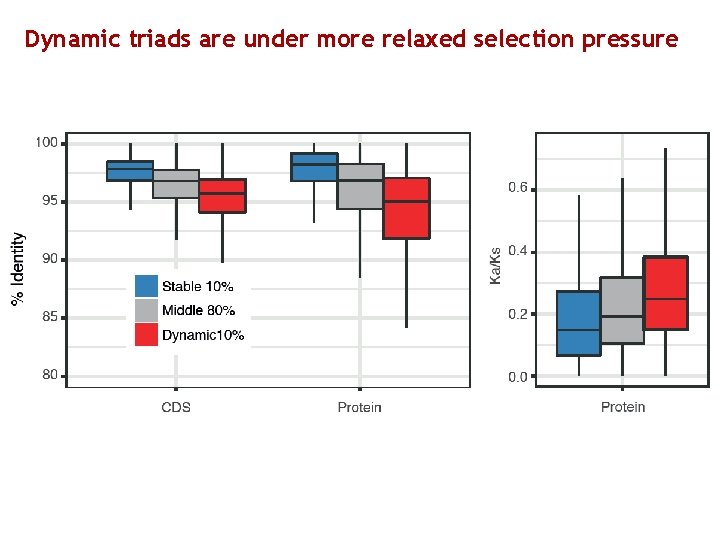
Dynamic triads are under more relaxed selection pressure

Final comments • • • Take advantage of what is out there Careful curation of data is essential Publish with ontologies to allow your data to be re-used Consider it a success if your data is reused! “pay” back to the community…. Community feedback very welcomed!

Ricardo Ramirez-Gonzalez Philippa Borrill Rob Davey Toni Etuk @Cristobal. Uauy Jemima Brinton Sophie Harrington Source code: Database and interface setup: https: //github. com/homonecloco/expvip-web Bio. JS visualisation component: http: //biojs. io/d/bio-vis-expression-bar Wheat (and human) diversity
- Slides: 36