Web Databases for Drosophila Introduction to Fly Base

Web Databases for Drosophila Introduction to Fly. Base and Ensembl Database Wilson Leung 6/06

Outline n n n Introduction to Fly. Base Introduction to Ensembl Using web databases to assist annotation of novel sequences

Introduction to Fly. Base Available at http: //www. flybase. org

Introduction to Fly. Base n Fly. Base is primarily funded by the National Institutes of Health n Fly. Base consortium includes Drosophila researchers and computer scientists at Harvard University, Indiana University, and University of Cambridge, plus scientists worldwide n In addition to the main site at www. flybase. org, there also many mirror sites

What is Fly. Base? n It is a comprehensive database of genetic and molecular data for many Drosophila species: n n n Information on genes and mutant alleles Expression and function of gene products Genetic, cytological, molecular map information Data from Berkeley Drosophila Genome Project Data from European Drosophila Genome Project
Introduction to Ensembl Available at http: //www. ensembl. org

What is Ensembl? n Ensembl is a joint project between the European Bioinformatics Institute (EBI) and the Wellcome Trust Sanger Institute n Ensembl seeks to develop an automated system for the production and maintenance of annotations on eukaryotic genomes n These annotations should also be easily accessible to researchers

What is Ensembl? n While originally developed for eukaryotes, the Ensembl system has also been used to analyze prokaryotic genomes n n EBI Genome Review (archaea and bacteria) Most recent version is v 38 (Apr 2006) n Genomes available include human, chimp, mouse, dog, C. elegans, fruit fly, honey bee, mosquito among others
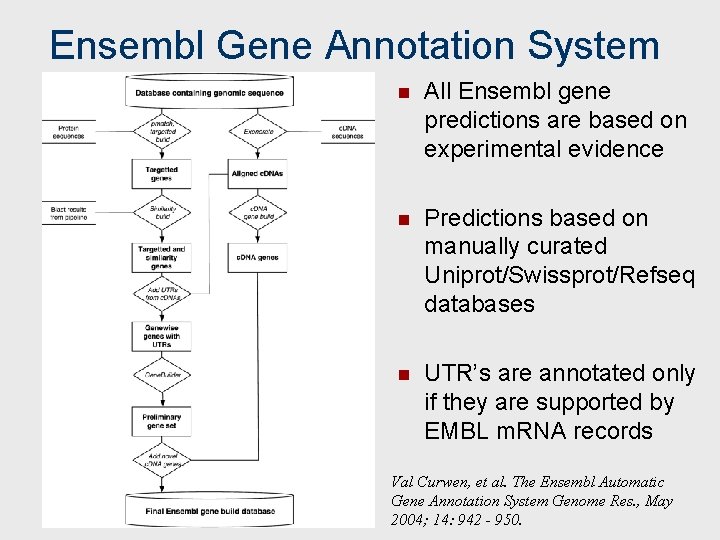
Ensembl Gene Annotation System n All Ensembl gene predictions are based on experimental evidence n Predictions based on manually curated Uniprot/Swissprot/Refseq databases n UTR’s are annotated only if they are supported by EMBL m. RNA records Val Curwen, et al. The Ensembl Automatic Gene Annotation System Genome Res. , May 2004; 14: 942 - 950.

Using Web Databases for Annotation List of available species in the Fly. Base BLAST service to use in a search for sequences homologous to your query Exon View in Ensembl: used to obtain sequence of a gene, exon-by-exon

Using Web Databases for Annotation n Motivations for using Fly. Base n n n Learn the biological functions of the gene of interest Use Fly. Base BLAST service to detect sequence homology to Drosophila species or species related to Drosophila Motivations for using Ensembl n n Obtain records of gene from multiple databases Obtain coding sequence of each exon of a gene

Walkthrough n Typical use of web databases is to identify putative homolog to a D. melanogaster gene n We have a novel 20 kb sequence from D. erecta n n n Using Repeat. Masker, we masked all drosophilaspecific repeats from the sequence Using blastx, we searched this sequence against the Swissprot database blastx results indicate our sequence is similar to the Paired-box protein (Pax 6) in D. melanogaster

Function of Pax-6 n Clicking on the accession number of the first hit in the blastx output shows that Pax-6 is also known as eyeless n We can learn more about eyeless using the Fly. Base web site @ http: //flybase. org n Type in eyeless in the search field, then click on the hit “ey” (#17)

Function of Pax-6 n This brings up the gene report for eyeless in D. melanogaster n We find that eyeless is important for brain and eye development n It is expressed in embryo, larva, and adult n Phenotypic changes in mutants include changes in the antenna, arista, and eye of the fruit fly

Finding Homologs in Other Species n Click on the BLAST button to access the BLAST service n Search our masked sequence against D. melanogaster, D. yakuba, D. mojavensis, D. virilis genome assemblies using blastn n Most of the species, other than D. melanogaster, are unannotated. n Nonetheless, this is useful for finding putative orthologs and for discovering regulatory regions using multiple sequence alignments

Using the Ensembl Database n Navigate to Ensembl @ http: //www. ensembl. org n Click on “Drosophila melanogaster” to access the data specific for this species n In the search box, type in the name “eyeless” then click “Go” n We find only one match - CG 1464 (the eyeless protein)

Transcripts of eyeless n There are four different isoforms of eyeless in D. melanogaster n We would typically annotate the most “comprehensive” isoform • In this case, isoform D n The Fruitfly Gene. View provides a general overview of the gene structure and function of eyeless n Links to Fly. Base, Ref. Seq, Swiss-Prot, EMBL records of eyeless are also available on this page.

Obtaining Transcript Sequence n Click on “Exon Info” for the transcript CG 1464 -RD n This bring us to the exon report for this transcript n n 9 exons, 3024 bps, 898 residues The sequence is shown with each exon in its own block. Sequence is color-coded: n n Purple = UTR’s Black = Coding DNA sequences (CDS) Blue = intronic sequences Green = upstream or downstream sequences

Obtaining Peptide Sequence n Click on the link “Protein Information” to obtain the peptide sequence of CG 1464 -RD n This bring us to the protein report for this transcript n n n “Protein Family” section shows that there are six gene members in this species Clicking on the link brings up the Family view - allows visualization of multiple sequence alignments of members of this family The peptide sequence has the following color-code: n n Black/Blue = Alternating text color for exons Red = Residue overlap splice site Green = Synonymous SNP Yellow = Non-synonymous SNP

Next Step n Annotate the exact boundaries of each exon in our D. erecta sequence based on sequence homology to D. melanogaster eyeless gene n Use exon-by-exon BLAST search with BLAST 2 Sequences (bl 2 seq)

Questions?

Walk- through example

Determining Exon Boundaries n Use bl 2 seq to determine exon boundaries of the putative ortholog in our D. erecta sequence n Go to www. ncbi. nlm. nih. gov/blast/ and select bl 2 seq n Copy D. erecta sequence and paste into the Sequence 1 box. Copy the first exon of DM eyeless and paste into the Sequence 2 box. n Change program to tblastx. Click “BLAST”

Determining Exon Boundaries n We find that the first exon corresponds to bases 1930719414 in our sequence n We can repeat the previous steps to locate the other exons in our sequence
- Slides: 24