Welcome Mass Spectrometry meets Cheminformatics WCMC Metabolomics Course

Welcome! Mass Spectrometry meets Cheminformatics WCMC Metabolomics Course 2013 Tobias Kind Course 4: Mass Spectrometry Tools & Concepts http: //fiehnlab. ucdavis. edu/staff/kind 1 CC-BY License
![Atomic Mass Correct unit is [u] – unified atomic mass unit or [Da] Dalton Atomic Mass Correct unit is [u] – unified atomic mass unit or [Da] Dalton](http://slidetodoc.com/presentation_image/dce1da431cd1b6629ed75b5285c6f387/image-2.jpg)
Atomic Mass Correct unit is [u] – unified atomic mass unit or [Da] Dalton see SI units 1 u = 1 Da = 1/12 th of mass of carbon 12 C = 1. 66053886 x 10 -27 kg Hexachlorobenzene (C 6 Cl 6) average mass - 284. 7804 u integer mass - 282. 0 u monoisotopic mass - 281. 81312 u Always (always) check molecular masses obtained from databases or publications. For mass spectrometry the monoisotopic massis used. In. Ch. IKey: CKAPSXZOOQJIBF-UHFFFAOYAV 2
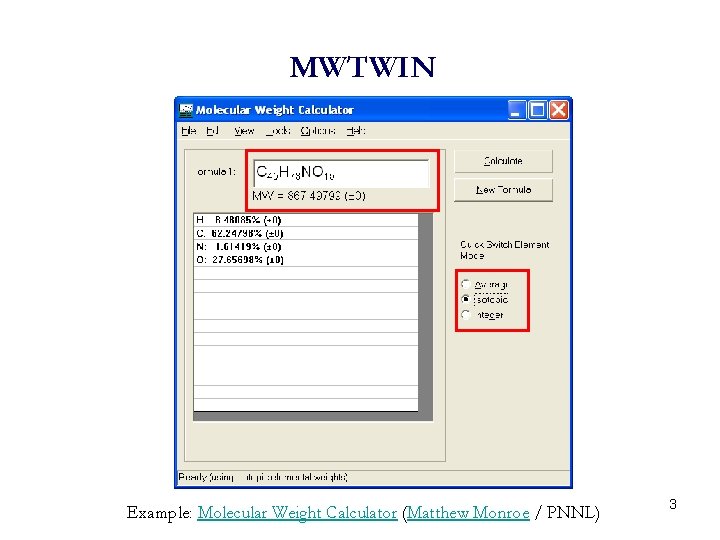
MWTWIN Example: Molecular Weight Calculator (Matthew Monroe / PNNL) 3

Mass Accuracy Instruments must be calibrated to obtain high mass accuracy. In case of FT-ICR-MS mass calibration can be stable over weeks. Post- mass calibration can be performed if calibrant was run with samples. Mass of electron becomes important at around 500 Da. Type Mass Accuracy FT-ICR-MS 0. 1 - 1 ppm Orbitrap 0. 5 - 1 ppm Magnetic Sector 1 - 2 ppm TOF-MS 3 - 5 ppm Q-TOF 1 - 5 ppm Triple Quad 3 - 5 ppm Linear Ion. Trap 50 -200 ppm (10 ppm in Ultra-Zoom) m(e-) = 0. 00054857990924 u = mass of electron m(1 H+) = 1. 00727645199076 u = mass of proton m(1 H) = 1. 0078250319 4

Resolving Power RP = 1700 High resolving power is helpful for separation of species with almost same mass (isobars). RP = 48, 250 High resolving power can not be used to distinguish between structural isomers. Example: C 8 H 10 N 2 O has 100, 082, 479 isomers. 5 Example Solanine (CID=30185)
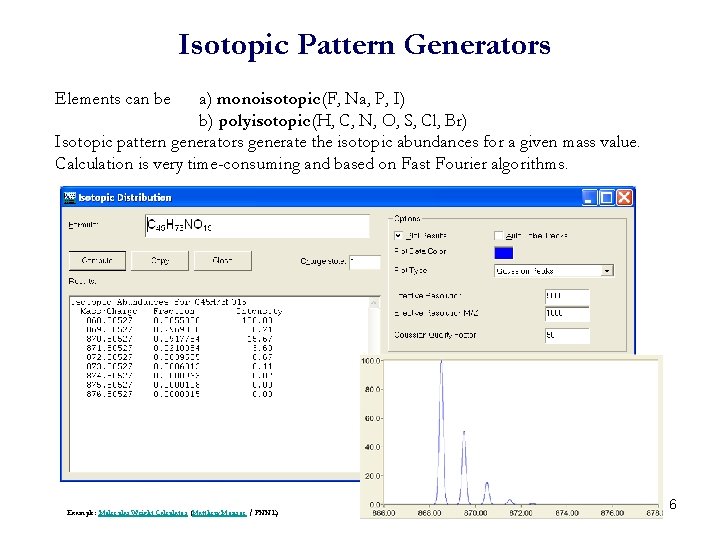
Isotopic Pattern Generators Elements can be a) monoisotopic (F, Na, P, I) b) polyisotopic (H, C, N, O, S, Cl, Br) Isotopic pattern generators generate the isotopic abundances for a given mass value. Calculation is very time-consuming and based on Fast Fourier algorithms. Example: Molecular Weight Calculator (Matthew Monroe / PNNL) 6

Isotopic pattern generators Example form Thermo Xcalibur with a very versatile isotopic pattern generator 7
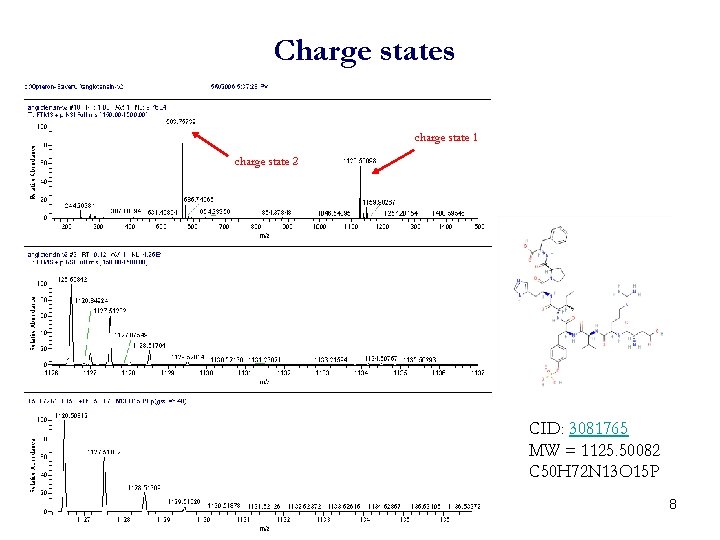
Charge states charge state 1 charge state 2 CID: 3081765 MW = 1125. 50082 C 50 H 72 N 13 O 15 P 8
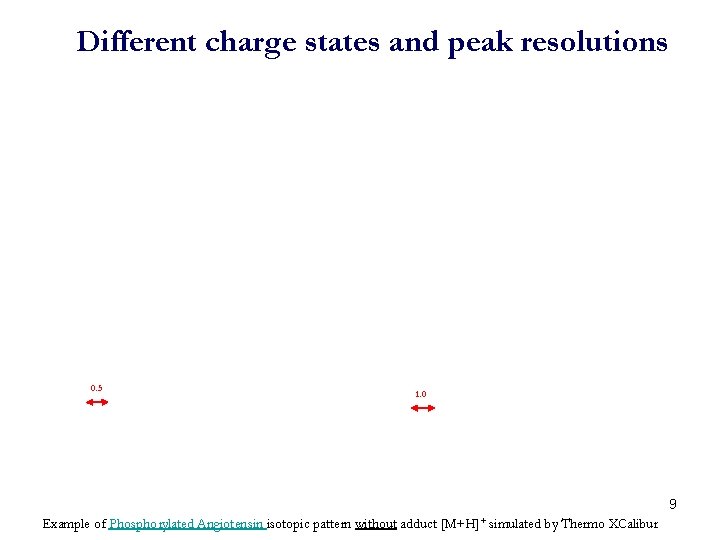
Different charge states and peak resolutions 0. 5 1. 0 9 Example of Phosphorylated Angiotensin isotopic pattern without adduct [M+H]+ simulated by Thermo XCalibur
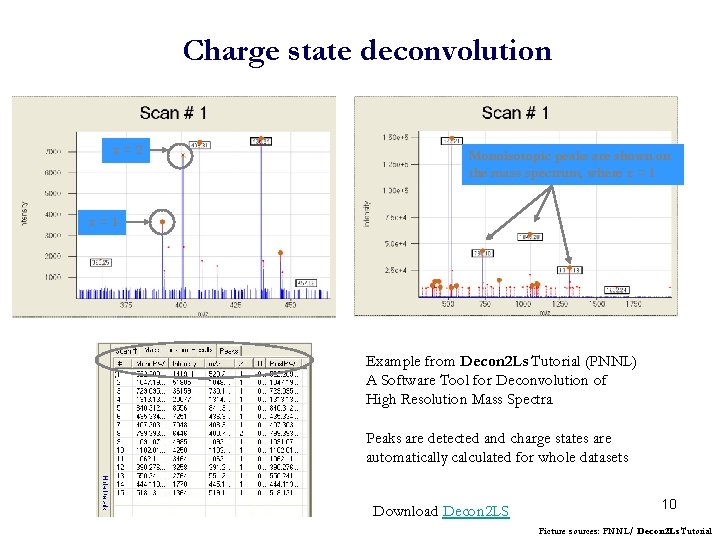
Charge state deconvolution z=2 Monoisotopic peaks are shown on the mass spectrum, where z = 1 z=1 Example from Decon 2 Ls Tutorial (PNNL) A Software Tool for Deconvolution of High Resolution Mass Spectra Peaks are detected and charge states are automatically calculated for whole datasets Download Decon 2 LS 10 Picture sources: PNNL/ Decon 2 Ls Tutorial

Adduct formation is observed in ESI, CI and APCI ionization modes (and others). Adduct detection is problematic for small molecules, can be influenced by solvent selection. Adduct detection can be automated if two or more adducts are detected in mass spectrum. Switching polarity (+/-) can be used for confirmation of adduct. Download the Mass Spectral Adduct Calculator Check: MZed. DB Tools for the annotation of High Resolution MS metabolomics data 11 http: //fiehnlab. ucdavis. edu/staff/kind/Metabolomics/MS-Adduct-Calculator/

Adduct formation – expect the unexpected …around 290 different adducts Statistics: Adducts in NIST 12 MS/MS DB (80, 000 spectra) Most common adducts for LC-MS ([M+H]+ [M+Na]+ [M+NH 4]+ [M+acetate]+) 12

Molecular Formula Generators Formula generators are used to create molecular formulae from accurate masses. Input requires 1) accurate isotopic mass (with or without adduct) and 2) error in ppm or m. Da (milli Dalton) Accurate mass Example MWTWIN 13
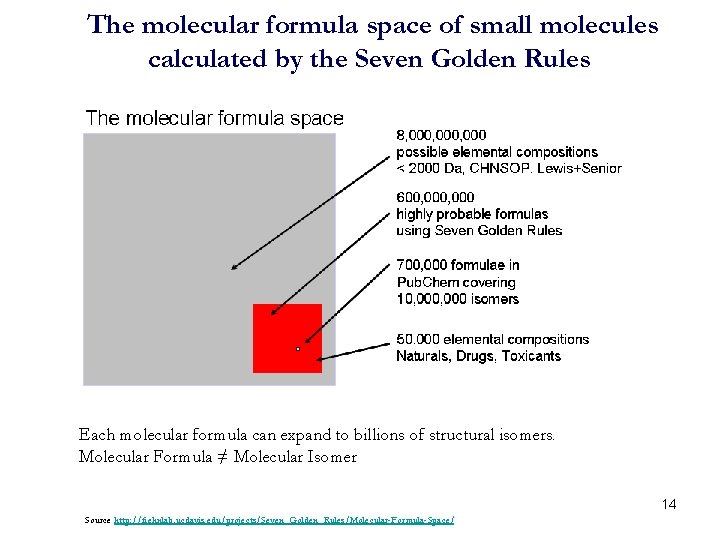
The molecular formula space of small molecules calculated by the Seven Golden Rules Each molecular formula can expand to billions of structural isomers. Molecular Formula ≠ Molecular Isomer 14 Source http: //fiehnlab. ucdavis. edu/projects/Seven_Golden_Rules/Molecular-Formula-Space/
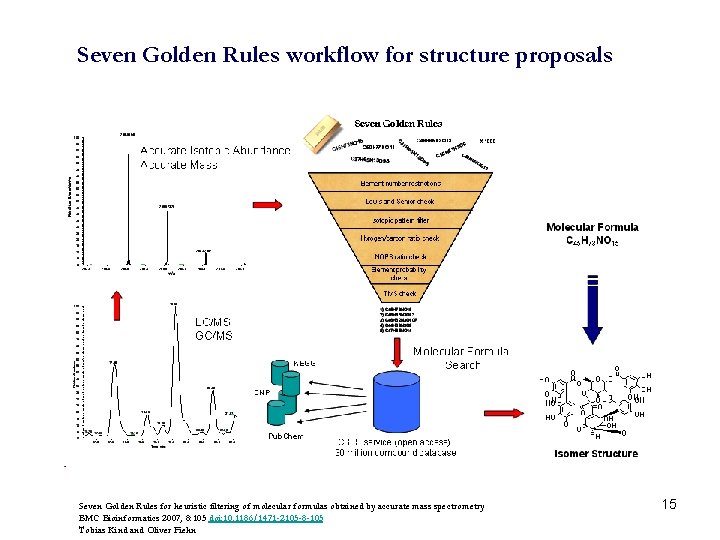
Seven Golden Rules workflow for structure proposals Seven Golden Rules for heuristic filtering of molecular formulas obtained by accurate mass spectrometry BMC Bioinformatics 2007, 8: 105 doi: 10. 1186/1471 -2105 -8 -105 Tobias Kind and Oliver Fiehn 15
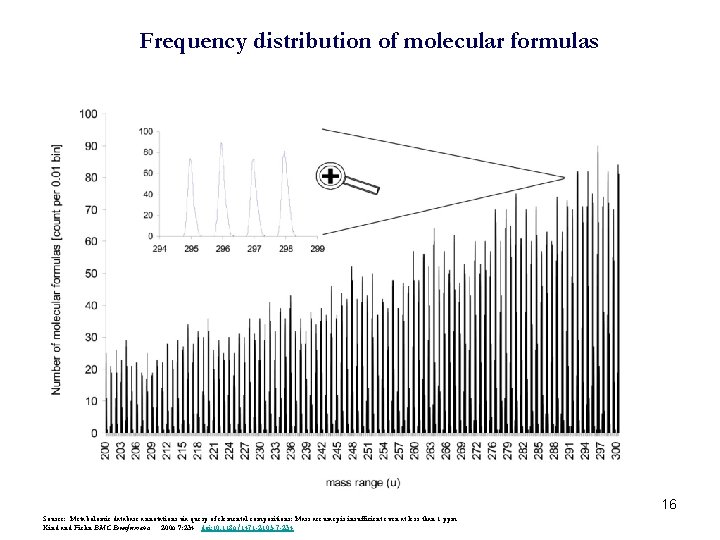
Frequency distribution of molecular formulas 16 Source: Metabolomic database annotations via query of elemental compositions: Mass accuracy is insufficient even at less than 1 ppm Kind and Fiehn BMC Bioinformatics 2006 7: 234 doi: 10. 1186/1471 -2105 -7 -234
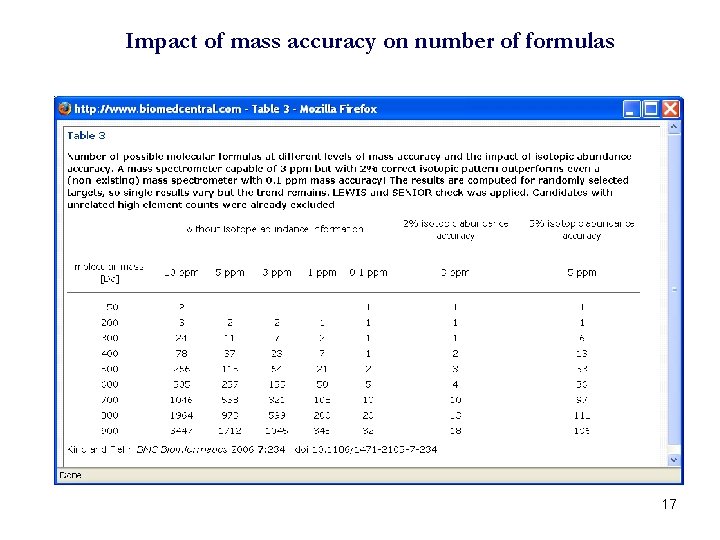
Impact of mass accuracy on number of formulas 17
![Mass accuracy and isotopic pattern [M+H]+ C 45 H 73 NO 15 MW = Mass accuracy and isotopic pattern [M+H]+ C 45 H 73 NO 15 MW =](http://slidetodoc.com/presentation_image/dce1da431cd1b6629ed75b5285c6f387/image-18.jpg)
Mass accuracy and isotopic pattern [M+H]+ C 45 H 73 NO 15 MW = 867. 49799 Example: ESI-MS (+) of Solanine on a LTQ Resolving Power: 1700 Mass Accuracy: 46 ppm Isotopic Abundance Error: ± 1. 46% 18 Example from: http: //fiehnlab. ucdavis. edu/projects/Seven_Golden_Rules/Examples/Solanine/
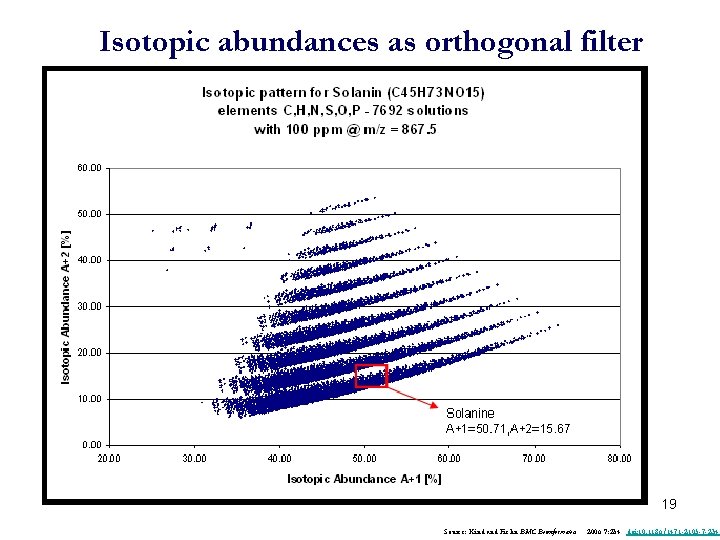
Isotopic abundances as orthogonal filter 19 Source: Kind and Fiehn BMC Bioinformatics 2006 7: 234 doi: 10. 1186/1471 -2105 -7 -234
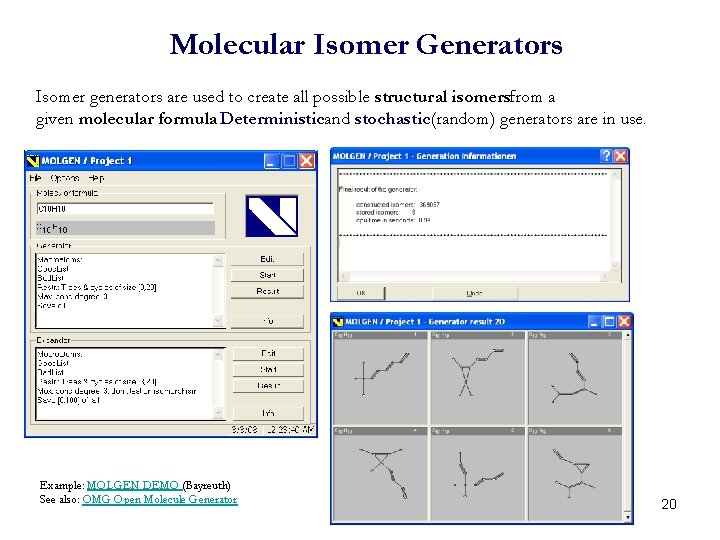
Molecular Isomer Generators Isomer generators are used to create all possible structural isomersfrom a given molecular formula. Deterministicand stochastic(random) generators are in use. Example: MOLGEN DEMO (Bayreuth) See also: OMG Open Molecule Generator 20
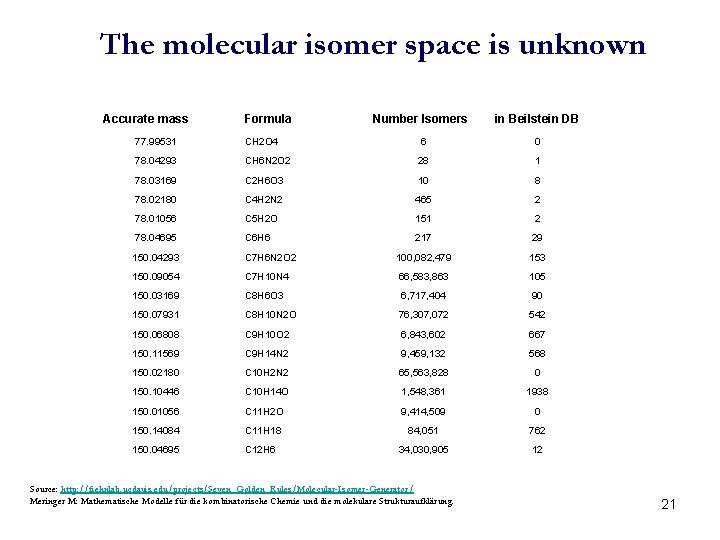
The molecular isomer space is unknown Accurate mass Formula Number Isomers in Beilstein DB 77. 99531 CH 2 O 4 6 0 78. 04293 CH 6 N 2 O 2 28 1 78. 03169 C 2 H 6 O 3 10 8 78. 02180 C 4 H 2 N 2 465 2 78. 01056 C 5 H 2 O 151 2 78. 04695 C 6 H 6 217 29 150. 04293 C 7 H 6 N 2 O 2 100, 082, 479 153 150. 09054 C 7 H 10 N 4 66, 583, 863 105 150. 03169 C 8 H 6 O 3 6, 717, 404 90 150. 07931 C 8 H 10 N 2 O 76, 307, 072 542 150. 06808 C 9 H 10 O 2 6, 843, 602 667 150. 11569 C 9 H 14 N 2 9, 459, 132 568 150. 02180 C 10 H 2 N 2 65, 563, 828 0 150. 10446 C 10 H 14 O 1, 548, 361 1938 150. 01056 C 11 H 2 O 9, 414, 509 0 150. 14084 C 11 H 18 84, 051 762 150. 04695 C 12 H 6 34, 030, 905 12 Source: http: //fiehnlab. ucdavis. edu/projects/Seven_Golden_Rules/Molecular-Isomer-Generator/ Meringer M: Mathematische Modelle für die kombinatorische Chemie und die molekulare Strukturaufklärung. 21
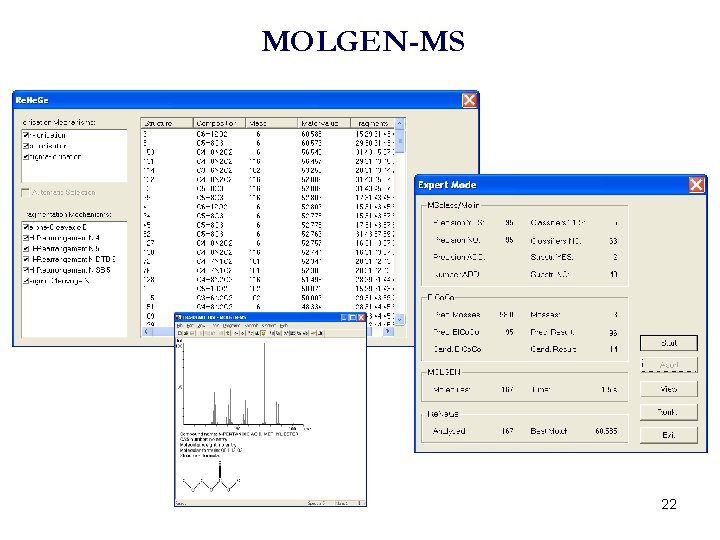
MOLGEN-MS 22

Substructure predictions Use computer algorithms (machine learning) for automated interpretation of fragments and corresponding substructures. Algorithms creates present/absent list of substructures. 23 Example from NIST-MS search
Simulation of mass spectra Why is simulation or prediction of mass spectra important? Molecular isomers (structures) can be generated very fast from molecular formulas Only certain mass spectra can be simulated (MS/MS of peptides, oligosaccharides, lipids) Problematic is abundance determination Problematic are all complex rearrangement reactions (gas phase ion chemistry) Simulation of mass spectra from small molecules is new and important research Experimental spectra Simulated spectra Perform comparison or matching 24
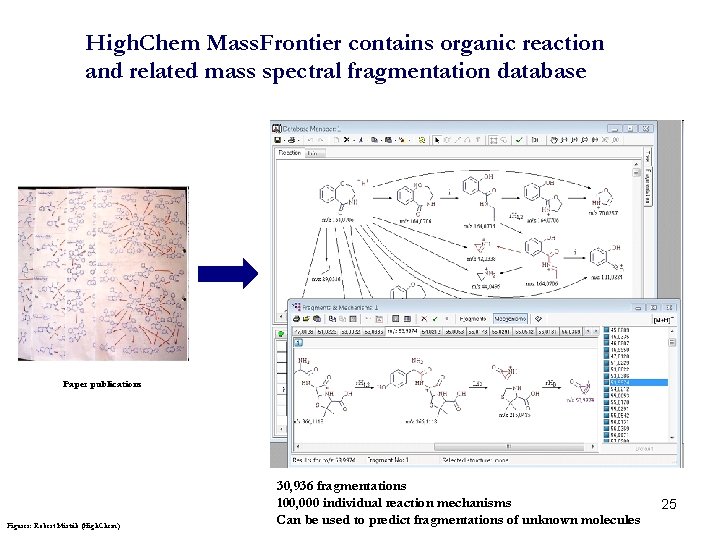
High. Chem Mass. Frontier contains organic reaction and related mass spectral fragmentation database Paper publications Figures: Robert Mistrik (High. Chem) 30, 936 fragmentations 100, 000 individual reaction mechanisms Can be used to predict fragmentations of unknown molecules 25
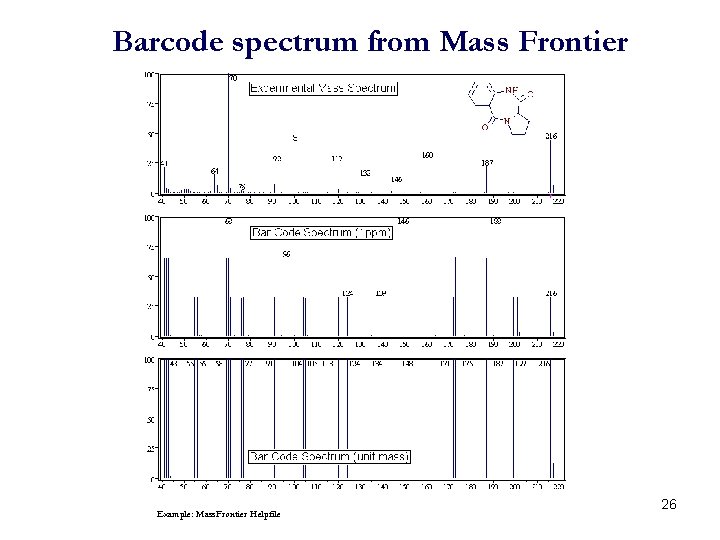
Barcode spectrum from Mass Frontier Example: Mass. Frontier Helpfile 26
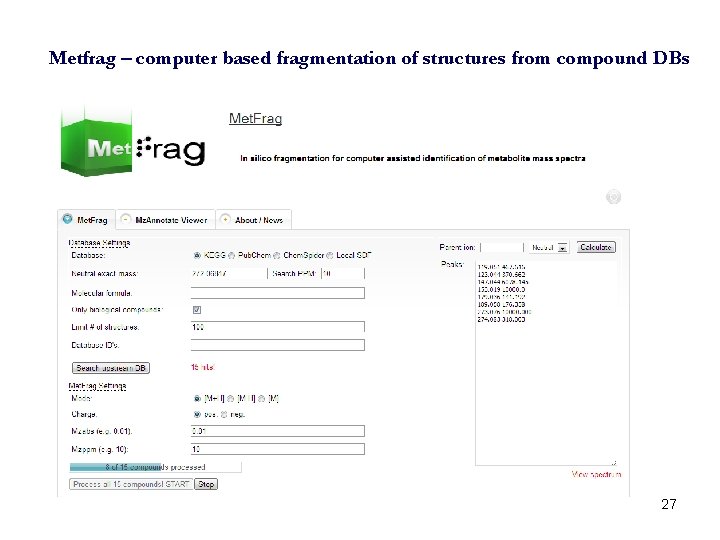
Metfrag – computer based fragmentation of structures from compound DBs 27
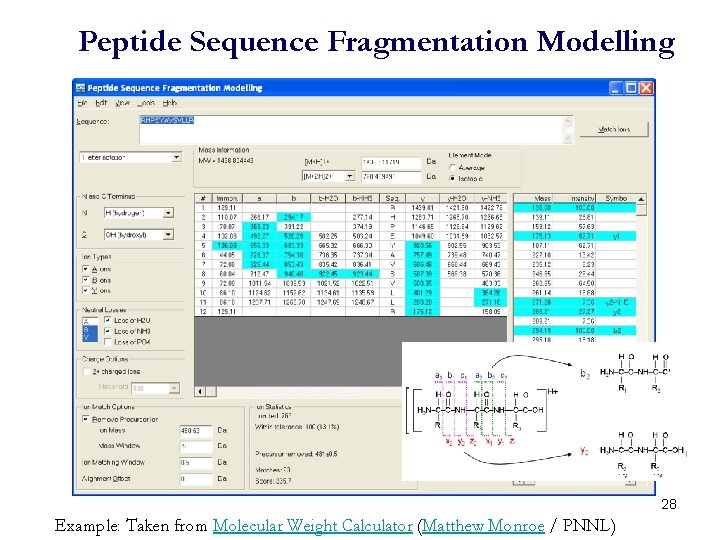
Peptide Sequence Fragmentation Modelling 28 Example: Taken from Molecular Weight Calculator (Matthew Monroe / PNNL)
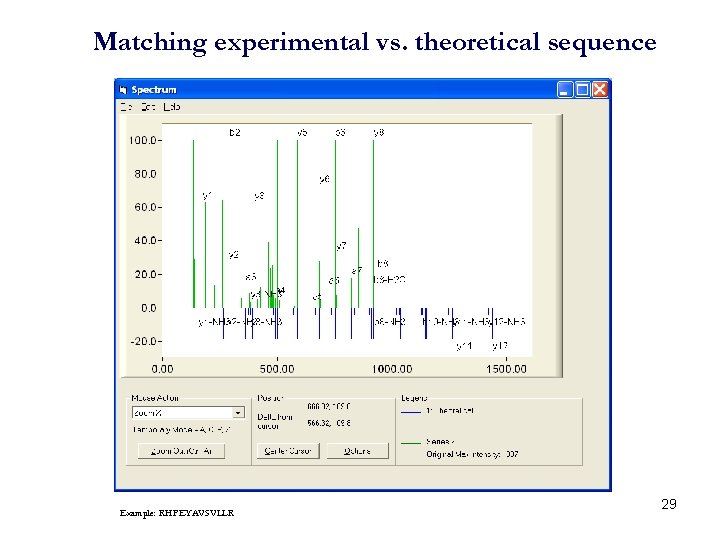
Matching experimental vs. theoretical sequence Example: RHPEYAVSVLLR 29
![Lipid. Blast in-silico MS/MS spectra Experimental MS/MS precursor m/z = 732. 55 [M+H]-sn 1 Lipid. Blast in-silico MS/MS spectra Experimental MS/MS precursor m/z = 732. 55 [M+H]-sn 1](http://slidetodoc.com/presentation_image/dce1da431cd1b6629ed75b5285c6f387/image-30.jpg)
Lipid. Blast in-silico MS/MS spectra Experimental MS/MS precursor m/z = 732. 55 [M+H]-sn 1 [M+H]-sn 2 [M+H]-sn 1 -H 2 O [M+H]-sn 2 -H 2 O [M+H]-C 5 H 14 NO 4 P (-183) in-silico MS/MS match precursor m/z = 732. 55 [M+H]-H 2 O (-18) [M+H]-C 3 H 9 N (-59) 30 Nature Methods. 2013 Aug; 10(8): 755 -8. doi: 10. 1038/nmeth. 2551. Lipid. Blast in silico tandem mass spectrometry database for lipid identification. Kind T, Liu KH, Lee do Y, Defelice B, Meissen JK, Fiehn O. Free download: Lipid. Blast

The Last Page - What is important to remember: Important performance parameters are accurate mass, resolving power, scan speed and accurate isotopic abundances Even high resolving power MS can not distinguish between structural isomers Accurate Mass Molecular Formula Structural Isomers MS/MS Mass spectra of only some substance classes can be simulated Only NMR can perform de-novo structure elucidation in an consistent manner Of general importance for this course: http: //fiehnlab. ucdavis. edu/staff/kind/Metabolomics/Structure_Elucidation/ Advances in structure elucidation of small molecules using mass spectrometry (Kind & Fiehn 2010) 31
- Slides: 31