What is hematopoiesis Hematopoiesis is a term describing
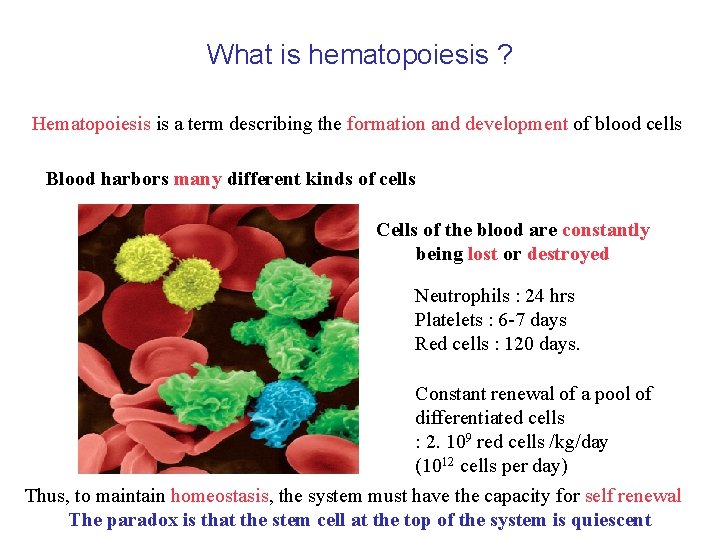
What is hematopoiesis ? Hematopoiesis is a term describing the formation and development of blood cells Blood harbors many different kinds of cells Cells of the blood are constantly being lost or destroyed Neutrophils : 24 hrs Platelets : 6 -7 days Red cells : 120 days. Constant renewal of a pool of differentiated cells : 2. 109 red cells /kg/day (1012 cells per day) Thus, to maintain homeostasis, the system must have the capacity for self renewal The paradox is that the stem cell at the top of the system is quiescent
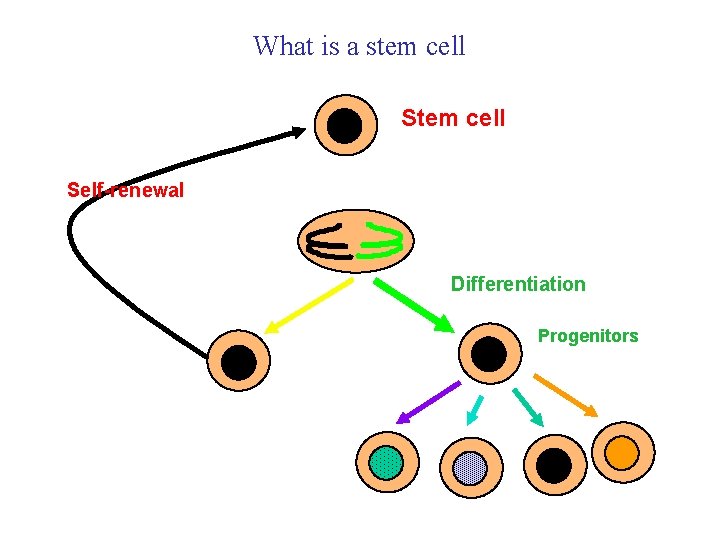
What is a stem cell Self-renewal Differentiation Progenitors
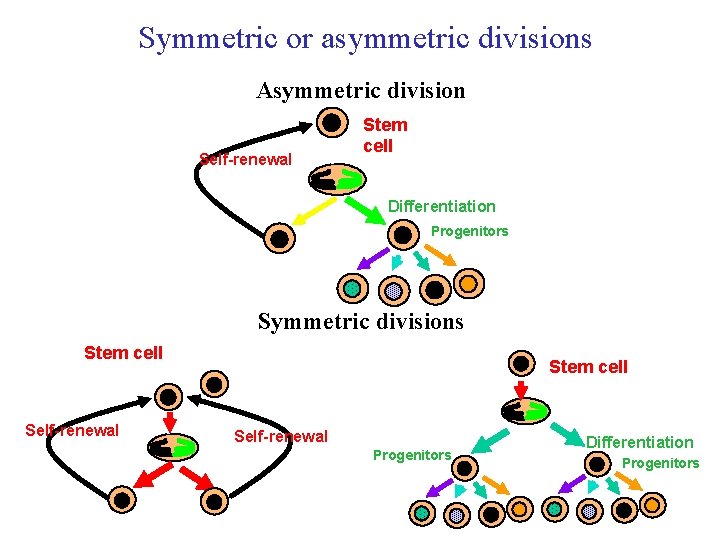
Symmetric or asymmetric divisions Asymmetric division Self-renewal Stem cell Differentiation Progenitors Symmetric divisions Stem cell Self-renewal Progenitors Differentiation Progenitors
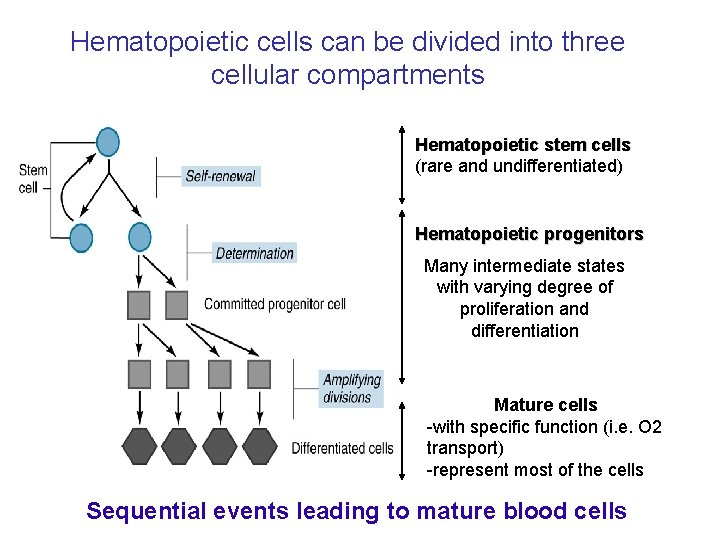
Hematopoietic cells can be divided into three cellular compartments Hematopoietic stem cells (rare and undifferentiated) Hematopoietic progenitors Many intermediate states with varying degree of proliferation and differentiation Mature cells -with specific function (i. e. O 2 transport) -represent most of the cells Sequential events leading to mature blood cells
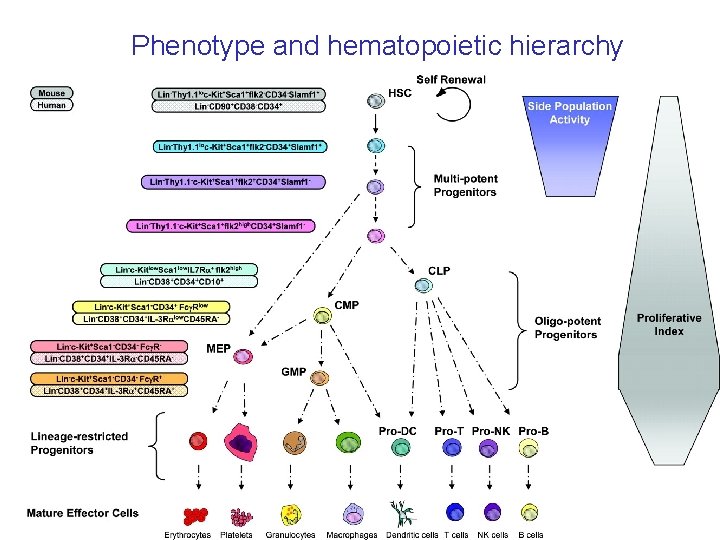
Phenotype and hematopoietic hierarchy
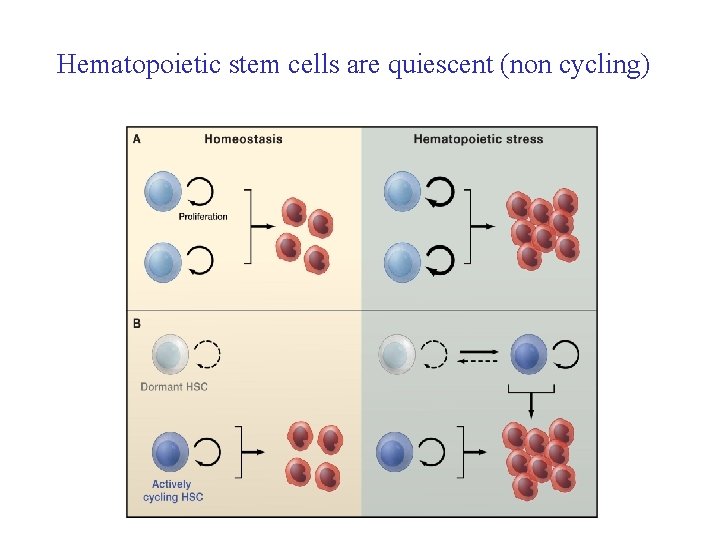
Hematopoietic stem cells are quiescent (non cycling)
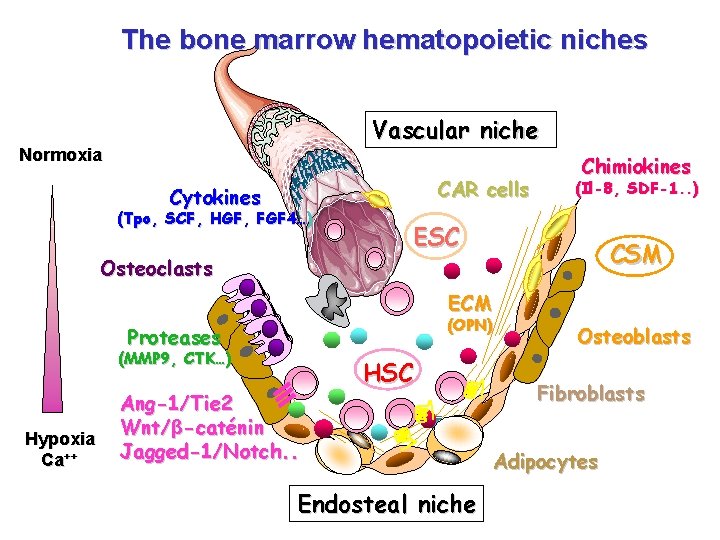
The bone marrow hematopoietic niches Vascular niche Normoxia CAR cells Cytokines (Tpo, SCF, HGF, FGF 4…) Osteoclasts Chimiokines (Il-8, SDF-1. . ) ESC CSM ECM (OPN) Proteases (MMP 9, CTK…) Hypoxia Ca++ Ang-1/Tie 2 Wnt/β-caténin Jagged-1/Notch. . HSC Endosteal niche Osteoblasts Fibroblasts Adipocytes
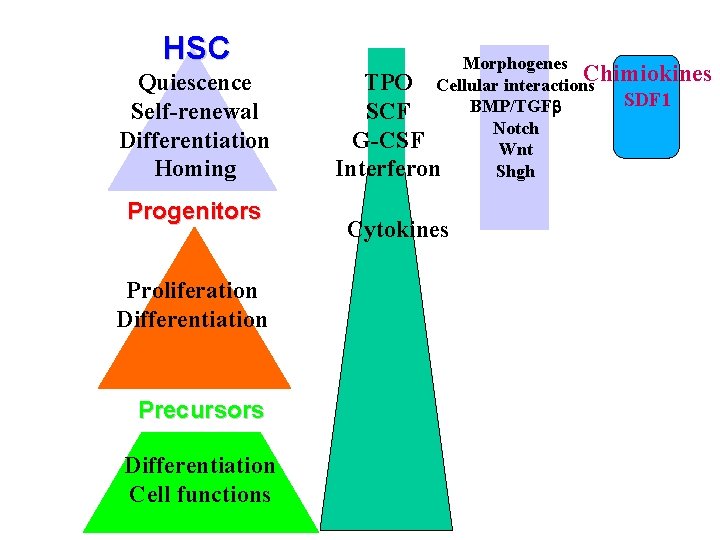
HSC Quiescence Self-renewal Differentiation Homing Progenitors Proliferation Differentiation Precursors Differentiation Cell functions Morphogenes TPO Cellular interactions. Chimiokines SDF 1 BMP/TGFb SCF Notch G-CSF Wnt Interferon Shgh Cytokines
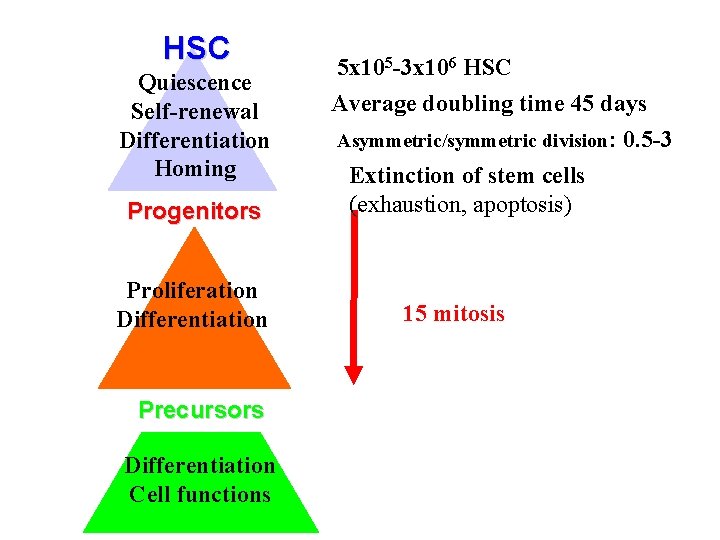
HSC Quiescence Self-renewal Differentiation Homing Progenitors Proliferation Differentiation Precursors Differentiation Cell functions 5 x 105 -3 x 106 HSC Average doubling time 45 days Asymmetric/symmetric division: 0. 5 -3 Extinction of stem cells (exhaustion, apoptosis) 15 mitosis
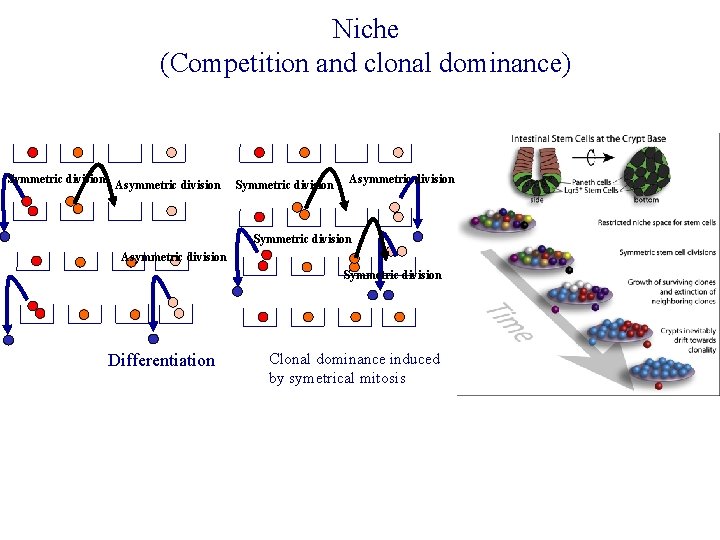
Niche (Competition and clonal dominance) Symmetric division Asymmetric division Symmetric division Differentiation Clonal dominance induced by symetrical mitosis
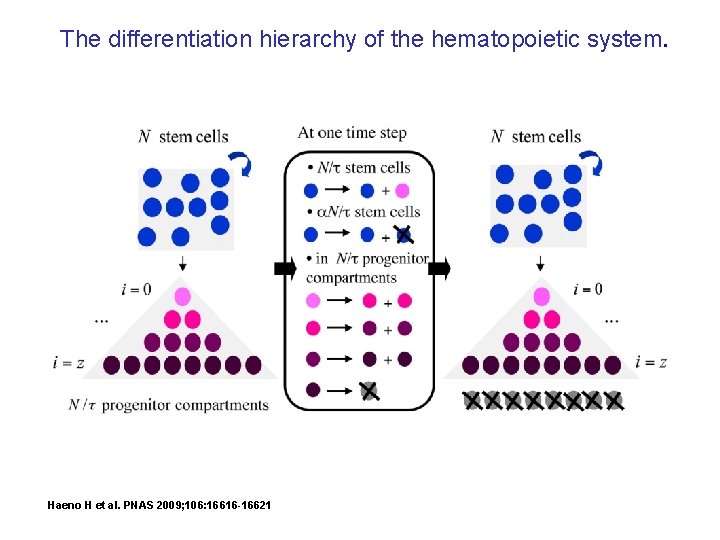
The differentiation hierarchy of the hematopoietic system. Haeno H et al. PNAS 2009; 106: 16616 -16621
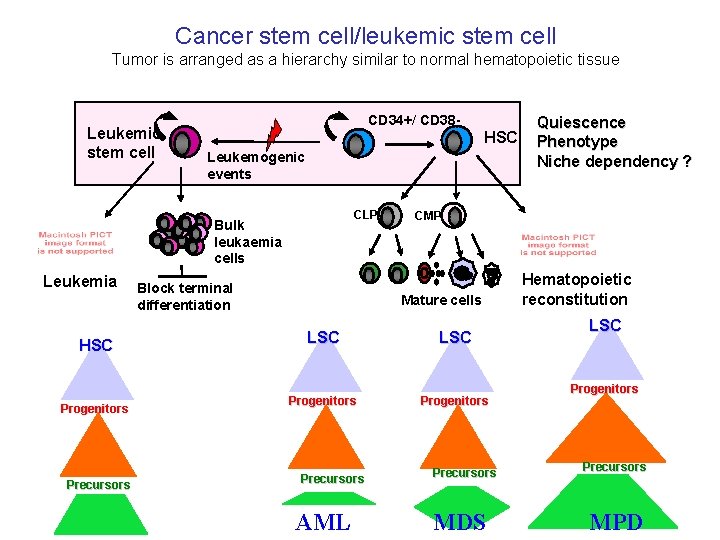
Cancer stem cell/leukemic stem cell Tumor is arranged as a hierarchy similar to normal hematopoietic tissue Leukemic stem cell CD 34+/ CD 38 - HSC Leukemogenic events CLP Bulk leukaemia cells Leukemia HSC Progenitors Precursors Block terminal differentiation CMP Mature cells LSC Progenitors Precursors AML Quiescence Phenotype Niche dependency ? Precursors MDS Hematopoietic reconstitution LSC Progenitors Precursors MPD
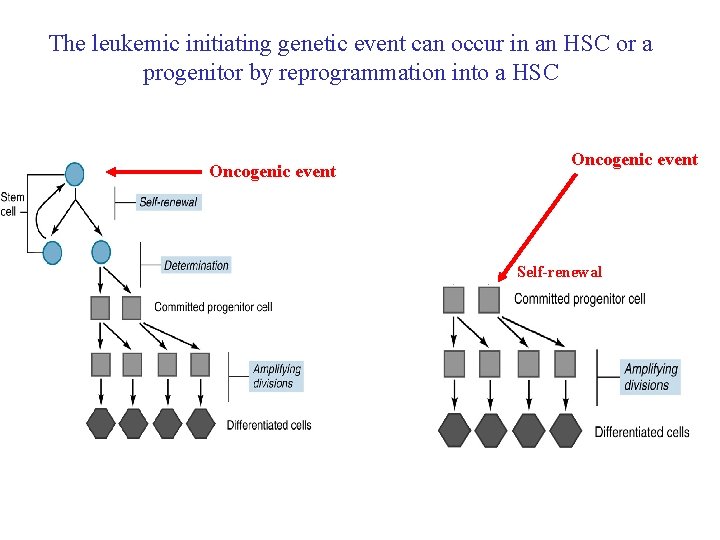
The leukemic initiating genetic event can occur in an HSC or a progenitor by reprogrammation into a HSC Oncogenic event Self-renewal
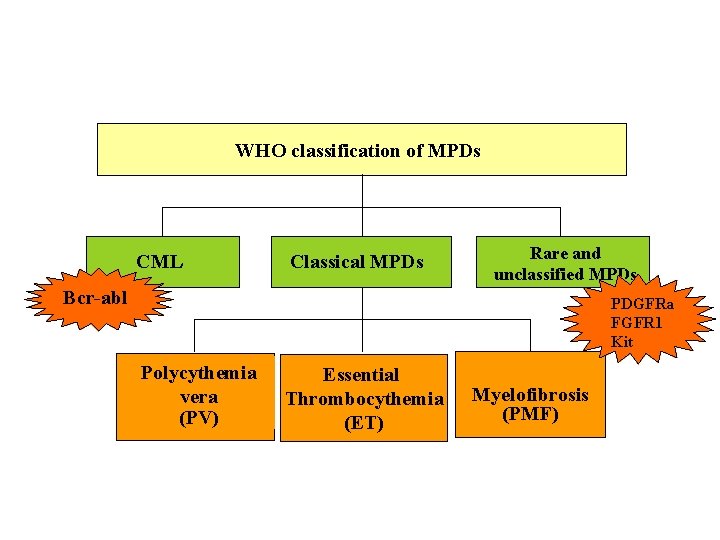
WHO classification of MPDs CML Classical MPDs Rare and unclassified MPDs Bcr-abl PDGFRa FGFR 1 Kit Polycythemia vera (PV) Essential Thrombocythemia (ET) Myelofibrosis (PMF)
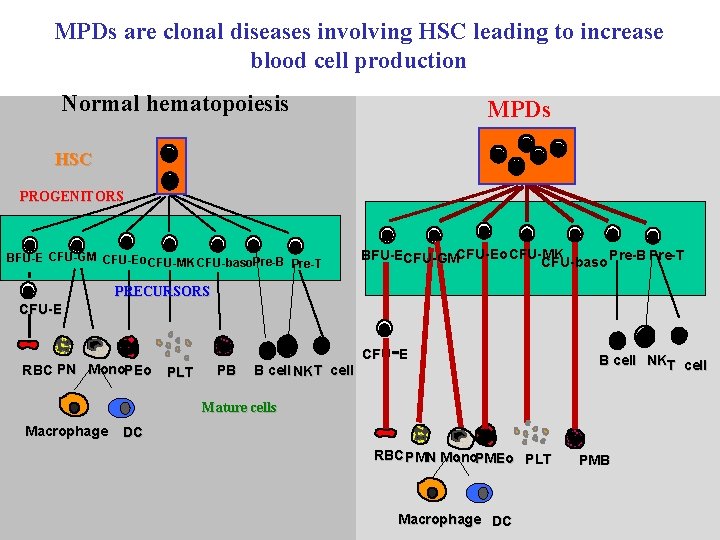
MPDs are clonal diseases involving HSC leading to increase blood cell production Normal hematopoiesis MPDs HSC PROGENITORS BFU-E CFU-GM CFU-Eo CFU-MK CFU-baso. Pre-B Pre-T BFU-ECFU-GMCFU-Eo CFU-MK Pre-B Pre-T CFU-baso PRECURSORS CFU-E RBC PN Mono. PEo - CFU E PLT PB B cell NK T cell B cell NKT cell Mature cells Macrophage DC RBC PMN Mono. PMEo PLT Macrophage DC PMB
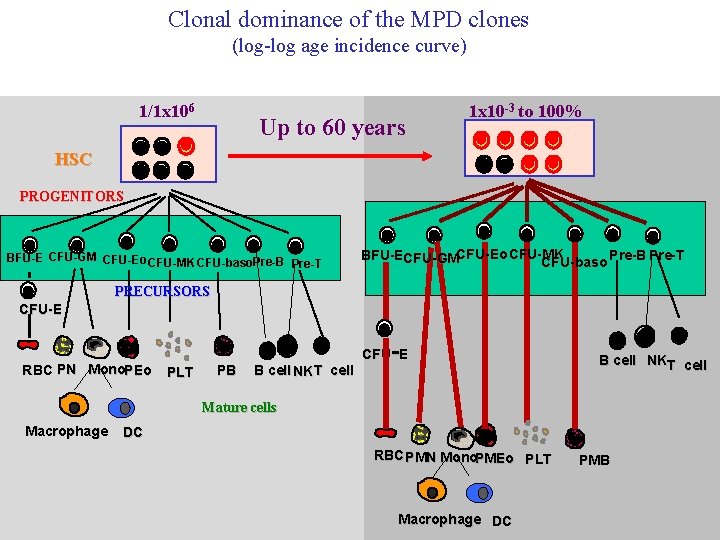
Clonal dominance of the MPD clones (log-log age incidence curve) 1/1 x 106 Up to 60 years 1 x 10 -3 to 100% HSC PROGENITORS BFU-E CFU-GM CFU-Eo CFU-MK CFU-baso. Pre-B Pre-T BFU-ECFU-GMCFU-Eo CFU-MK Pre-B Pre-T CFU-baso PRECURSORS CFU-E RBC PN Mono. PEo - CFU E PLT PB B cell NK T cell B cell NKT cell Mature cells Macrophage DC RBC PMN Mono. PMEo PLT Macrophage DC PMB
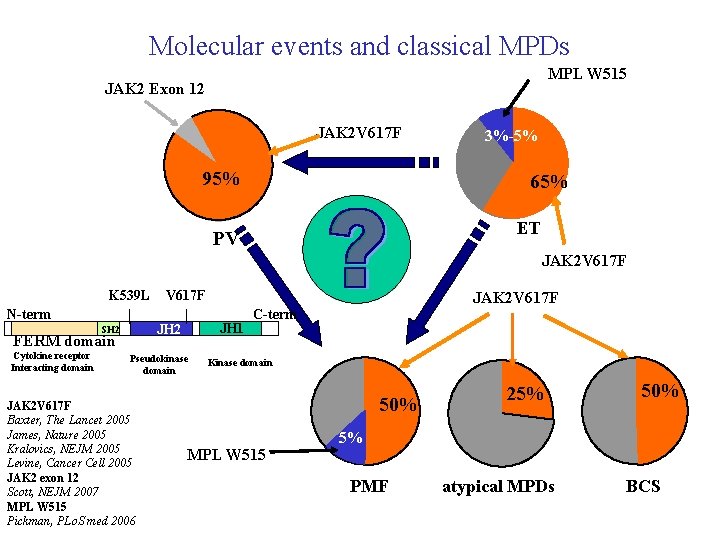
Molecular events and classical MPDs MPL W 515 JAK 2 Exon 12 JAK 2 V 617 F 3%-5% 30% 95% 65% ET PV JAK 2 V 617 F K 539 L N-term Pseudokinase domain JAK 2 V 617 F Baxter, The Lancet 2005 James, Nature 2005 Kralovics, NEJM 2005 Levine, Cancer Cell 2005 JAK 2 exon 12 Scott, NEJM 2007 MPL W 515 Pickman, PLo. S med 2006 JAK 2 V 617 F JH 1 JH 2 SH 2 FERM domain Cytokine receptor Interacting domain V 617 F C-term Kinase domain 45% 50% MPL W 515 25% 50% 5% PMF atypical MPDs BCS
A proposed structure of JAK 2 V 617 F Lee et al. Cancer 15, 1692, 2009
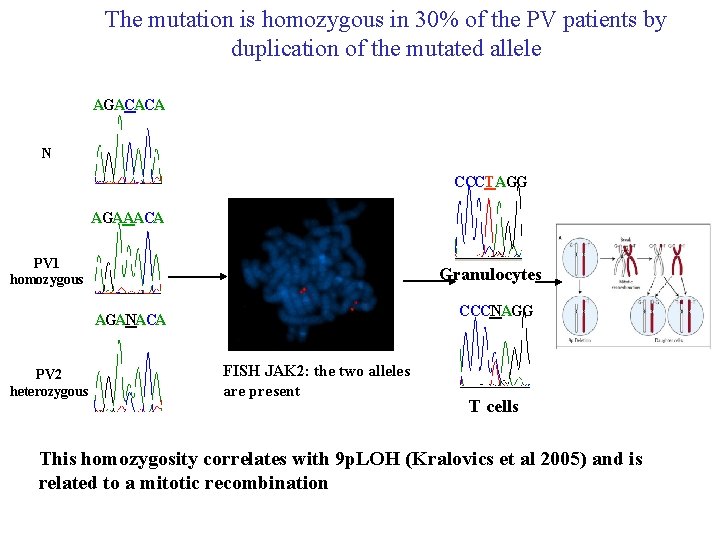
The mutation is homozygous in 30% of the PV patients by duplication of the mutated allele AGACACA N CCCTAGG AGAAACA PV 1 homozygous Granulocytes CCCNAGG AGANACA PV 2 heterozygous FISH JAK 2: the two alleles are present T cells This homozygosity correlates with 9 p. LOH (Kralovics et al 2005) and is related to a mitotic recombination
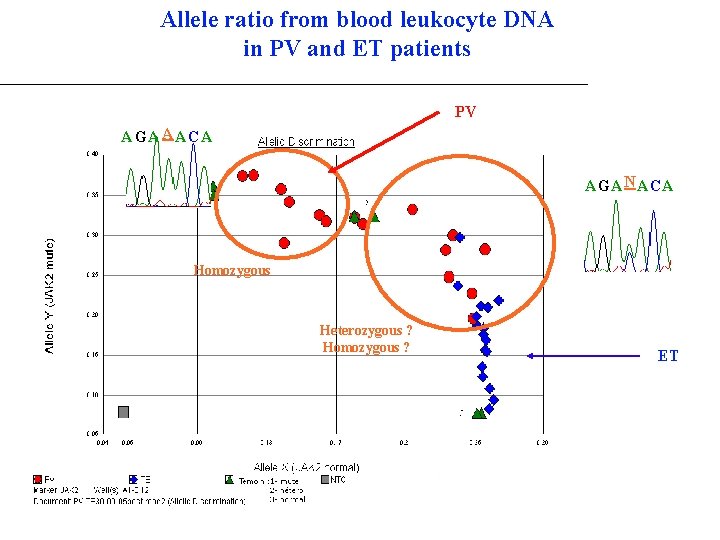
Allele ratio from blood leukocyte DNA in PV and ET patients PV A GA AA CA A GA N A CA Homozygous Heterozygous ? Homozygous ? ET
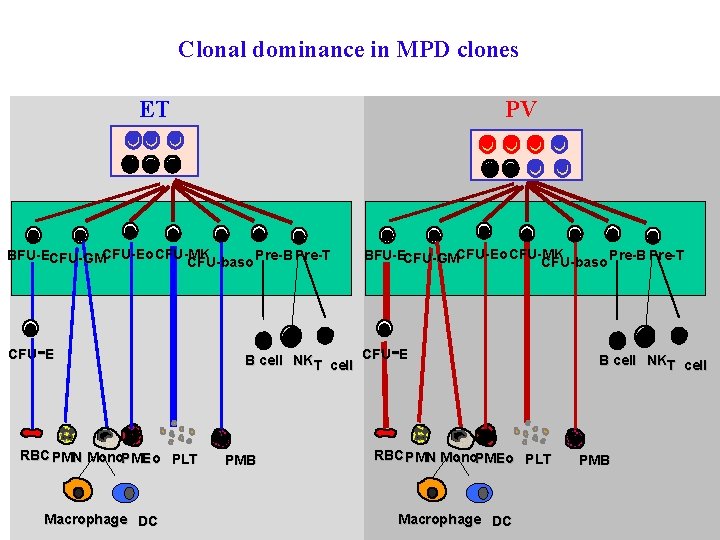
Clonal dominance in MPD clones ET PV BFU-ECFU-GMCFU-Eo CFU-MK Pre-B Pre-T CFU-baso - CFU E RBC PMN Mono. PMEo PLT Macrophage DC BFU-ECFU-GMCFU-Eo CFU-MK Pre-B Pre-T CFU-baso - B cell NKT cell CFU E PMB RBC PMN Mono. PMEo PLT Macrophage DC B cell NKT cell PMB
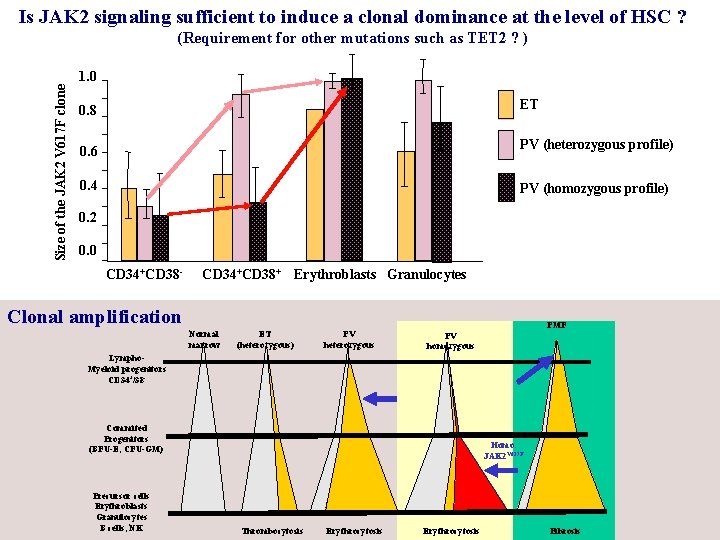
Is JAK 2 signaling sufficient to induce a clonal dominance at the level of HSC ? Size of the JAK 2 V 617 F clone (Requirement for other mutations such as TET 2 ? ) 1. 0 0. 8 ET 0. 6 PV (heterozygous profile) 0. 4 PV (homozygous profile) 0. 2 0. 0 CD 34+CD 38 - CD 34+CD 38+ Erythroblasts Granulocytes Clonal amplification Normal marrow ET (heterozygous) PV heterozygous PMF PV homozygous Lympho. Myeloid progenitors CD 34+/38 - Commited Progenitors (BFU-E, CFU-GM) Precursor cells Erythroblasts Granulocytes B cells, NK Homo. JAK 2 V 617 F Thrombocytosis Erythrocytosis Fibrosis
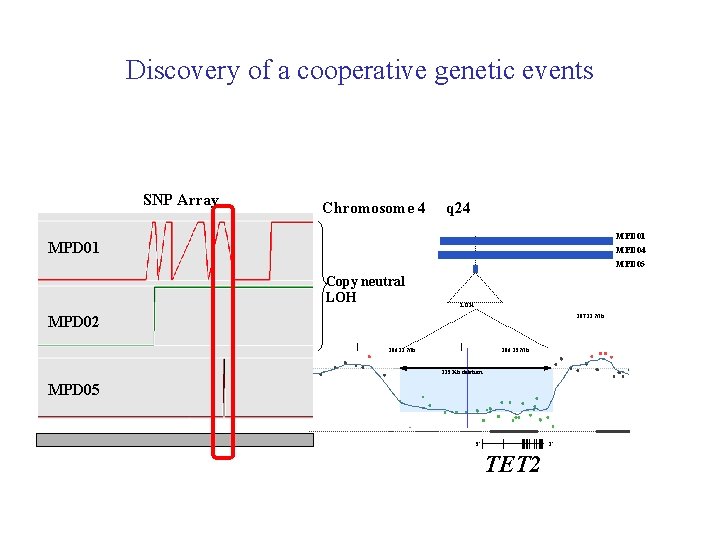
Discovery of a cooperative genetic events SNP Array Chromosome 4 q 24 MPD 01 MPD 04 MPD 05 Copy neutral LOH MPD 02 107. 33 Mb 106. 11 Mb 106. 35 Mb 325 Kb deletion MPD 05 5' 3' TET 2
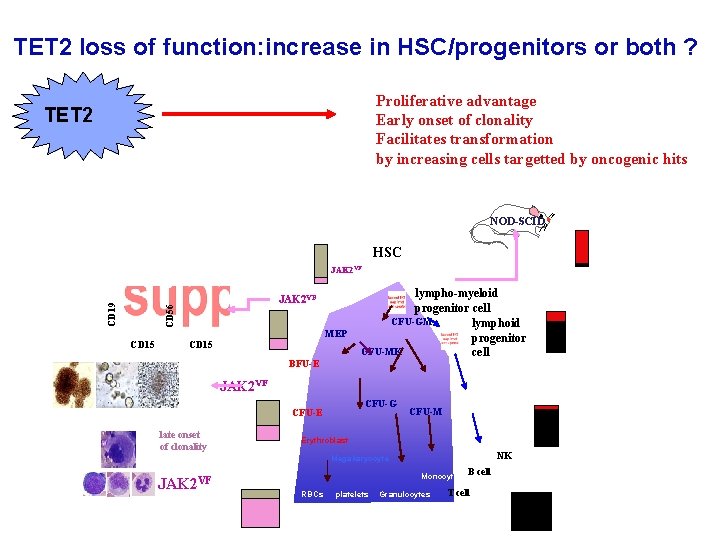
TET 2 loss of function: increase in HSC/progenitors or both ? Proliferative advantage Early onset of clonality Facilitates transformation by increasing cells targetted by oncogenic hits TET 2 NOD-SCID JAK 2 VF CD 56 CD 19 HSC JAK 2 VF CD 15 5 MEP CD 15 5 BFU-E lympho-myeloid progenitor cell CFU-GM lymphoid progenitor CFU-MK cell JAK 2 VF CFU-G CFU-E late onset of clonality CFU-M Erythroblast NK Megakaryocyte JAK 2 VF Monocytes RBCs platelets Granulocytes B cell T cell
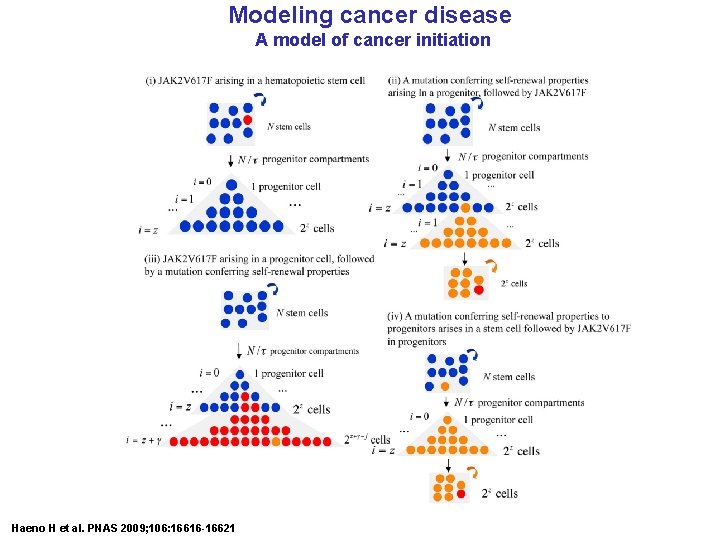
Modeling cancer disease A model of cancer initiation Haeno H et al. PNAS 2009; 106: 16616 -16621
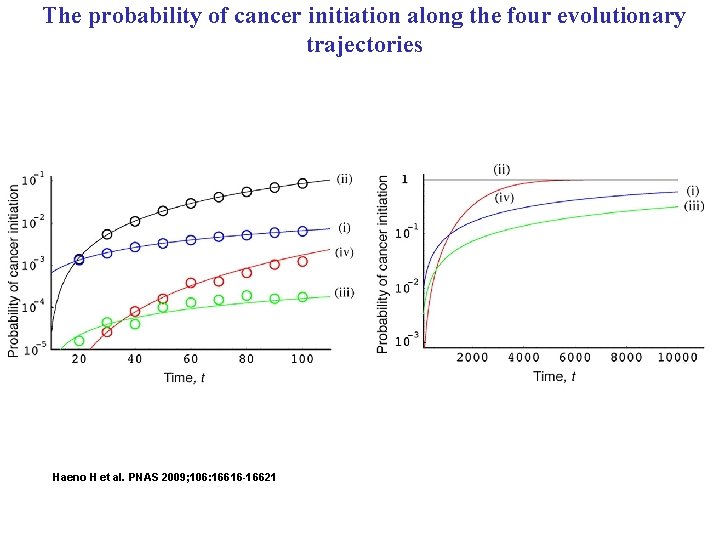
The probability of cancer initiation along the four evolutionary trajectories Haeno H et al. PNAS 2009; 106: 16616 -16621
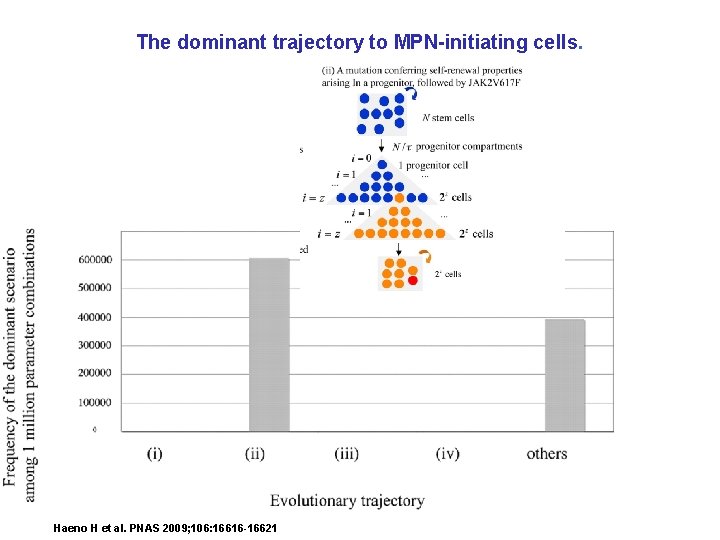
The dominant trajectory to MPN-initiating cells. Haeno H et al. PNAS 2009; 106: 16616 -16621
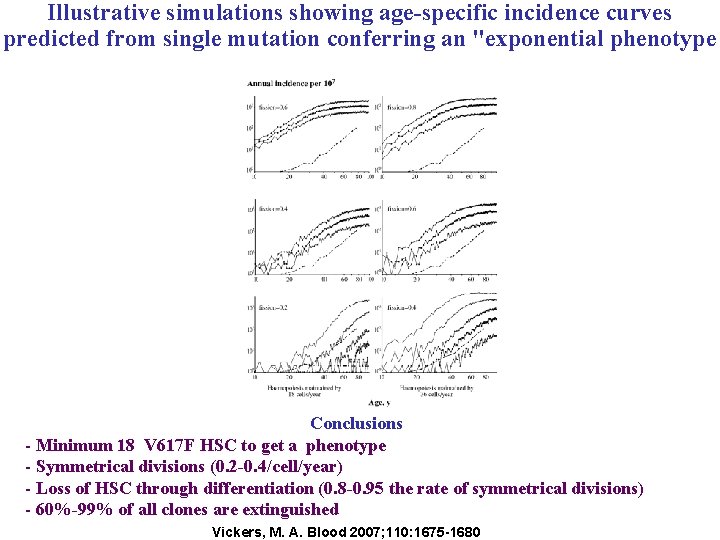
Illustrative simulations showing age-specific incidence curves predicted from single mutation conferring an "exponential phenotype Conclusions - Minimum 18 V 617 F HSC to get a phenotype - Symmetrical divisions (0. 2 -0. 4/cell/year) - Loss of HSC through differentiation (0. 8 -0. 95 the rate of symmetrical divisions) - 60%-99% of all clones are extinguished Vickers, M. A. Blood 2007; 110: 1675 -1680
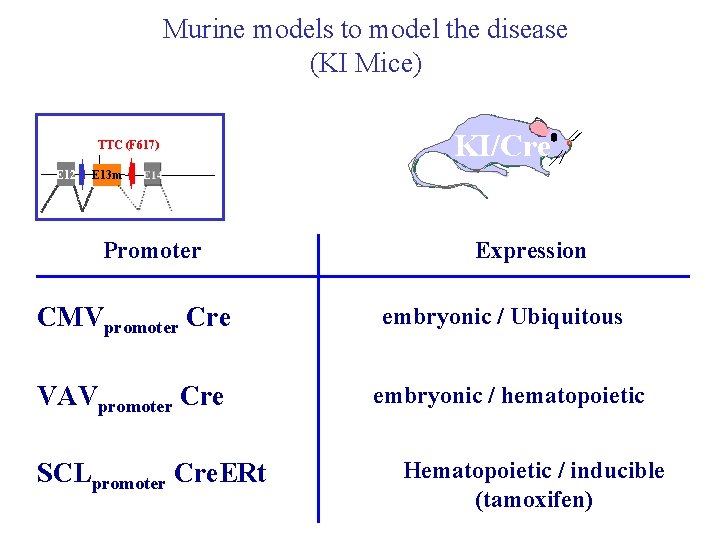
Murine models to model the disease (KI Mice) TTC (F 617) E 12 KI/Cre E 13 m Promoter Expression CMVpromoter Cre embryonic / Ubiquitous VAVpromoter Cre embryonic / hematopoietic SCLpromoter Cre. ERt Hematopoietic / inducible (tamoxifen)
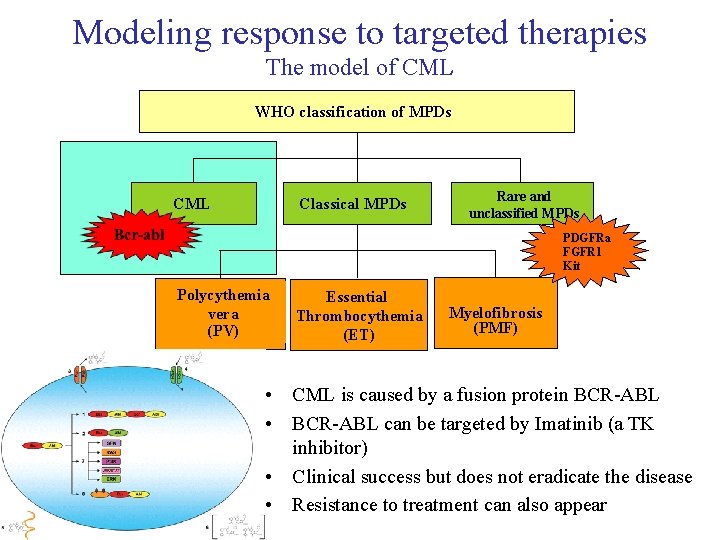
Modeling response to targeted therapies The model of CML WHO classification of MPDs Classical MPDs CML Rare and unclassified MPDs PDGFRa FGFR 1 Kit Polycythemia vera (PV) Essential Thrombocythemia (ET) Myelofibrosis (PMF) • CML is caused by a fusion protein BCR-ABL • BCR-ABL can be targeted by Imatinib (a TK inhibitor) • Clinical success but does not eradicate the disease • Resistance to treatment can also appear
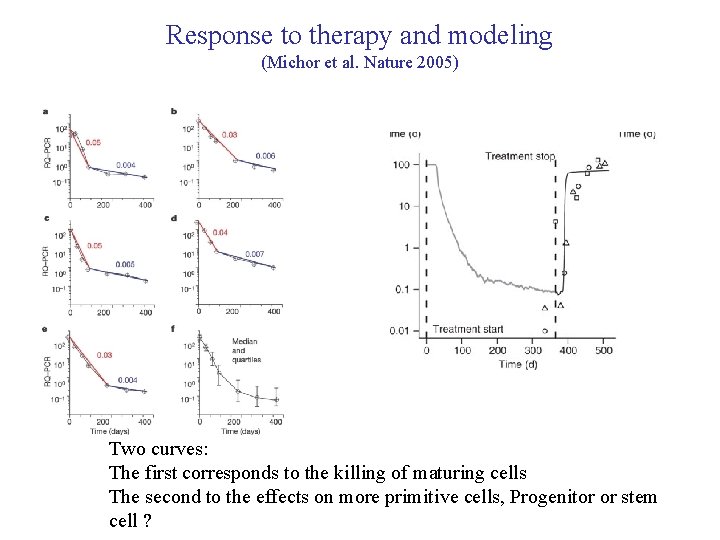
Response to therapy and modeling (Michor et al. Nature 2005) Two curves: The first corresponds to the killing of maturing cells The second to the effects on more primitive cells, Progenitor or stem cell ?
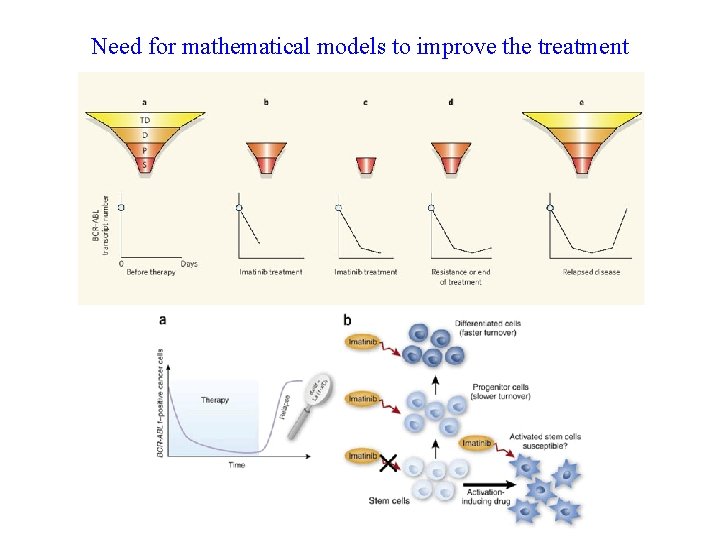
Need for mathematical models to improve the treatment
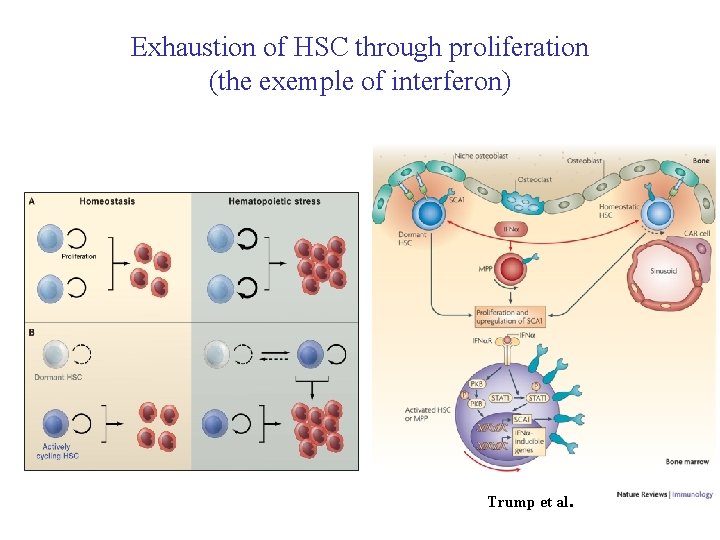
Exhaustion of HSC through proliferation (the exemple of interferon) Trump et al.
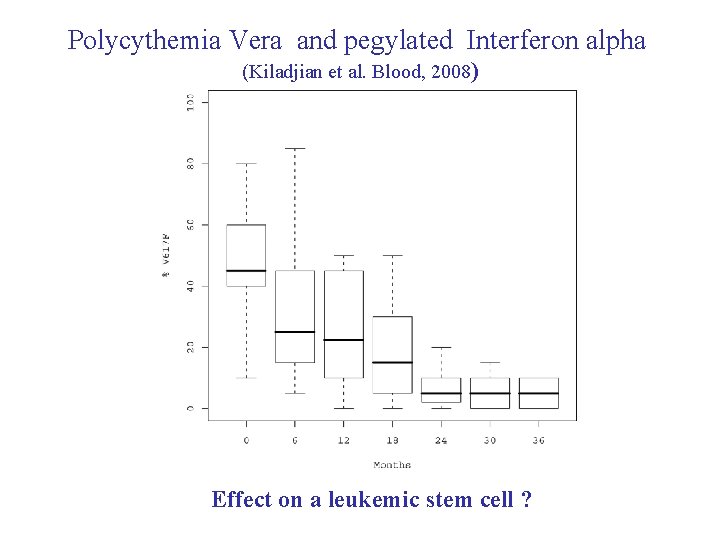
Polycythemia Vera and pegylated Interferon alpha (Kiladjian et al. Blood, 2008) Effect on a leukemic stem cell ?
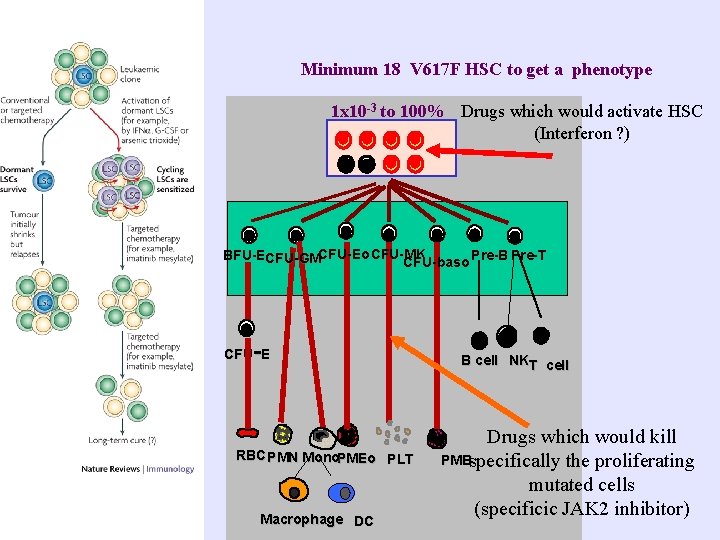
Minimum 18 V 617 F HSC to get a phenotype 1 x 10 -3 to 100% Drugs which would activate HSC (Interferon ? ) BFU-ECFU-GMCFU-Eo CFU-MK Pre-B Pre-T CFU-baso - CFU E RBC PMN Mono. PMEo PLT Macrophage DC B cell NKT cell Drugs which would kill PMBspecifically the proliferating mutated cells (specificic JAK 2 inhibitor)

Conclusions • Modeling of normal hematopoiesis may be a powerful tool to predict complex biologic processes that will be validated by experimental procedures • Modeling of malignant hematopoiesis may help to understand the clonal dominance and the cooperative effects of several genetic alterations • Modeling might be a powerful tool to prospectively in vivo monitor the response to a drug and a combination of drugs • MPD appear to be disorders whose pathogenesis and treatment might be greatly improved by such an approach

Acknowledgments INSERM U 1009 Villejuif Eric Solary François Delhommeau Nicole Casadevall Stéphane Giraudier Rodolphe Besancenot Jean-Pierre Le Couédic Isabelle Plo Jean Luc Villeval Ludwig Institute for Cancer Research (Brussels) Stefan Constantinescu INSERM U 985 Villejuif Olivier Bernard Véronique Della Valle Roland Berger
- Slides: 37