Web Databases for Drosophila An introduction to web

Web Databases for Drosophila An introduction to web tools, databases and NCBI BLAST Wilson Leung 08/2019

Agenda • GEP annotation project overview • Web databases for Drosophila annotation – UCSC Genome Browser – NCBI / BLAST – Fly. Base – Gene Record Finder
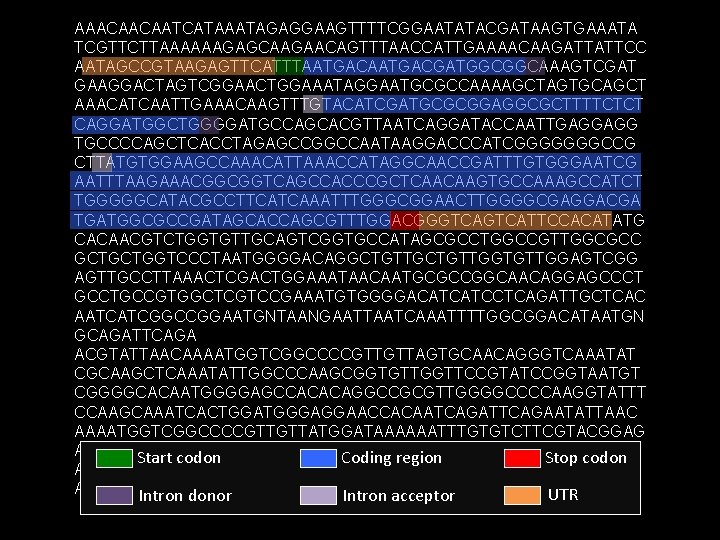
AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATA TCGTTCTTAAAAAAGAGCAAGAACAGTTTAACCATTGAAAACAAGATTATTCC AATAGCCGTAAGAGTTCATTTAATGACGATGGCGGCAAAGTCGAT GAAGGACTAGTCGGAACTGGAAATAGGAATGCGCCAAAAGCTAGTGCAGCT AAACATCAATTGAAACAAGTTTGTACATCGATGCGCGGAGGCGCTTTTCTCT CAGGATGGCTGGGGATGCCAGCACGTTAATCAGGATACCAATTGAGGAGG TGCCCCAGCTCACCTAGAGCCGGCCAATAAGGACCCATCGGGGGGGCCG CTTATGTGGAAGCCAAACATTAAACCATAGGCAACCGATTTGTGGGAATCG AATTTAAGAAACGGCGGTCAGCCACCCGCTCAACAAGTGCCAAAGCCATCT TGGGGGCATACGCCTTCATCAAATTTGGGCGGAACTTGGGGCGAGGACGA TGATGGCGCCGATAGCACCAGCGTTTGGACGGGTCATTCCACATATG CACAACGTCTGGTGTTGCAGTCGGTGCCATAGCGCCTGGCCGTTGGCGCC GCTGCTGGTCCCTAATGGGGACAGGCTGTTGGTGTTGGAGTCGG AGTTGCCTTAAACTCGACTGGAAATAACAATGCGCCGGCAACAGGAGCCCT GCCTGCCGTGGCTCGTCCGAAATGTGGGGACATCATCCTCAGATTGCTCAC AATCATCGGCCGGAATGNTAANGAATTAATCAAATTTTGGCGGACATAATGN GCAGATTCAGA ACGTATTAACAAAATGGTCGGCCCCGTTGTTAGTGCAACAGGGTCAAATAT CGCAAGCTCAAATATTGGCCCAAGCGGTGTTGGTTCCGTATCCGGTAATGT CGGGGCACAATGGGGAGCCACACAGGCCGCGTTGGGGCCCCAAGGTATTT CCAAGCAAATCACTGGATGGGAGGAACCACAATCAGATTCAGAATATTAAC AAAATGGTCGGCCCCGTTGTTATGGATAAAAAATTTGTGTCTTCGTACGGAG ATTATGTTGTTAATCAATTTTATTAAGATATTTAAATATGTGTACCTTTC Start codon Stop codon Coding region ACGAGAAATTTGCTTACCTTTTCGACACACTTATACAGGTAATA ATTACCTTTTGAGCAATTCGATTTTCATAAAATATACCTAAATCGCATCGTC UTR Intron donor Intron acceptor

Annotation – adding labels to a sequence • • • Genes: Novel or known genes, pseudogenes Regulatory Elements: Promoters, enhancers, silencers Non-coding RNA: t. RNAs, mi. RNAs, sno. RNAs Repeats: Transposable elements, simple repeats Structural: Origins of replication Experimental Results: – DNase I Hypersensitive sites – Ch. IP-chip and Ch. IP-Seq datasets (e. g. , mod. ENCODE)
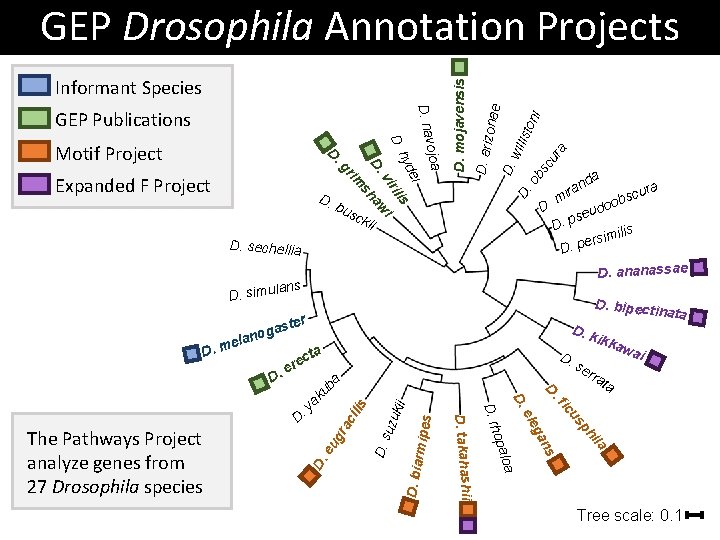
Informant Species D. ei hyd ilis sck ii D i bu da n ira . m aw D. sh Expanded F Project vir im gr D. Motif Project ojoa D. nav GEP Publications D. mojavensis D. ari zonae D. wil D. list oni ob sc ur a GEP Drosophila Annotation Projects D. sechellia se D. p imilis s r e D. p D. ananassae lans D. simu ter gas D. k ta rmipe s kii D. bia ila D. s uzu ilis ata ph ra c wai us s an D. eu g ikka fic D. y ata se rr D. leg e D. loa opa D. takahashii The Pathways Project analyze genes from 27 Drosophila species D. rh ak D. c ere a D D. bipectin ub o lan. me a cur obs o d u Tree scale: 0. 1
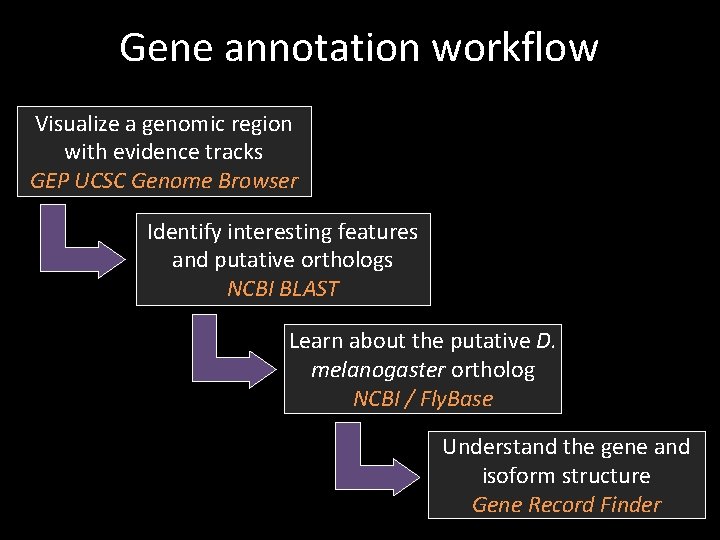
Gene annotation workflow Visualize a genomic region with evidence tracks GEP UCSC Genome Browser Identify interesting features and putative orthologs NCBI BLAST Learn about the putative D. melanogaster ortholog NCBI / Fly. Base Understand the gene and isoform structure Gene Record Finder

UCSC Genome Browser • Provide graphical view of genomic regions – Sequence conservation – Gene and splice site predictions – RNA-Seq and splice junction predictions • BLAT – BLAST-Like Alignment Tool – Map protein or nucleotide sequences against an assembly – Faster but less sensitive than BLAST • Table Browser – Access data used to create the graphical browser
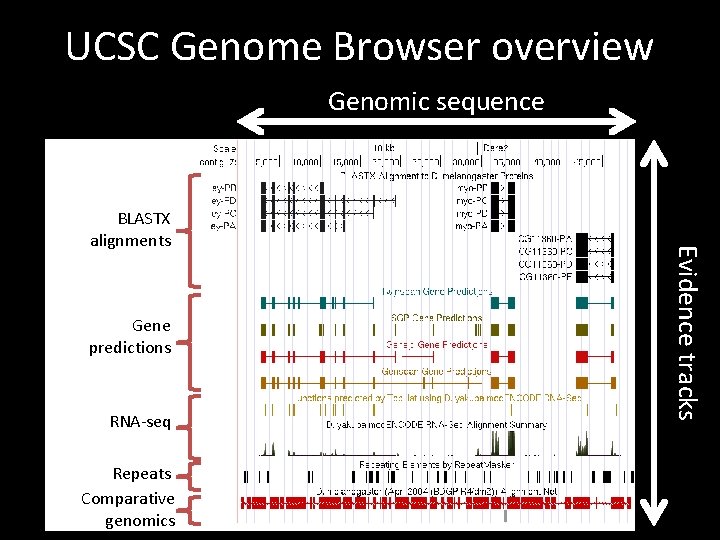
UCSC Genome Browser overview Genomic sequence Gene predictions RNA-seq Repeats Comparative genomics Evidence tracks BLASTX alignments

Control how evidence tracks are displayed on the Genome Browser • Most evidence tracks have five display modes: – Hide: track is hidden – Dense: all features (including overlapping features) are displayed on a single line – Squish: overlapping features are drawn on separate lines, features are half the height compared to full mode – Pack: overlapping features are drawn on separate lines, features are the same height as full mode – Full: Each feature is displayed on its own line • Some annotation tracks (e. g. , Repeat. Masker) only have a subset of these display modes
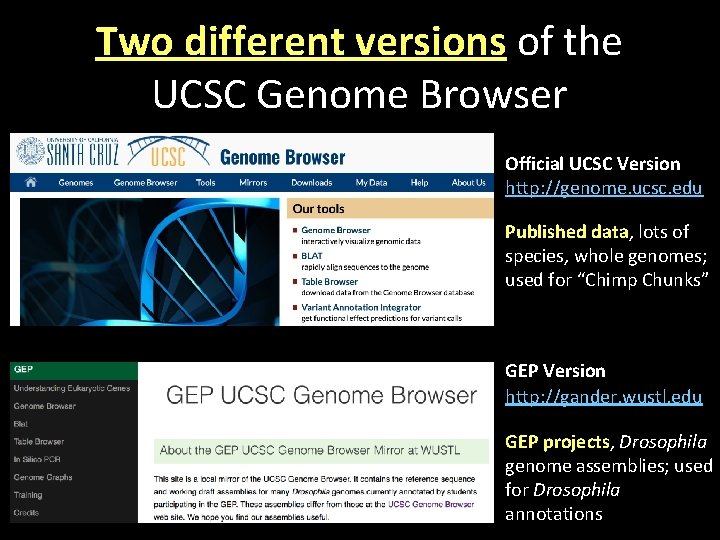
Two different versions of the UCSC Genome Browser Official UCSC Version http: //genome. ucsc. edu Published data, lots of species, whole genomes; used for “Chimp Chunks” GEP Version http: //gander. wustl. edu GEP projects, Drosophila genome assemblies; used for Drosophila annotations

Additional resources for the UCSC Genome Browser • Training section on the UCSC web site – http: //genome. ucsc. edu/training/index. html – Video tutorials – User guides – Mailing lists • Biostars – https: //www. biostars. org/t/ucsc/ – Questions with the “ucsc” tag

UCSC GENOME BROWSER DEMO

Use BLAST to detect sequence similarity • BLAST = Basic Local Alignment Search Tool • Why is BLAST popular? – Provide statistical significance for each match – Good balance of sensitivity and speed • Find local regions of similarity irrespective of where they are in the sequence

Common types of BLAST programs • Except for BLASTN, all alignments are based on comparisons of protein sequences • Decide which BLAST program to use based on the type of query and subject sequences: Program BLASTN BLASTP BLASTX Query Nucleotide Protein Nucleotide → Protein Database (Subject) Nucleotide Protein TBLASTN Protein Nucleotide → Protein TBLASTX Nucleotide → Protein

Common BLAST programs use cases • • • BLASTN: Map m. RNAs against genomic assemblies BLASTP: Search for proteins similar to predicted genes BLASTX: Map proteins / CDS against genomic sequence TBLASTN: Map proteins against genomic assemblies TBLASTX: Identify genes in unannotated sequences • See the “Guide to BLAST home and search pages” document for details: • ftp: //ftp. ncbi. nlm. nih. gov/pub/factsheets/How. To_BLASTGuide. pdf

NCBI BLAST nucleotide databases • Gen. Bank Non-Redundant Nucleotide Database (nr/nt) – Most comprehensive but some entries are low quality – Exclude sequences from whole genome assemblies • Ref. Seq RNA Database – m. RNA and non-coding RNA entries from the NCBI Reference Sequence Project – Include real and computationally predicted sequences • Transcriptome Shotgun Assembly (TSA) Database – Transcripts assembled from expressed sequence tags (ESTs) and RNA-Seq reads

NCBI BLAST protein databases • Gen. Bank Non-Redundant Protein Database (nr) – Most comprehensive but some entries are low quality – Include sequences from Ref. Seq and Uni. Prot. KB • Ref. Seq Protein Database – Sequences from the NCBI Reference Sequence Project – Higher quality than the nr database – Include real and computationally predicted sequences • Uni. Prot. KB / Swiss-Protein Database – Manually curated proteins from literature – Real proteins with known functions – Much smaller database than either Ref. Seq or nr

Where can I run BLAST? • NCBI BLAST web service – https: //blast. ncbi. nlm. nih. gov/Blast. cgi • EBI BLAST web service – https: //www. ebi. ac. uk/Tools/sss/ • Fly. Base BLAST (Drosophila and other insects) – http: //flybase. org/blast/

NCBI BLAST DEMO
National Center for Biotechnology Information (NCBI) https: //www. ncbi. nlm. nih. gov

Key features of NCBI • Strengths – Most comprehensive among publicly available databases – Pub. Med for literature searches – Comprehensive BLAST web service • Weaknesses – Web site is large and complex – Quality of Gen. Bank records may vary • Use cases – Perform BLAST searches against Refseq, nr/nt databases – Use BLAST (bl 2 seq) to align two or more sequences

Fly. Base — Database for the Drosophila research community http: //flybase. org/

Key Features of Fly. Base • Lots of ancillary data for each gene in Drosophila • Curation of literature for each gene • Reference Drosophila annotations for all the other databases (including NCBI) • Fast release cycle (6 -8 releases per year) • Use cases: – BLAST searches against Drosophila genome assemblies – Genome browsers (GBrowse/JBrowse) – RNA-Seq expression profile and similarity searches

Web databases and tools • Many genome databases available – Be aware of different annotation releases – Use Fly. Base as the canonical reference • Web databases are being updated constantly – Update GEP materials before semester starts – Discrepancies in exercise screenshots – Minor changes in search results – Let us know about errors or revisions

FLYBASE DEMO
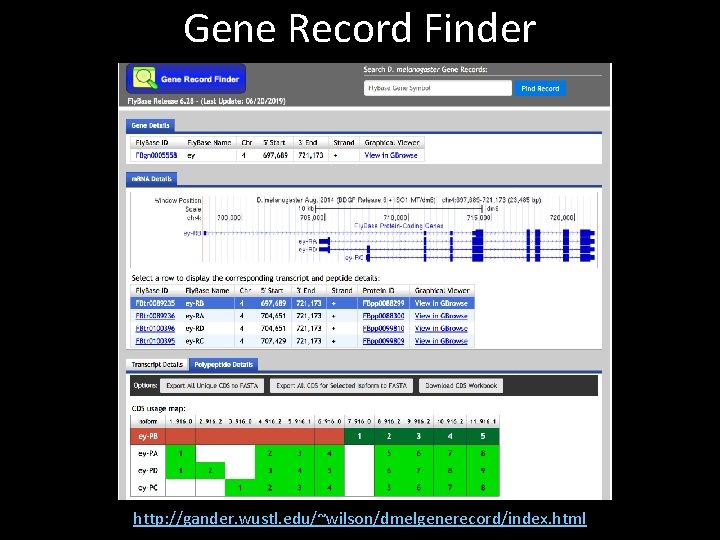
Gene Record Finder http: //gander. wustl. edu/~wilson/dmelgenerecord/index. html

Key features of the Gene Record Finder • List of unique coding and non-coding exons for each gene in D. melanogaster • CDS and exon usage maps for each isoform • Optimized for exon-by-exon annotation strategy • Slower update release cycle than Fly. Base – Database is updated every semester • Use cases: – Get amino acid sequences and nucleotide sequences of each exon for BLAST 2 Sequences (bl 2 seq) searches

GENE RECORD FINDER DEMO

Summary • GEP annotation project seeks to generate high quality manually curated gene models for multiple Drosophila species • Use BLAST to characterize a genomic sequence • Use web databases to gather information on a gene – UCSC Genome Browser – NCBI – Fly. Base – Gene Record Finder

Questions? https: //www. flickr. com/photos/jac_opo/240254763/sizes/l/

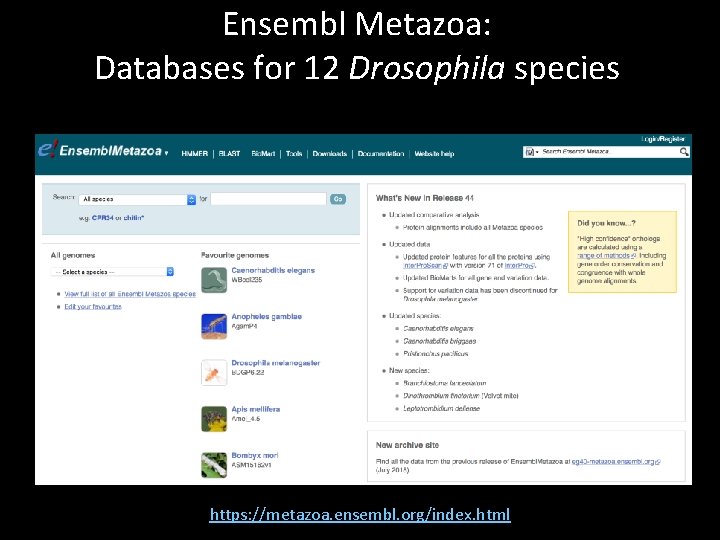
Ensembl Metazoa: Databases for 12 Drosophila species https: //metazoa. ensembl. org/index. html

Key features of Ensembl Metazoa • Lots of ancillary data for each gene • Detailed information on each gene available at the transcript, peptide, and exon level • Not always up-to-date – Annotations are from Fly. Base Release 6. 22 • Data for 12 Drosophila species available • Use cases: – Get amino acid sequences and nucleotide sequences of each exon for bl 2 seq searches – Perform species-specific BLAST searches
- Slides: 33