Welcome webinar instructions The webinar will start soon

Welcome - webinar instructions • The webinar will start soon • Go. Training works best in Chrome or on Linux, Firefox • All microphones will be muted while the trainer is speaking • If you have a question please use the chat box at the bottom of the Go. Training box • Please complete the feedback survey which will launch at the end of the webinar • The webinar will be recorded and added to Train online

An Introductory Webinar Wojtek Bazant & Faye Rodgers https: //parasite. wormbase. org

Outline • Why Worm. Base Para. Site? • Our genomes • Data available • Bio. Mart • Questions

Why Worm. Base Para. Site? ● Helminths (parasitic roundworms and flatworms) are the causative agents of many diseases of humans, animals and plants ● Increasing amounts of genomic data are becoming available to the helminth research community ● Worm. Base Para. Site processes and presents that data in a consistent and accessible way
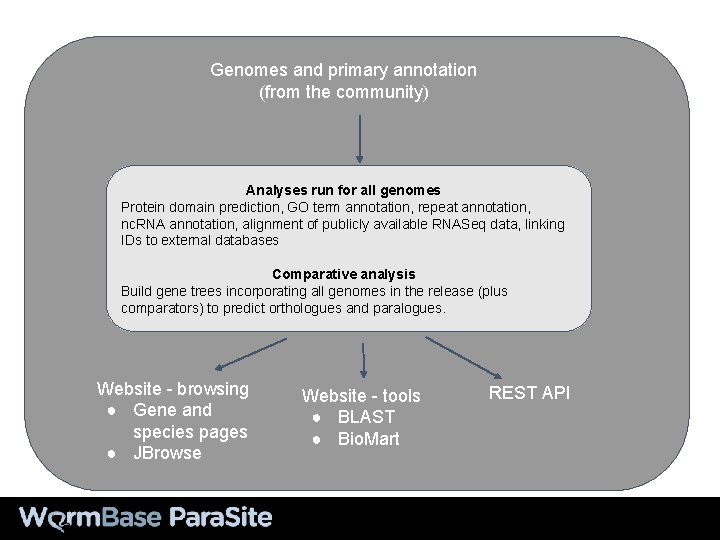
Genomes and primary annotation (from the community) Analyses run for all genomes Protein domain prediction, GO term annotation, repeat annotation, nc. RNA annotation, alignment of publicly available RNASeq data, linking IDs to external databases Comparative analysis Build gene trees incorporating all genomes in the release (plus comparators) to predict orthologues and paralogues. Website - browsing ● Gene and species pages ● JBrowse Website - tools ● BLAST ● Bio. Mart REST API
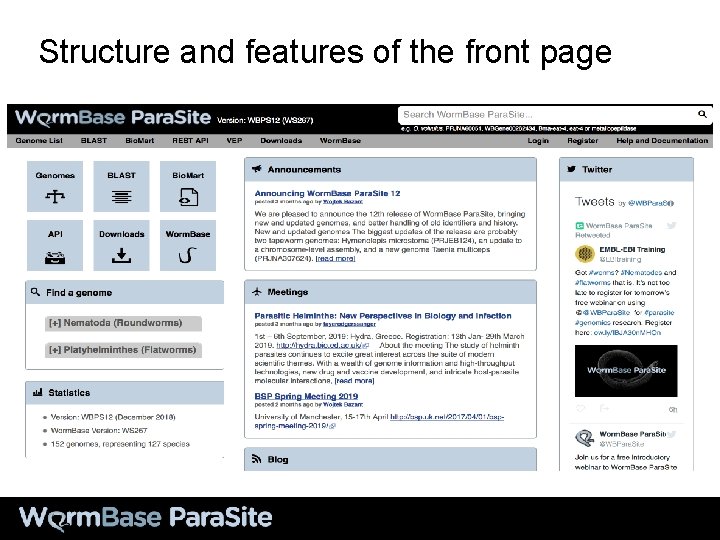
Structure and features of the front page
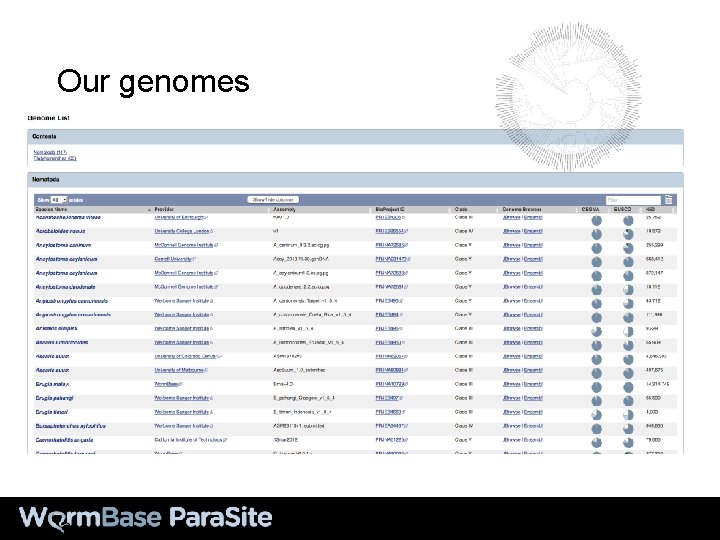
Our genomes
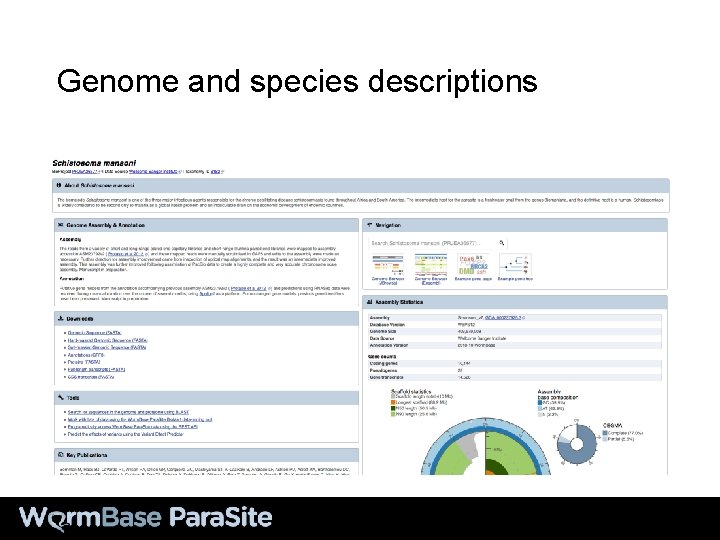
Genome and species descriptions

Finding information related to your scientific question If you know the gene name or ID, it’s just a search task! Otherwise, it more like research. Common avenues: • BLAST the sequence • Text search to try match a gene description • Search through a protein feature or GO term • Navigate through an orthologous gene in other species
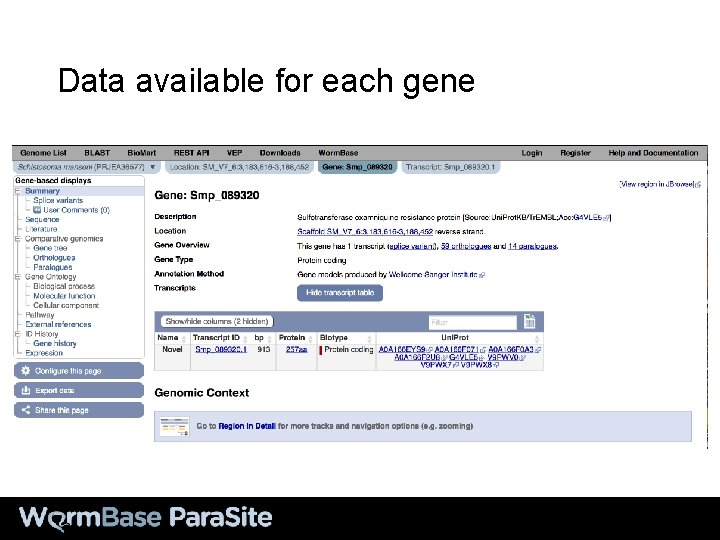
Data available for each gene
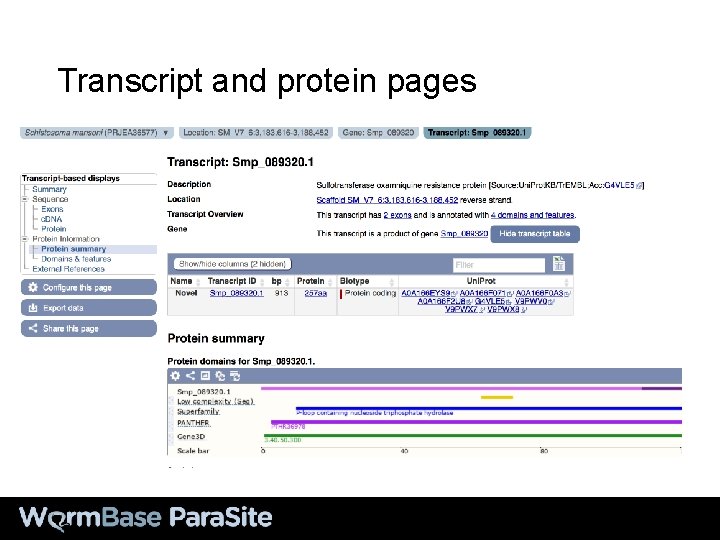
Transcript and protein pages
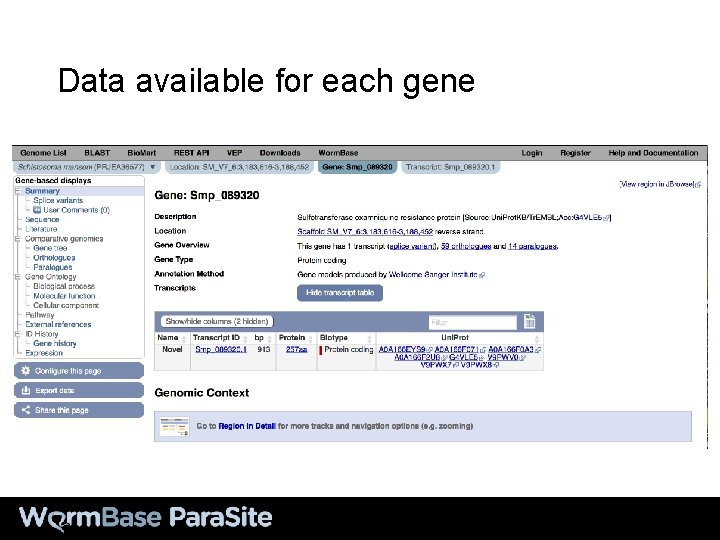
Data available for each gene

“Region in detail” - embedded genome browser
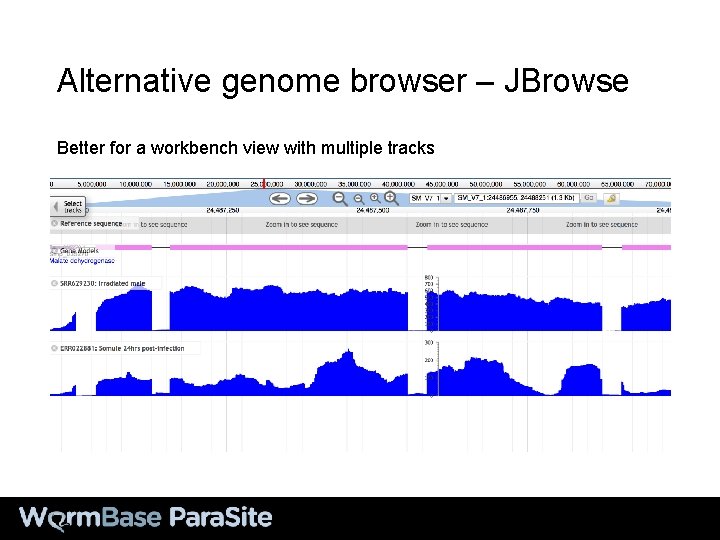
Alternative genome browser – JBrowse Better for a workbench view with multiple tracks
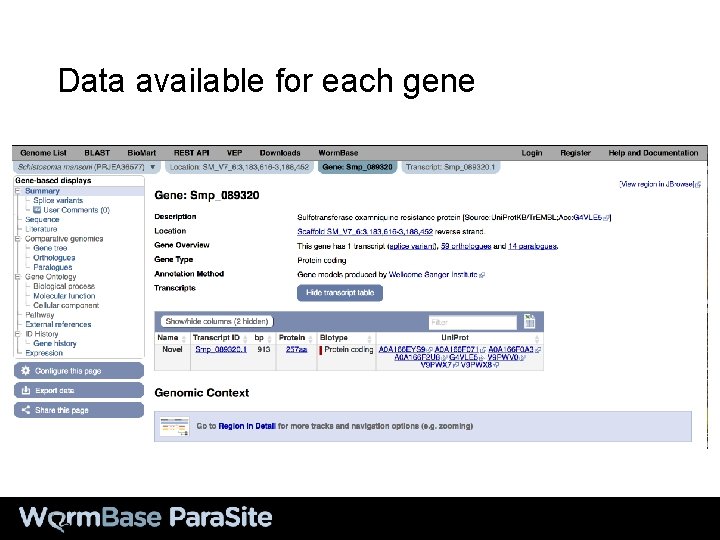
Data available for each gene

Links and references - Uni. Prot etc.
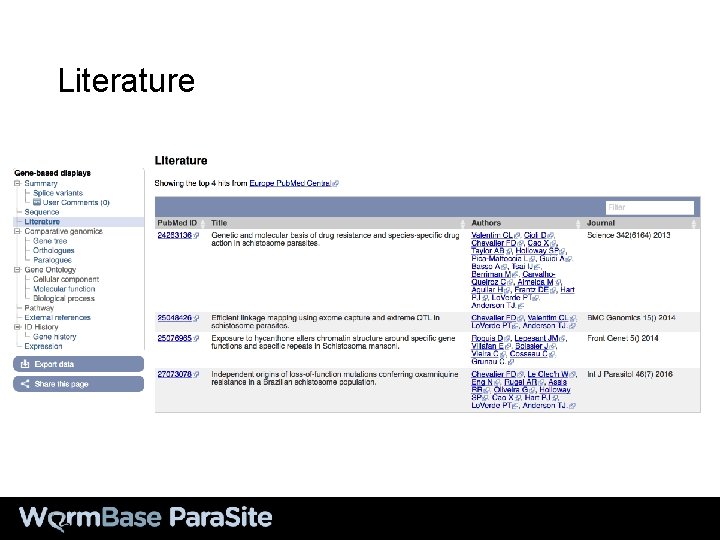
Literature

Comparative Genomics Gene trees are computed with every release, classifying genes into families. These are reconciled with the species trees to infer orthologous and paralogous relationships. Speciation node Duplication node Tree views can be configured for exploring the gene family https: //www. ensembl. org/info/genome/compara/homology_method. html
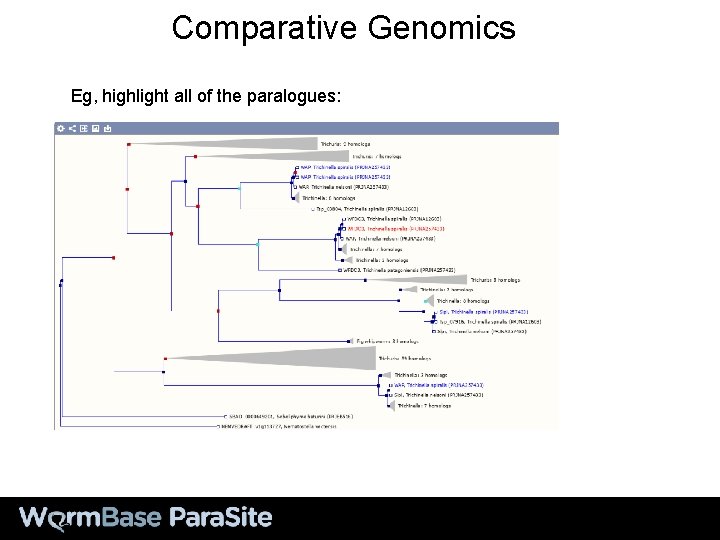
Comparative Genomics Eg, highlight all of the paralogues:

Comparative Genomics Orthologues and paralogues are also available in tabular format: ● Lists can be exported from Bio. Mart ● Full gene trees can be accessed programmatically via the API

Bio. Mart A very powerful tool for accessing data in bulk without any programming knowledge. Filters Values The data type you’re basing your query on, eg: The actual data you’re basing your query on, eg: Attributes The data you want, eg: Genome Schistosoma mansoni PRJEA 36577 Protein stable IDs Genomic region Schistosoma mansoni Sm_V 7_1 c. DNA sequences A list of gene IDs Smp_035270, Smp_010250, Smp_244010… Uniprot IDs All genes annotated with a protein domain or a GO term All genes that have an orthologue in a species Signal. P Genes with an orthologue in Schistosoma haematobium Protein domains Orthologue names, % identity Filters can be combined to build more complex queries

Bio. Mart Walk-through example: using Bio. Mart to retrieve S. mansoni genes from the ZW chromosome that have an orthologue in S. japonicum and S. haematobium. Want to return the S. mansoni, S. haematobium and S. japonicum gene IDs.
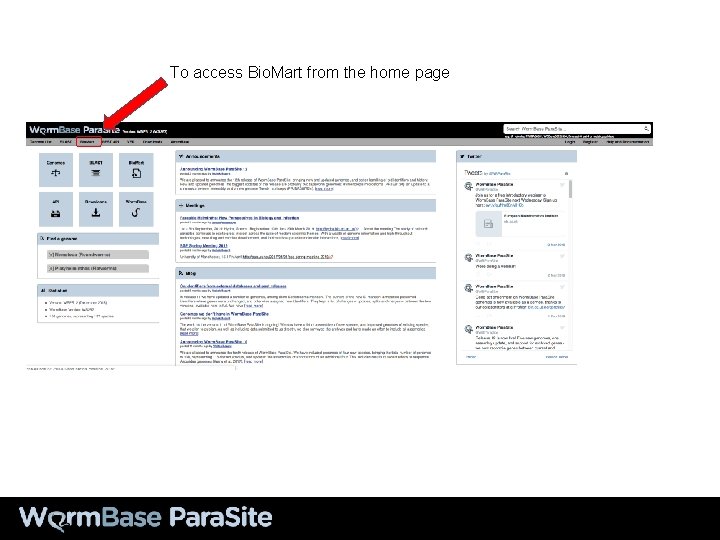
To access Bio. Mart from the home page
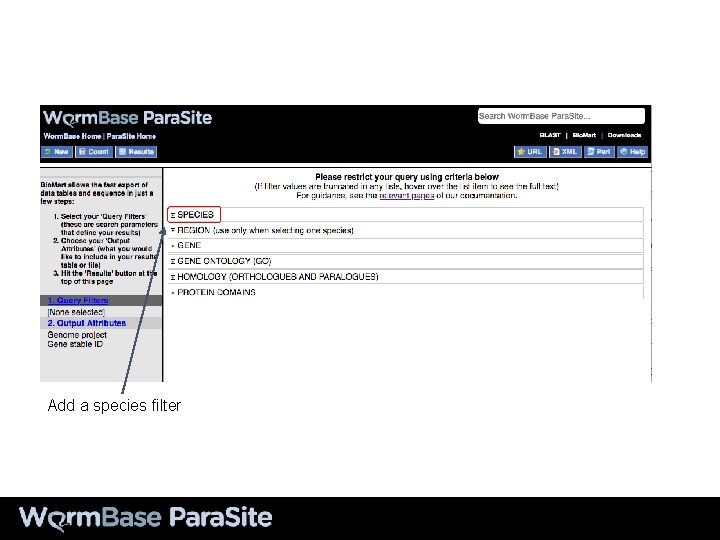
Add a species filter
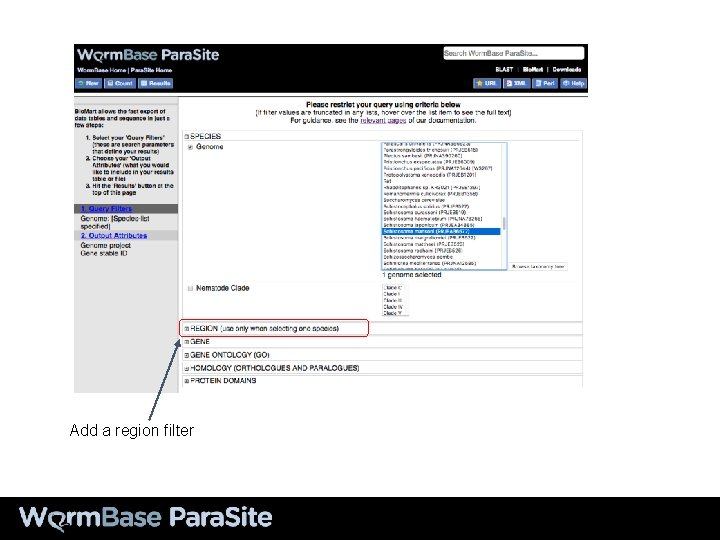
Add a region filter
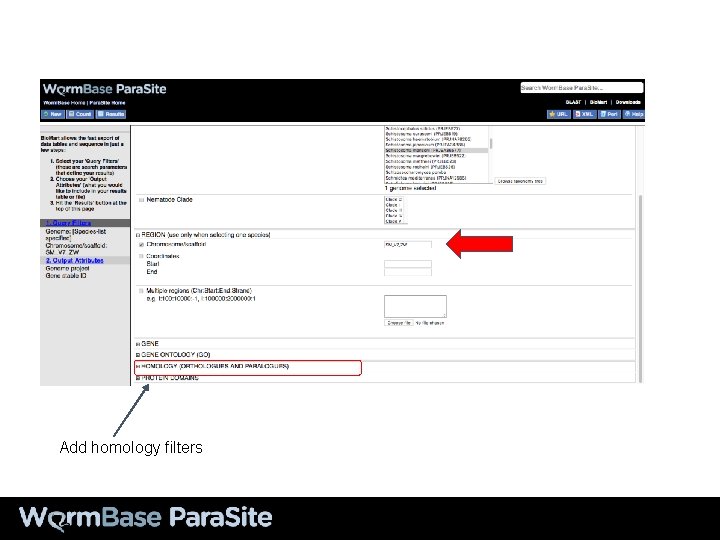
Add homology filters
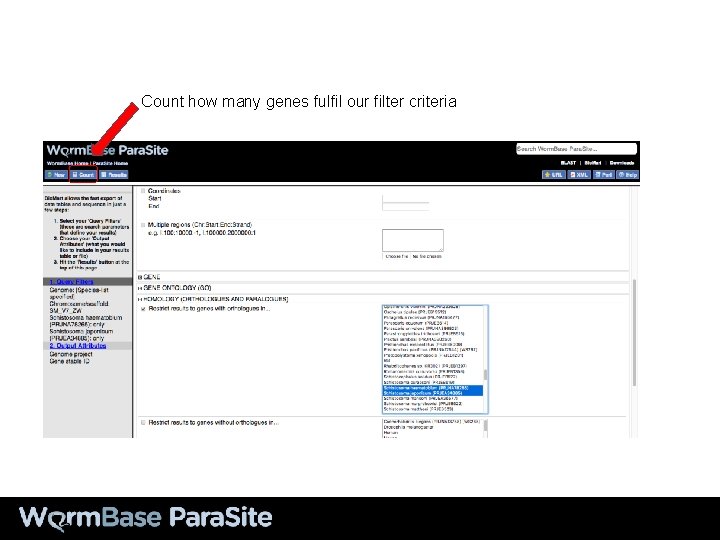
Count how many genes fulfil our filter criteria
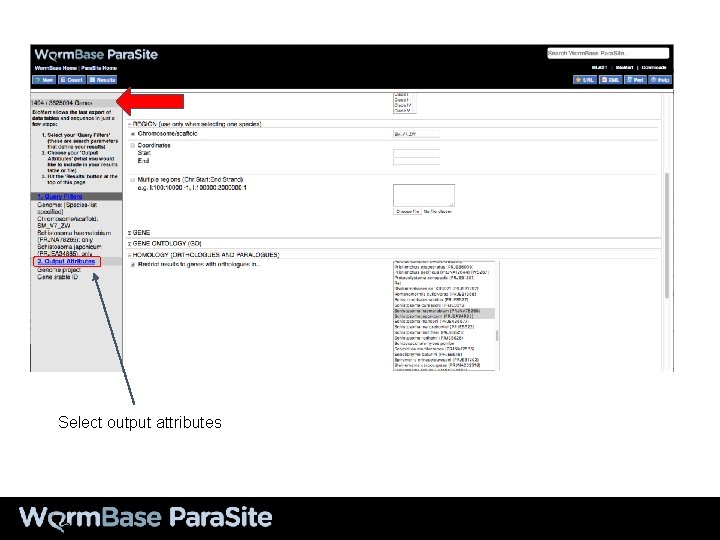
Select output attributes
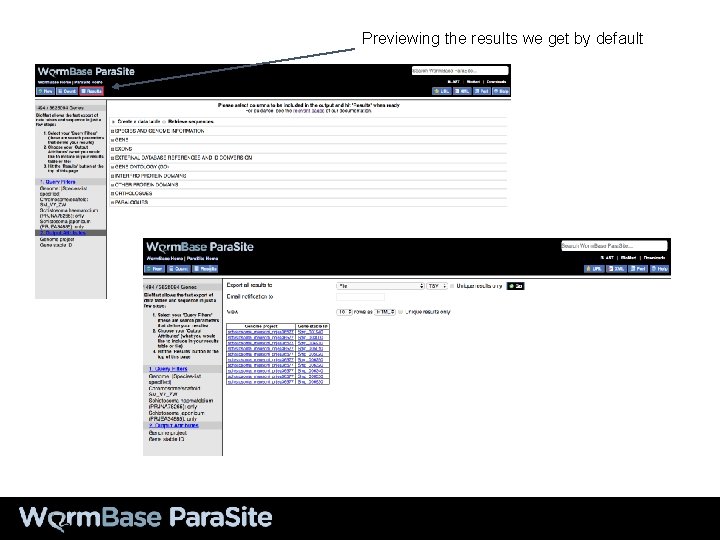
Previewing the results we get by default
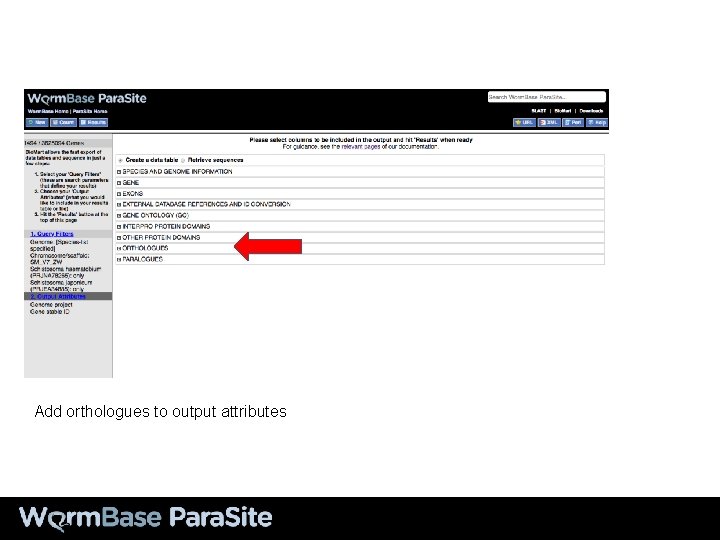
Add orthologues to output attributes

Scroll down to find the species that we’re interested in
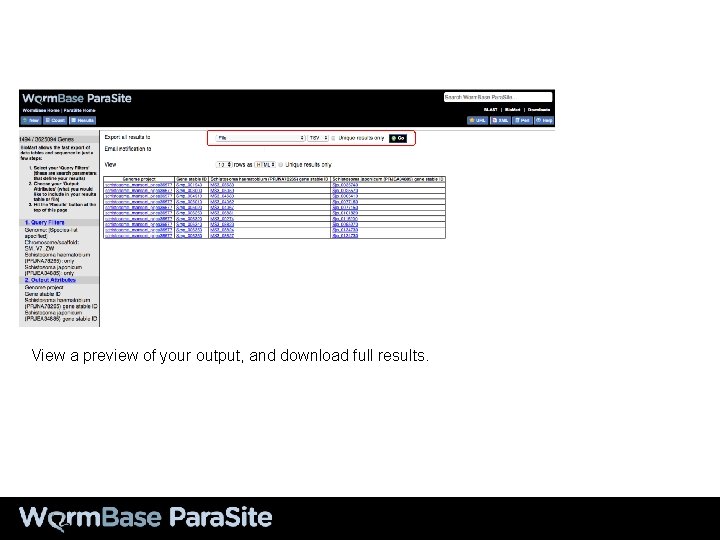
View a preview of your output, and download full results.

Bio. Mart Other examples of questions that can be answered with Bio. Mart: ● For a list of gene IDs: ○ Convert to other types of identifier (Uniprot, Ref. Seq, NCBI) ○ Retrieve associated protein domains, GO terms ○ Retrieve their genomic coordinates ○ Generate FASTA files of protein, c. DNA, UTR, flanking region sequences etc ● Retrieve a list of genes that: ○ Have a given protein domain/GO term ○ Have/do not have orthologues in species X, Y, Z. ○ Are on genomic region X For R users, Worm. Base Para. Site Bio. Mart supports the bioma. Rt R package: see our help and documentation pages to get started.

Outline • Why Worm. Base Para. Site? • Our genomes • Data available • Bio. Mart • Questions

Outline • Why Worm. Base Para. Site? • Our genomes • Data available • Bio. Mart • Questions If we don’t get to your question: email parasite-help@sanger. ac. uk

Sample question I need the sequences for a set of Schistosoma mansoni genes. I have the chromosome, start, and stop for each. The suggested option Other, more creative approaches? - download the GFF and the sequence files from the FTP, and write a program - check the cases one by one - use the API, first „region” endpoint to get gene IDs, then „sequence” endpoint - email the helpdesk ( it might work )

Bio. Mart Example 2 Using Bio. Mart to generate a protein FASTA file from a list of gene IDs
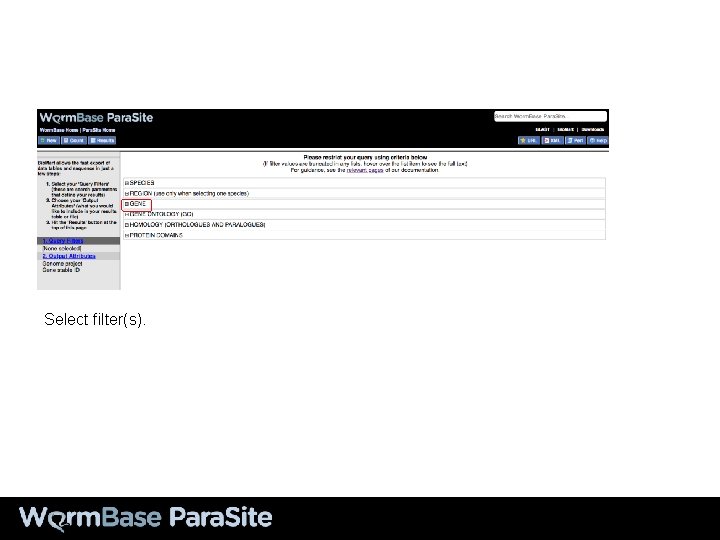
Select filter(s).
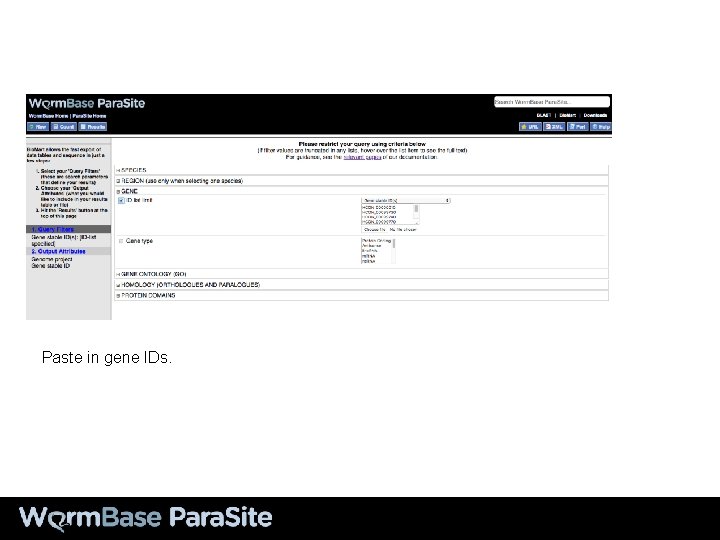
Paste in gene IDs.
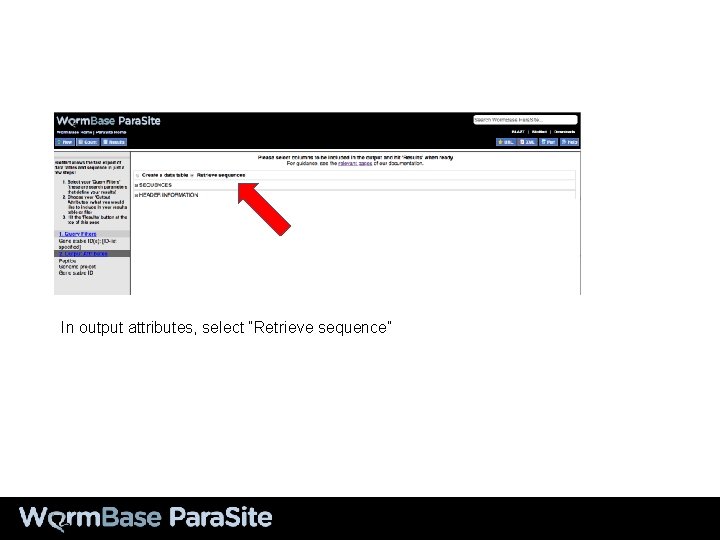
In output attributes, select “Retrieve sequence”
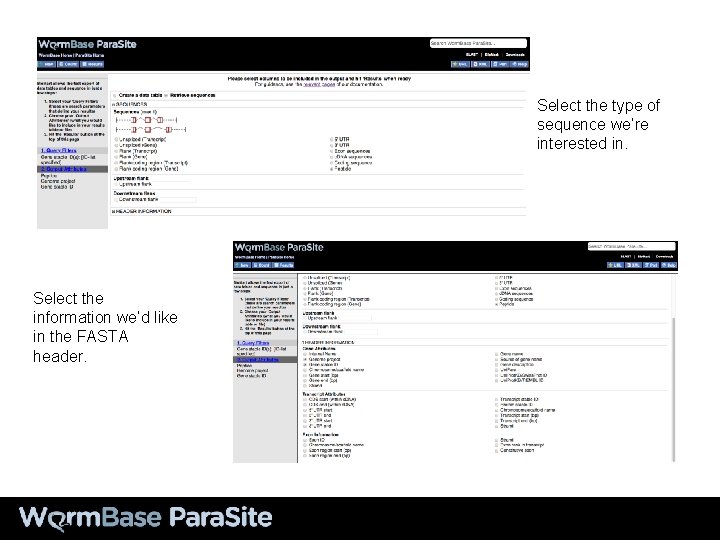
Select the type of sequence we’re interested in. Select the information we’d like in the FASTA header.
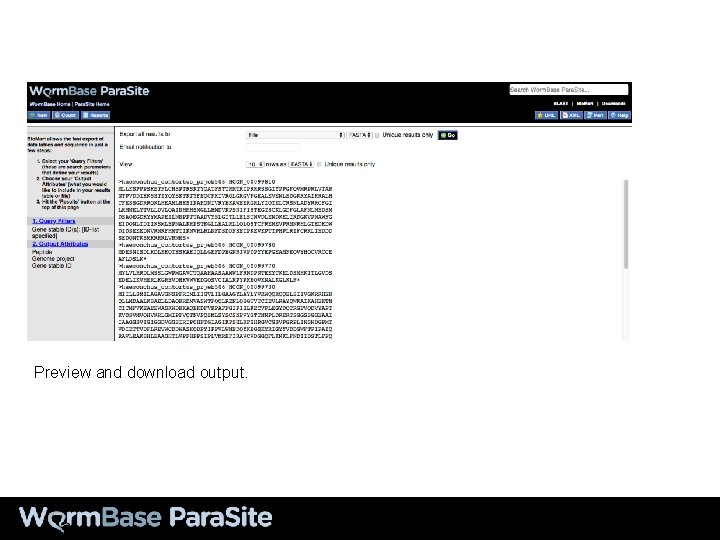
Preview and download output.
Upcoming webinars See the full list of upcoming webinars at https: //www. ebi. ac. uk/training/webinars Don’t forget! Please fill in the survey that launches after the webinar – thanks!
- Slides: 43