Volumetric Intersubject Registration John Ashburner Wellcome Department of

Volumetric Intersubject Registration John Ashburner Wellcome Department of Imaging Neuroscience, 12 Queen Square, London, UK.

Intersubject registration for f. MRI * Inter-subject averaging * Increase sensitivity with more subjects * Fixed-effects analysis * Extrapolate findings to the population as a whole * Mixed-effects analysis * Standard coordinate system * e. g. , Talairach & Tournoux space
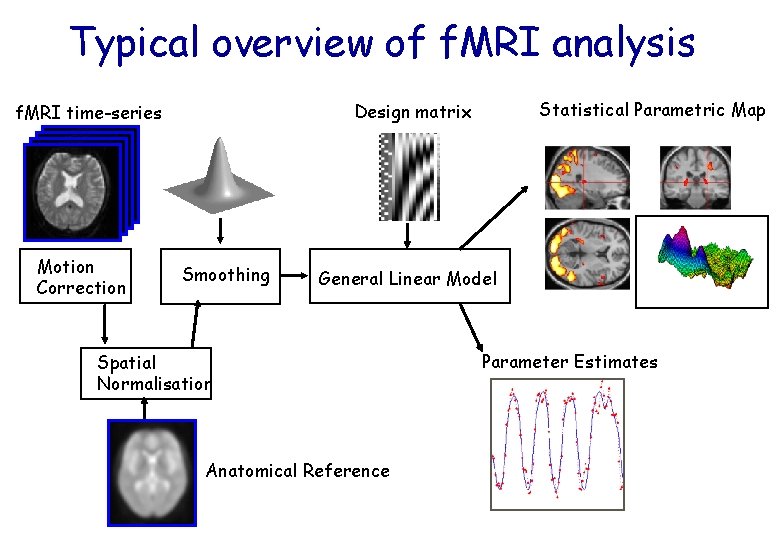
Typical overview of f. MRI analysis Motion Correction Statistical Parametric Map Design matrix f. MRI time-series Smoothing General Linear Model Spatial Normalisation Anatomical Reference Parameter Estimates

Overview * Part I: General Inter-subject registration * Spatial transformations * Affine * Global nonlinear * Local nonlinear * Objective functions for registration * Likelihood Models * Mean squared difference * Information Theoretic measures * Prior Models * Part II: The Segmentation Method in SPM 5

Image Registration z. Registration - i. e. Optimise the parameters that describe a spatial transformation between the source and reference (template) images z. Transformation - i. e. Re-sample according to the determined transformation parameters
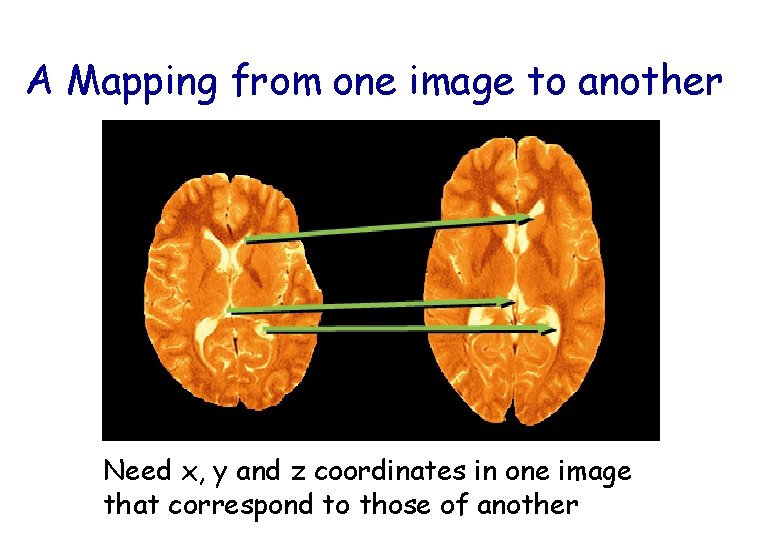
A Mapping from one image to another Need x, y and z coordinates in one image that correspond to those of another

Affine Transforms * Rigid-body transformations are a subset * Parallel lines remain parallel * Operations can be represented by: x’ = m 11 x + m 12 y + m 13 z + m 14 y’ = m 21 x + m 22 y + m 23 z + m 24 z’ = m 31 x + m 32 y + m 33 z + m 34 * Or as matrices: Y=Mx
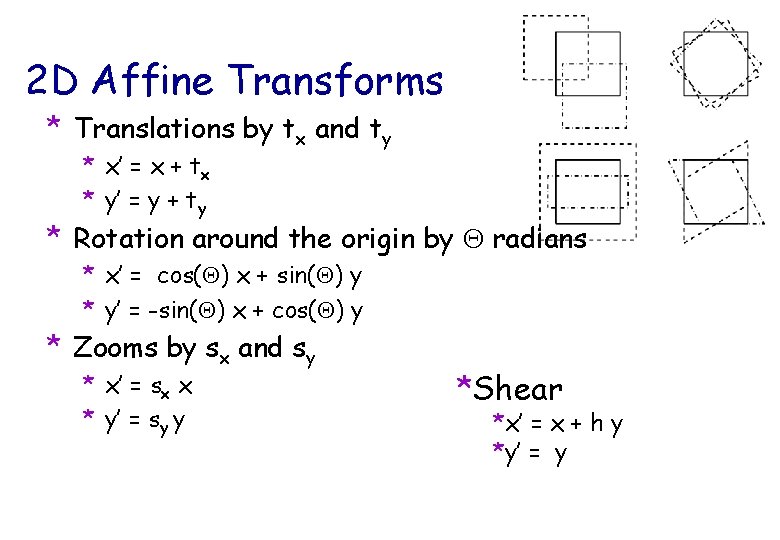
2 D Affine Transforms * Translations by tx and ty * x’ = x + tx * y’ = y + ty * Rotation around the origin by radians * x’ = cos( ) x + sin( ) y * y’ = -sin( ) x + cos( ) y * Zooms by sx and sy * x’ = sx x * y’ = sy y *Shear *x’ = x + h y *y’ = y

2 D Affine Transforms * Translations by tx and ty * x’ = 1 x + 0 y + tx * y’ = 0 x + 1 y + ty * Rotation around the origin by radians * x’ = cos( ) x + sin( ) y + 0 * y’ = -sin( ) x + cos( ) y + 0 * Zooms by sx and sy: * x’ = sx x + 0 y + 0 * y’ = 0 x + sy y + 0 *Shear *x’ = 1 x + h y + 0 *y’ = 0 x + 1 y + 0

Polynomial Basis Functions As used by Roger Woods’ AIR Software

Cosine Transform Basis Functions As used by SPM software
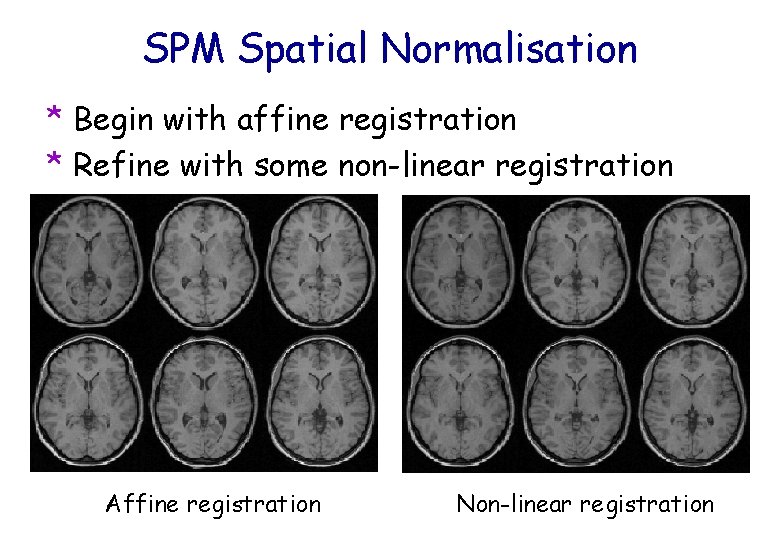
SPM Spatial Normalisation * Begin with affine registration * Refine with some non-linear registration Affine registration Non-linear registration

Accuracy of Automated Volumetric Inter-subject Registration Hellier et al. Inter subject registration of functional and anatomical data using SPM. MICCAI'02 LNCS 2489 (2002) Hellier et al. Retrospective evaluation of inter-subject brain registration. MIUA (2001)

Local Basis Functions * More detailed deformations use lots of basis functions with local support. * Local support means that the basis functions are mostly all zero * Faster computations

Simple addition of displacements Notice that there is no longer a one-to-one mapping

Generating large one-to-one deformations Y 1 Y 2 = Y 1 Y 3 = Y 1 Y 2 Y 4 = Y 1 Y 3 Y 5 = Y 1 Y 4 Y 6 = Y 1 Y 5 Y 7 = Y 1 Y 6 Y 8 = Y 1 Y 7 The principle behind the one-to-one mappings of viscous fluid registration

Faster to repeatedly square the deformation Y 1 Y 4 = Y 2 Y 2 = Y 1 Y 8 = Y 4 Note that this is analogous to computing a matrix exponential (c. f. Lie Groups and exponential mappings)
Y 16 = Y 8

One-to-One Mappings * One-to-one mappings break down beyond a certain scale * The concept of a single “best” mapping may be meaningless at higher resolution Pictures taken from http: //www. messybeast. com/freak-face. htm

Optimisation * Optimisation involves finding some “best” parameters according to an “objective function”, which is either minimised or maximised * The “objective function” is often related to a probability based on some model Objective function Most probable solution (global optimum) Local optimum Value of parameter

Bayes Rule * Most registration procedures maximise a joint probability of the deformation (warp) and the images (data). * P(Warp, Data) = P(Warp | Data) x P(Data) = P(Data | Warp) x P(Warp) * In practice, this can be by minimising * -log P(Warp, Data) = -log P(Data | Warp) -log P(Warp) Likelihood Prior
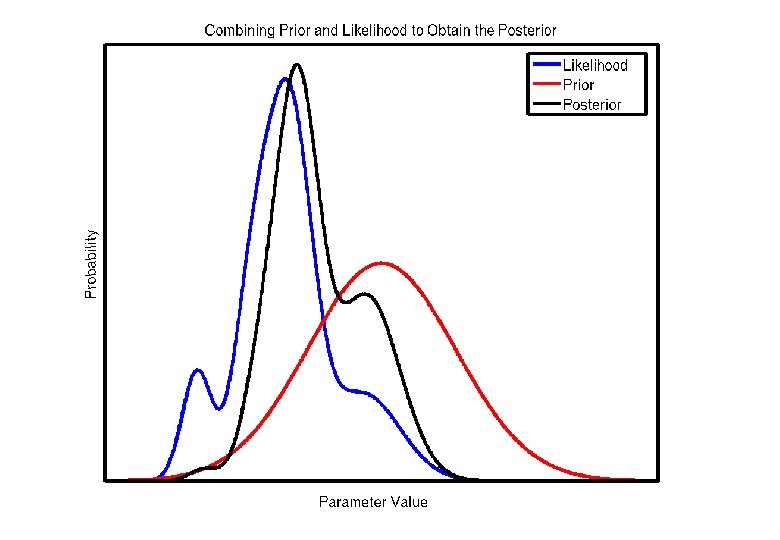

Mean Squared Difference Objective Function * Assumes one image is a warped version of the other with Gaussian noise added… * P(fi|t) = (2 ps 2)-1/2 exp(-(fi-gi(t))2/(2 s 2)) so * -log P(fi |t) = (fi-gi(t))2 /(2 s 2) + 1/2 log(2 ps 2) * Assumes that voxels are independent. . . * P(f 1, f 2, …, f. N, . . . ) = P(f 1) P(f 2) … P(f. N) so * -log P(f 1, f 2, …, f. N) = ((f 1 -g 1(t))2 + (f 2 -g 2(t))2 +…+ (f. N-g. N(t))2)/(2 s 2) + 1/2 N log(2 ps 2)
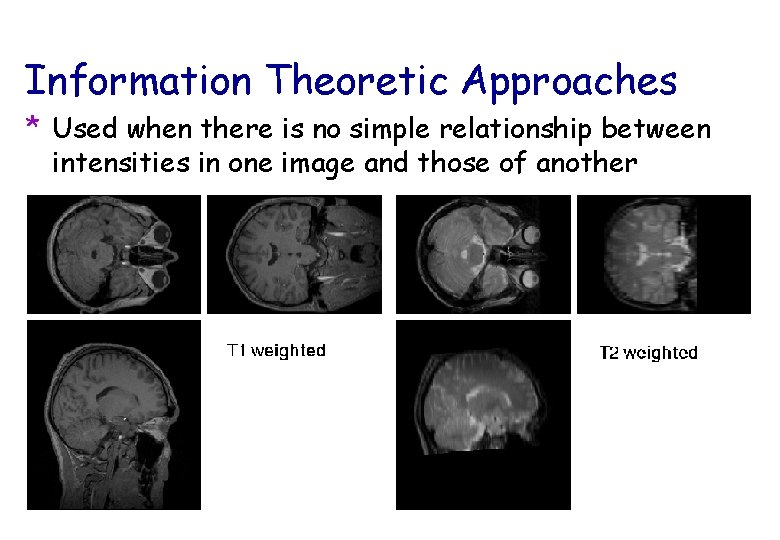
Information Theoretic Approaches * Used when there is no simple relationship between intensities in one image and those of another
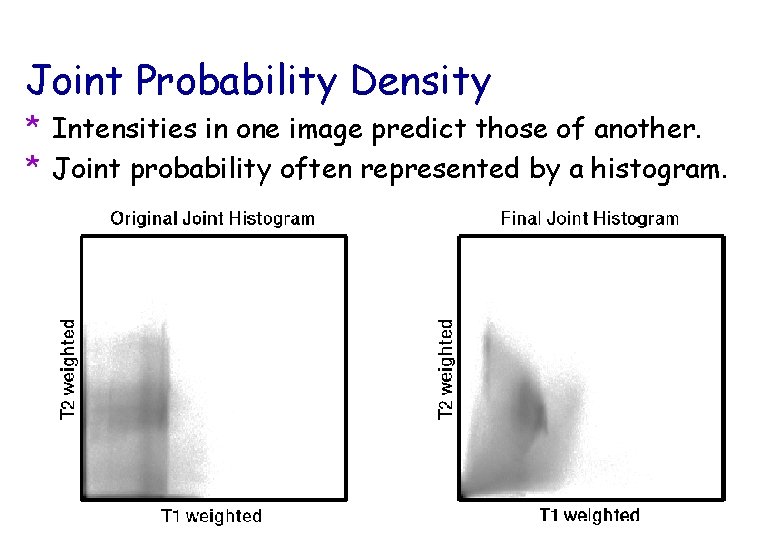
Joint Probability Density * Intensities in one image predict those of another. * Joint probability often represented by a histogram.
![Mutual Information * MI= ab P(a, b) log 2 [P(a, b)/( P(a) P(b) )] Mutual Information * MI= ab P(a, b) log 2 [P(a, b)/( P(a) P(b) )]](http://slidetodoc.com/presentation_image/dcf854f7b20f771fa680d6cb7b736607/image-26.jpg)
Mutual Information * MI= ab P(a, b) log 2 [P(a, b)/( P(a) P(b) )] * Related to entropy: MI = -H(a, b) + H(a) + H(b) * H(a) = - a P(a) log P(a) da * H(a, b) = - a P(a, b) log P(a, b) da
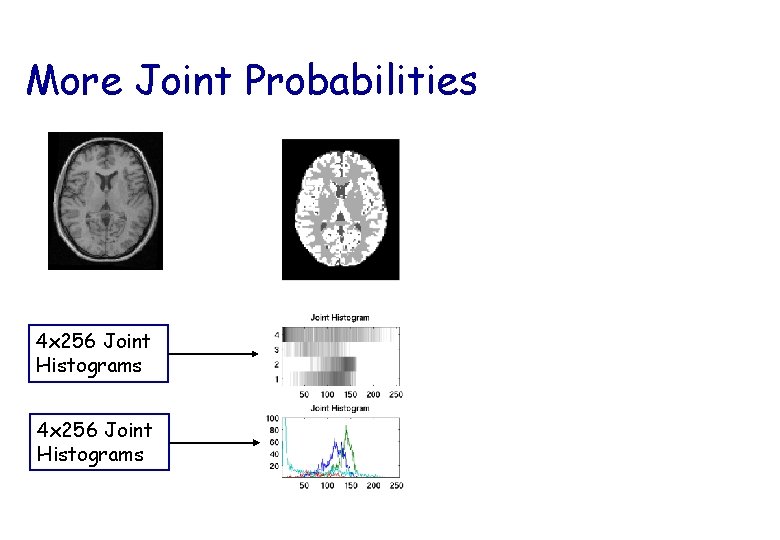
More Joint Probabilities 4 x 256 Joint Histograms
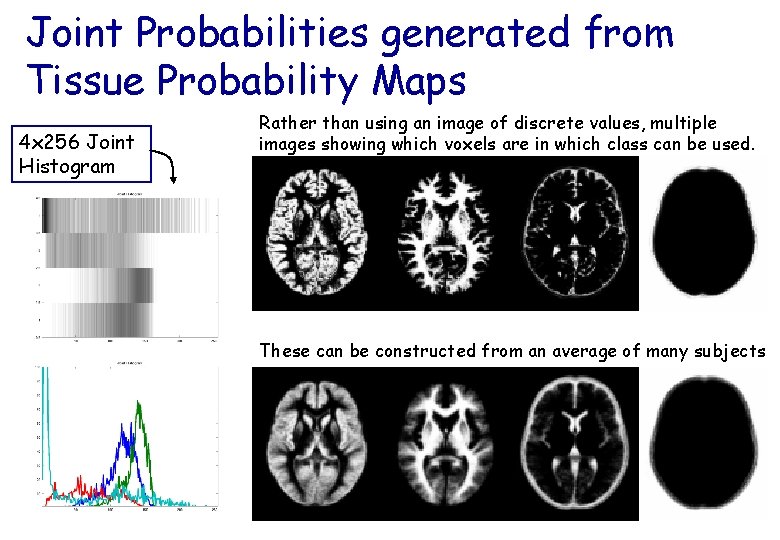
Joint Probabilities generated from Tissue Probability Maps 4 x 256 Joint Histogram Rather than using an image of discrete values, multiple images showing which voxels are in which class can be used. These can be constructed from an average of many subjects

Priors enforce “smooth” deformations * Membrane Energy * Bending Energy * Linear Elastic Energy

Priors enforce “smooth” deformations * The form of prior determines how the deformations behave in regions with no matching information

Overview * Part I: General Inter-subject registration * Part II: The Segmentation Method in SPM 5 * * Modelling intensities by a Mixture of Gaussians Bias correction Tissue Probability Maps to assist the segmentation Warping the tissue probability maps to match the image
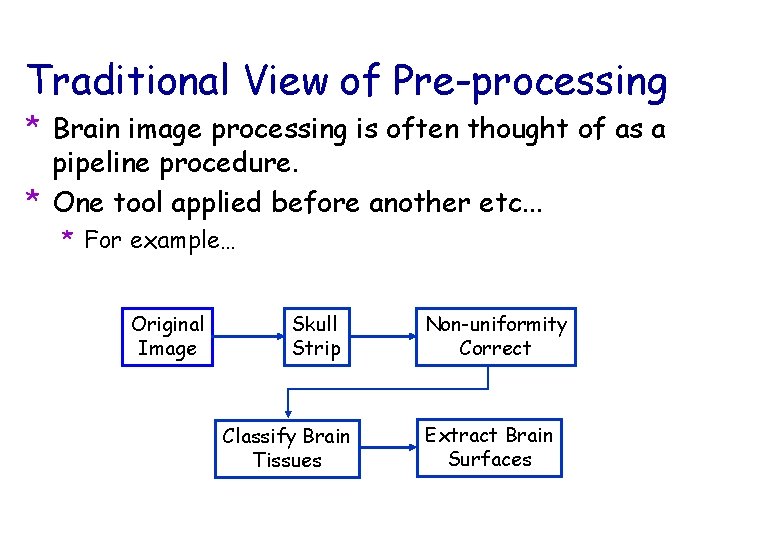
Traditional View of Pre-processing * Brain image processing is often thought of as a pipeline procedure. * One tool applied before another etc. . . * For example… Original Image Skull Strip Classify Brain Tissues Non-uniformity Correct Extract Brain Surfaces
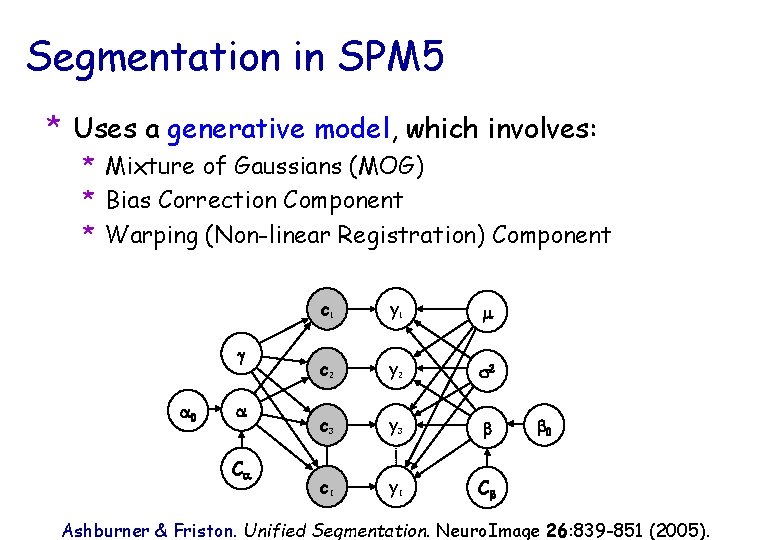
Segmentation in SPM 5 * Uses a generative model, which involves: * Mixture of Gaussians (MOG) * Bias Correction Component * Warping (Non-linear Registration) Component g a 0 a Ca c 1 y 1 m c 2 y 2 s 2 c 3 y 3 b c. I y. I Cb b 0 Ashburner & Friston. Unified Segmentation. Neuro. Image 26: 839 -851 (2005).
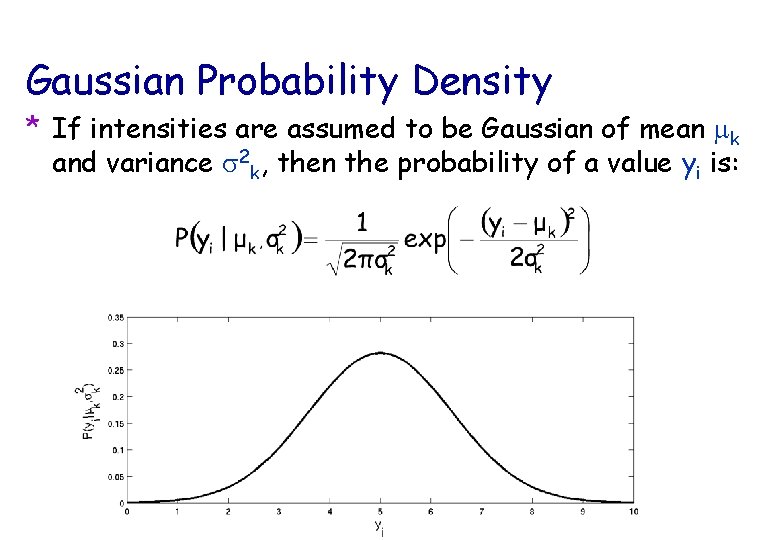
Gaussian Probability Density * If intensities are assumed to be Gaussian of mean mk and variance s 2 k, then the probability of a value yi is:
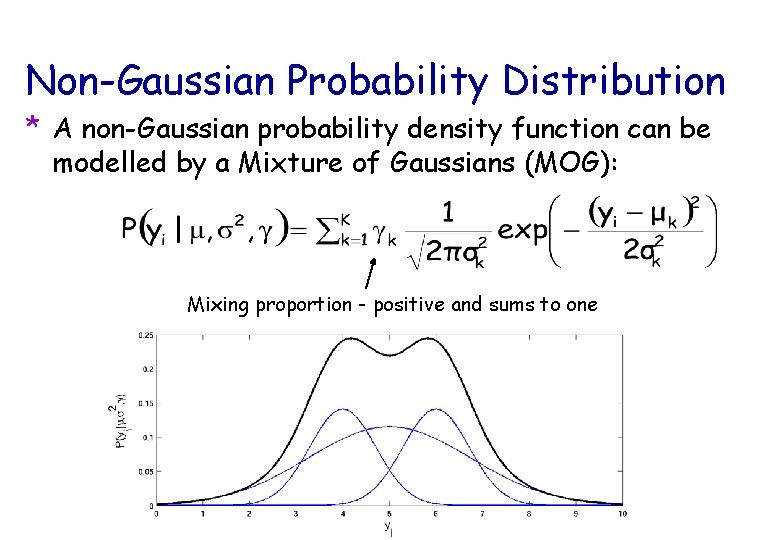
Non-Gaussian Probability Distribution * A non-Gaussian probability density function can be modelled by a Mixture of Gaussians (MOG): Mixing proportion - positive and sums to one
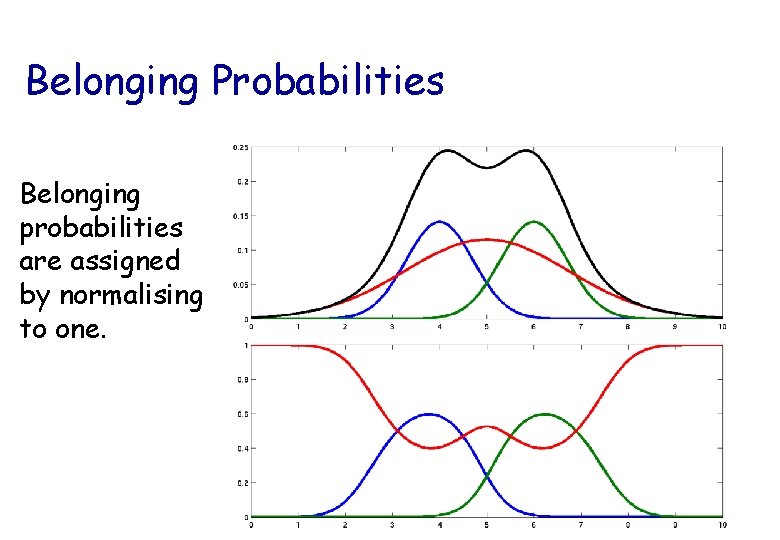
Belonging Probabilities Belonging probabilities are assigned by normalising to one.

Mixing Proportions * The mixing proportion gk represents the prior probability of a voxel being drawn from class k irrespective of its intensity. * So:
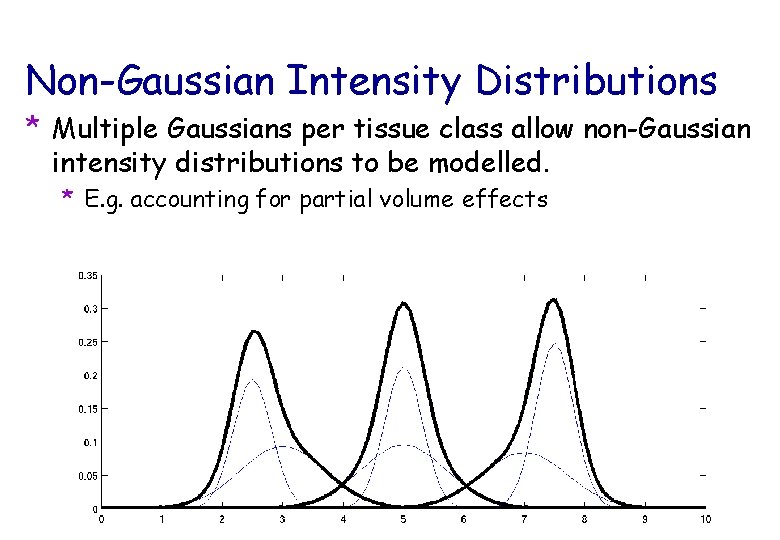
Non-Gaussian Intensity Distributions * Multiple Gaussians per tissue class allow non-Gaussian intensity distributions to be modelled. * E. g. accounting for partial volume effects

Probability of Whole Dataset * If the voxels are assumed to be independent, then the probability of the whole image is the product of the probabilities of each voxel: * A maximum-likelihood solution can be found by minimising the negative log-probability:

Modelling a Bias Field * A bias field is included, such that the required scaling at voxel i, parameterised by b, is ri(b). * Replace the means by mk/ri(b) * Replace the variances by (sk/ri(b))2

Modelling a Bias Field * After rearranging. . . y c 1 y 1 m g c 2 y 2 s 2 a 0 a c 3 y 3 b c. I y. I Cb Ca r(b) y r(b) b 0
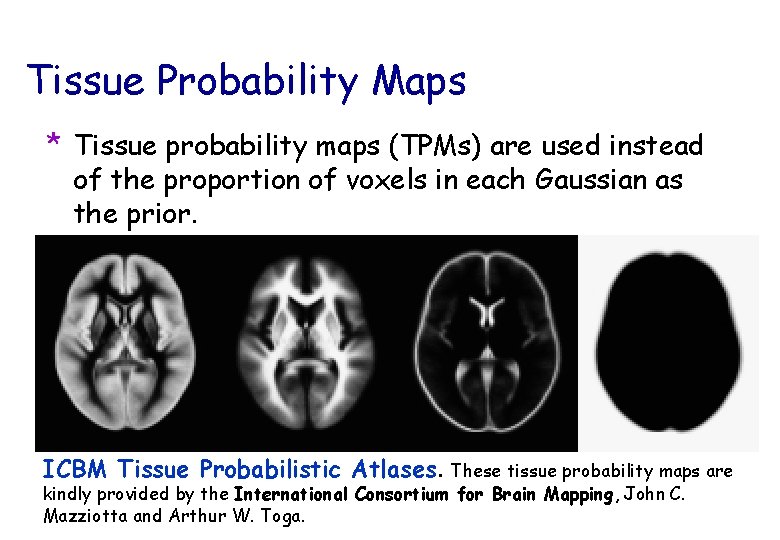
Tissue Probability Maps * Tissue probability maps (TPMs) are used instead of the proportion of voxels in each Gaussian as the prior. ICBM Tissue Probabilistic Atlases. These tissue probability maps are kindly provided by the International Consortium for Brain Mapping, John C. Mazziotta and Arthur W. Toga.
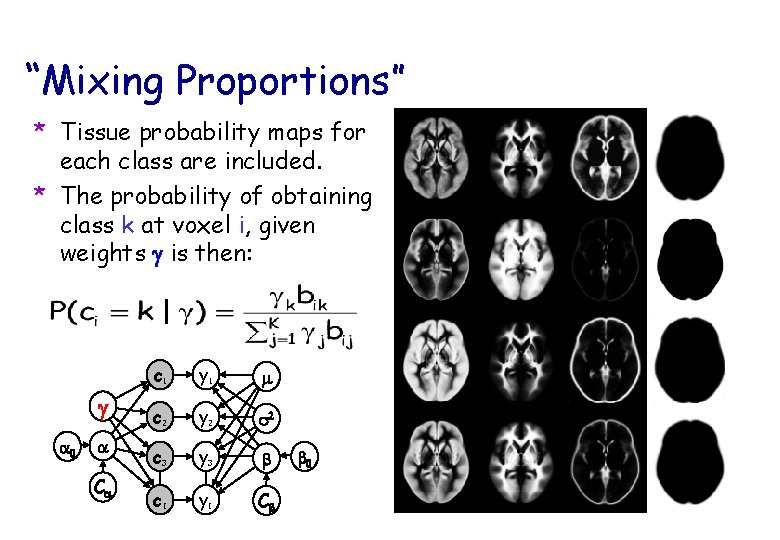
“Mixing Proportions” * Tissue probability maps for each class are included. * The probability of obtaining class k at voxel i, given weights g is then: g a 0 a Ca c 1 y 1 m c 2 y 2 s 2 c 3 y 3 b c. I y. I Cb b 0
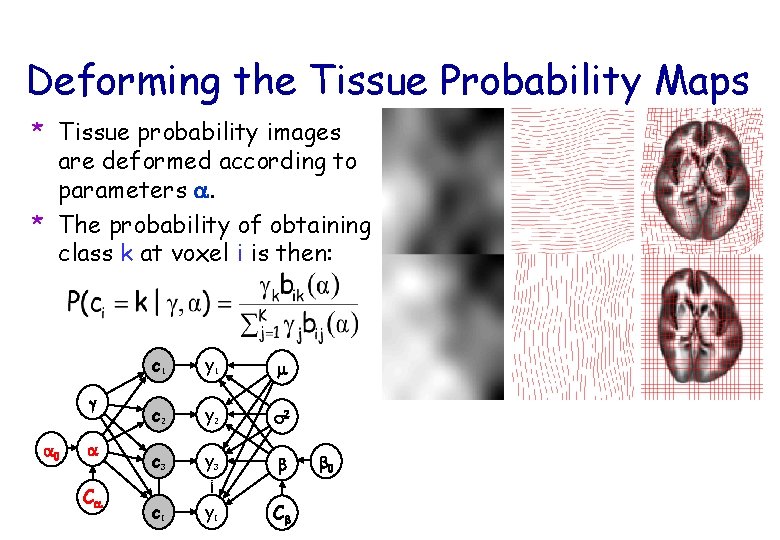
Deforming the Tissue Probability Maps * Tissue probability images are deformed according to parameters a. * The probability of obtaining class k at voxel i is then: g a 0 a Ca c 1 y 1 m c 2 y 2 s 2 c 3 y 3 b c. I y. I Cb b 0

The Extended Model * By combining the modified P(ci=k|q) and P(yi|ci=k, q), the overall objective function (E) becomes: The Objective Function

Optimisation * The “best” parameters are those that minimise this objective function. * Optimisation involves finding them. * Begin with starting estimates, and repeatedly change them so that the objective function decreases each time.

Steepest Descent Start Optimum Alternate between optimising different groups of parameters

Schematic of optimisation Repeat until convergence… Hold g, m, s 2 and a constant, and minimise E w. r. t. b - Levenberg-Marquardt strategy, using d. E/db and d 2 E/db 2 Hold g, m, s 2 and b constant, and minimise E w. r. t. a - Levenberg-Marquardt strategy, using d. E/da and d 2 E/da 2 Hold a and b constant, and minimise E w. r. t. g, m and s 2 -Use an Expectation Maximisation (EM) strategy. end

Levenberg-Marquardt Optimisation * LM optimisation is used for nonlinear registration (a) and bias correction (b). * Requires first and second derivatives of the objective function (E). * Parameters a and b are updated by * Increase l to improve stability (at expense of decreasing speed of convergence).

Expectation Maximisation is used to update m, s 2 and g * For iteration (n), alternate between: * E-step: Estimate belonging probabilities by: * M-step: Set q(n+1) to values that reduce:

Regularisation * Some bias fields and warps are more probable (a priori) than others. * Encoded using Bayes rule (for a maximum a posteriori solution): * Prior probability distributions modelled by a multivariate normal distribution. * * Mean vector ma and mb Covariance matrix Sa and Sb -log[P(a)] = (a-ma)TSa-1(a-ma) + const -log[P(b)] = (b-mb)TSb-1(b-mb) + const
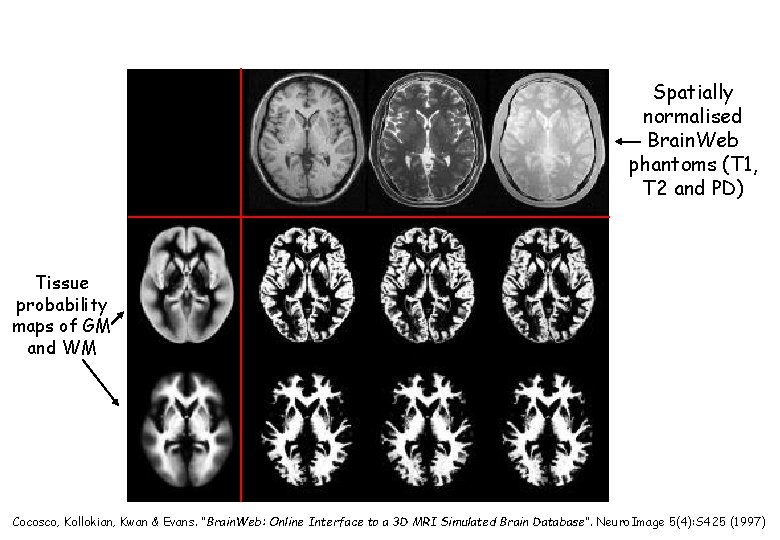
Spatially normalised Brain. Web phantoms (T 1, T 2 and PD) Tissue probability maps of GM and WM Cocosco, Kollokian, Kwan & Evans. “Brain. Web: Online Interface to a 3 D MRI Simulated Brain Database”. Neuro. Image 5(4): S 425 (1997)

Summary * Part I: General Inter-subject registration * Spatial transformations * Affine * Global nonlinear * Local nonlinear * Objective functions for registration * Likelihood Models * Mean squared difference * Information Theoretic measures * Prior Models * Part II: The Segmentation Method in SPM 5 * * Modelling intensities by a Mixture of Gaussians Bias correction Tissue Probability Maps to assist the segmentation Warping the tissue probability maps to match the image

References * Friston et al. Spatial registration and normalisation of images. Human Brain Mapping 3: 165 -189 (1995). * Collignon et al. Automated multi-modality image registration based on information theory. IPMI’ 95 pp 263 -274 (1995). * Ashburner et al. Incorporating prior knowledge into image registration. Neuro. Image 6: 344 -352 (1997). * Ashburner & Friston. Nonlinear spatial normalisation using basis functions. Human Brain Mapping 7: 254 -266 (1999). * Thévenaz et al. Interpolation revisited. IEEE Trans. Med. Imaging 19: 739 -758 (2000). * Andersson et al. Modeling geometric deformations in EPI time series. Neuroimage 13: 903 -919 (2001). * Ashburner & Friston. Unified Segmentation. Neuro. Image in press (2005).



Spare slides
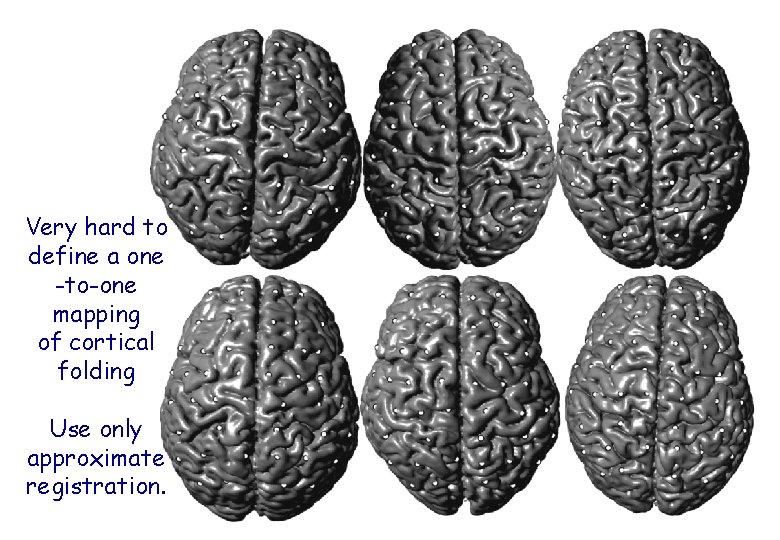
Very hard to define a one -to-one mapping of cortical folding Use only approximate registration.
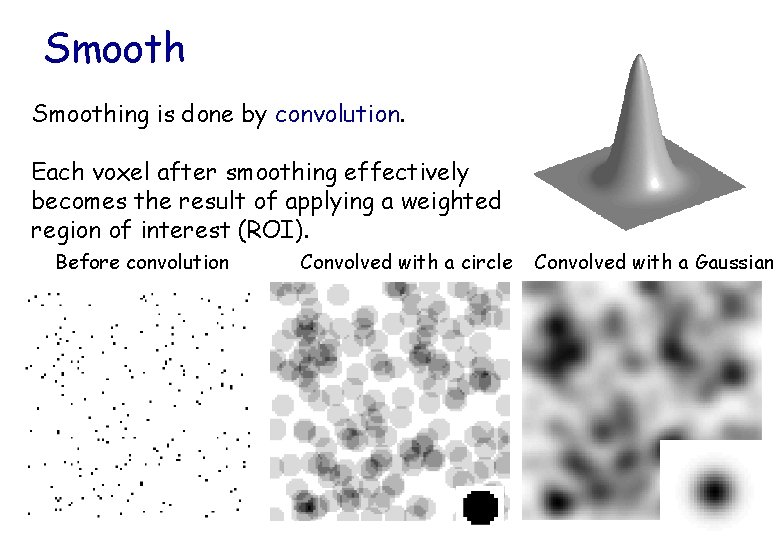
Smoothing is done by convolution. Each voxel after smoothing effectively becomes the result of applying a weighted region of interest (ROI). Before convolution Convolved with a circle Convolved with a Gaussian

Voxel-to-world Transforms * Affine transform associated with each image * Maps from voxels (x=1. . nx, y=1. . ny, z=1. . nz) to some world co -ordinate system. e. g. , * Scanner co-ordinates - images from DICOM toolbox * T&T/MNI coordinates - spatially normalised * Registering image B (source) to image A (target) will update B’s voxel-to-world mapping * Mapping from voxels in A to voxels in B is by * A-to-world using MA, then world-to-B using MB-1 * MB-1 MA
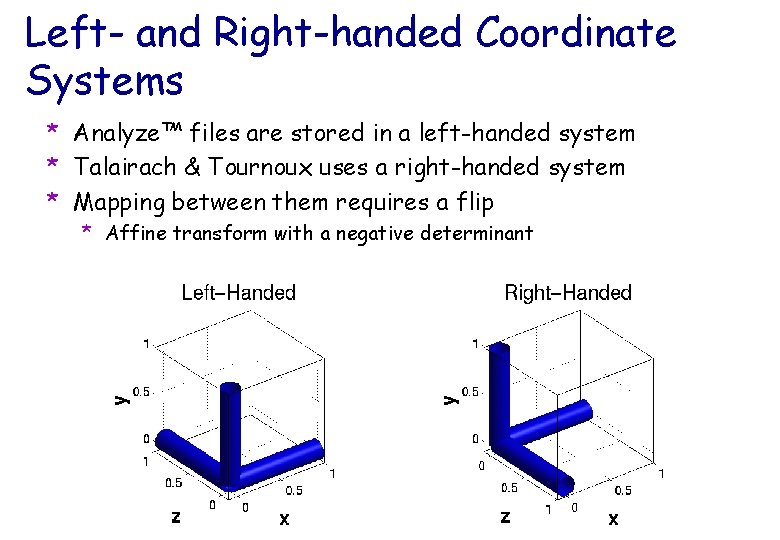
Left- and Right-handed Coordinate Systems * Analyze™ files are stored in a left-handed system * Talairach & Tournoux uses a right-handed system * Mapping between them requires a flip * Affine transform with a negative determinant

Transforming an image * Images are re-sampled. An example in 2 D: for y=1. . ny % loop over rows for x=1. . nx % loop over pixels in row x’= tx(x, y, a) % transform according to a y’= ty(x, y, a) if 1 x’ nx & 1 y’ ny then % voxel in range f (x, y) = f’(x’, y’) % assign re-sampled value end % voxel in range end % loop over pixels in row end % loop over rows * What happens if x’ and y’ are not integers?
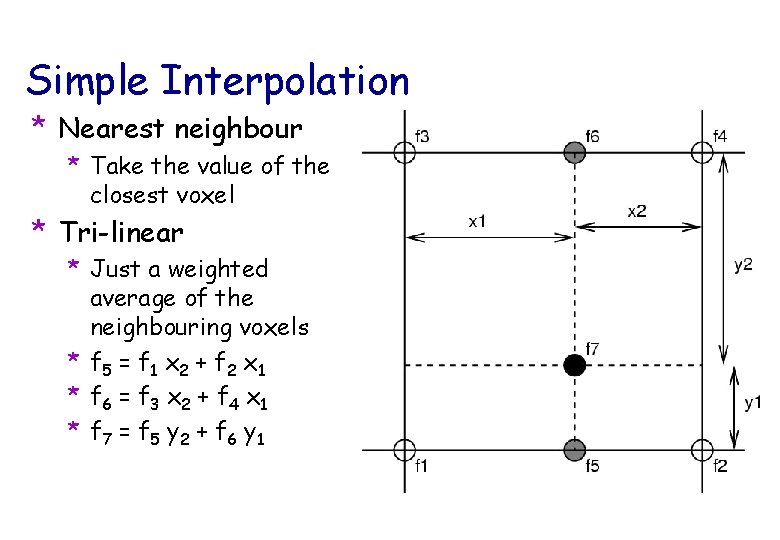
Simple Interpolation * Nearest neighbour * Take the value of the closest voxel * Tri-linear * Just a weighted average of the neighbouring voxels * f 5 = f 1 x 2 + f 2 x 1 * f 6 = f 3 x 2 + f 4 x 1 * f 7 = f 5 y 2 + f 6 y 1
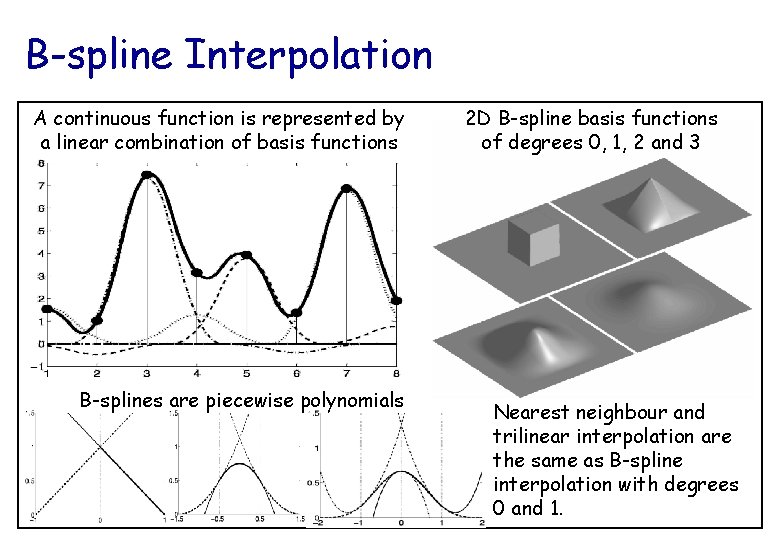
B-spline Interpolation A continuous function is represented by a linear combination of basis functions B-splines are piecewise polynomials 2 D B-spline basis functions of degrees 0, 1, 2 and 3 Nearest neighbour and trilinear interpolation are the same as B-spline interpolation with degrees 0 and 1.
Inverse
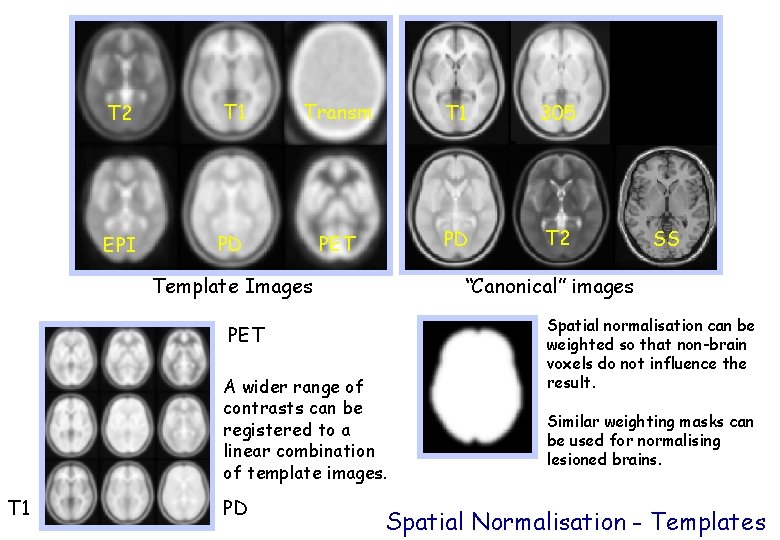
T 2 T 1 Transm T 1 305 EPI PD PET PD T 2 Template Images “Canonical” images PET A wider range of contrasts can be registered to a linear combination of template images. T 1 PD SS Spatial normalisation can be weighted so that non-brain voxels do not influence the result. Similar weighting masks can be used for normalising lesioned brains. Spatial Normalisation - Templates
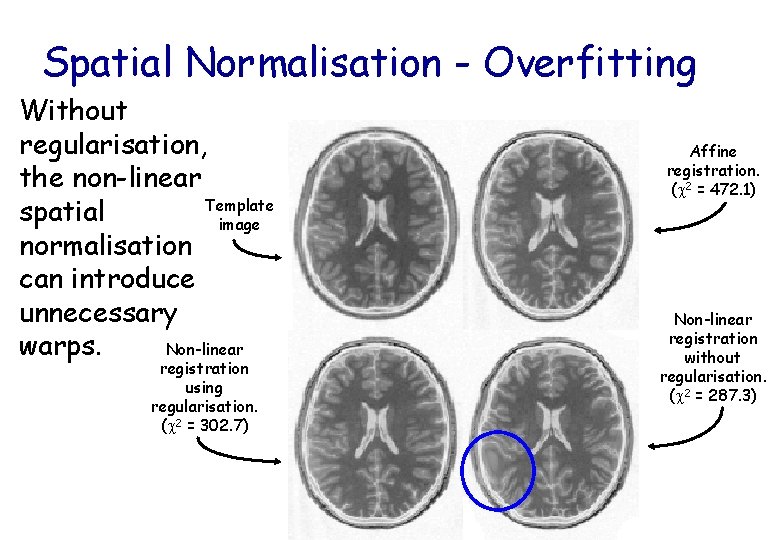
Spatial Normalisation - Overfitting Without regularisation, the non-linear Template spatial image normalisation can introduce unnecessary Non-linear warps. registration using regularisation. ( 2 = 302. 7) Affine registration. ( 2 = 472. 1) Non-linear registration without regularisation. ( 2 = 287. 3)

A Growing Trend * Larger and more complex models are being produced to explain brain imaging data. * Bigger and better computers * allow more powerful models to be used * More experience among software developers * Older and wiser * More engineers - rather than e. g. psychiatrists & biochemists * This presentation is about combining various preprocessing procedures for anatomical images into a single generative model.
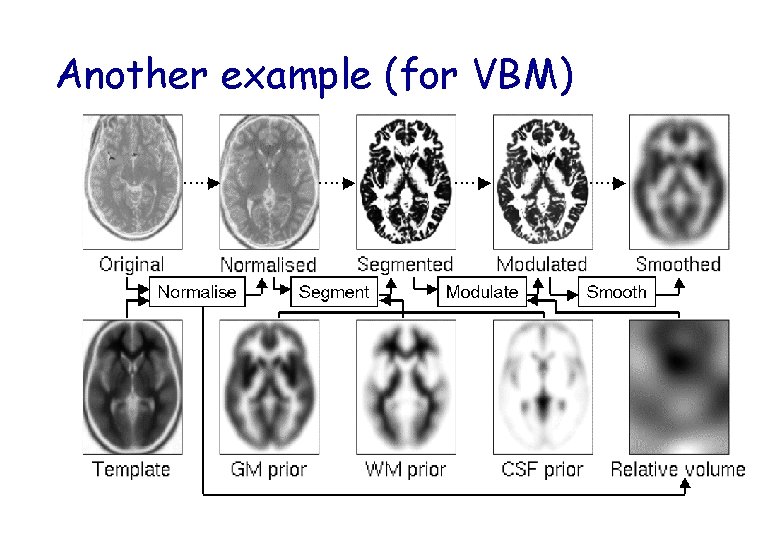
Another example (for VBM)
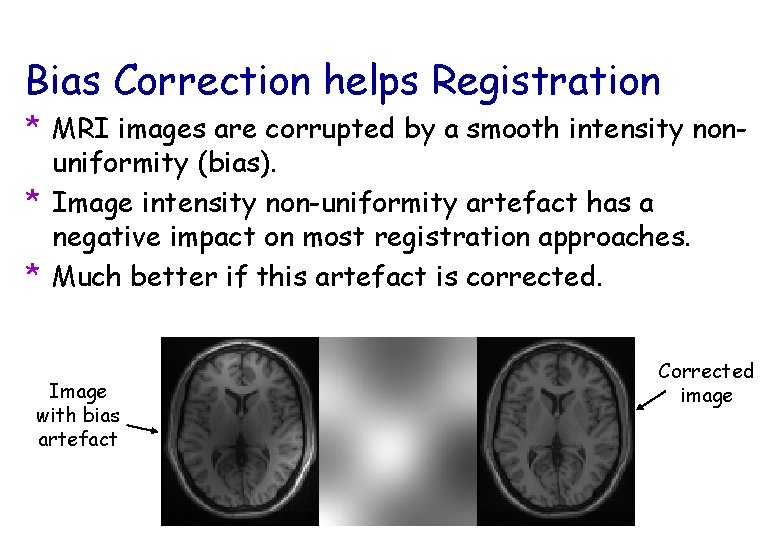
Bias Correction helps Registration * MRI images are corrupted by a smooth intensity nonuniformity (bias). * Image intensity non-uniformity artefact has a negative impact on most registration approaches. * Much better if this artefact is corrected. Image with bias artefact Corrected image
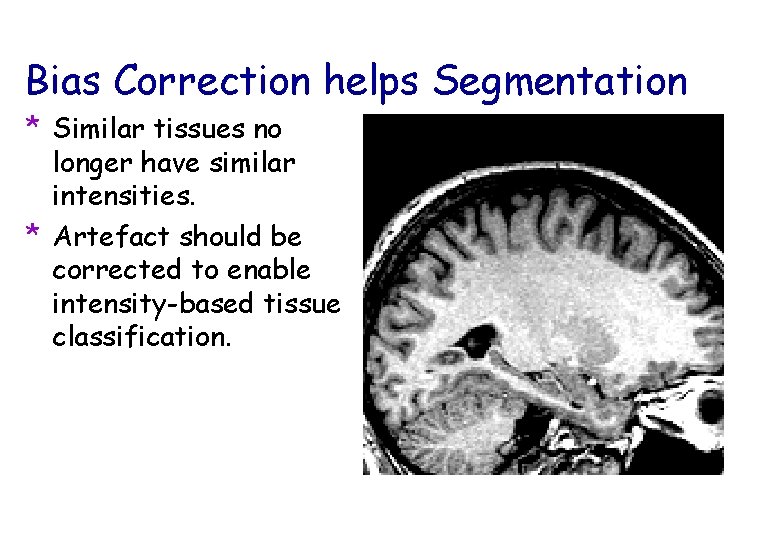
Bias Correction helps Segmentation * Similar tissues no longer have similar intensities. * Artefact should be corrected to enable intensity-based tissue classification.
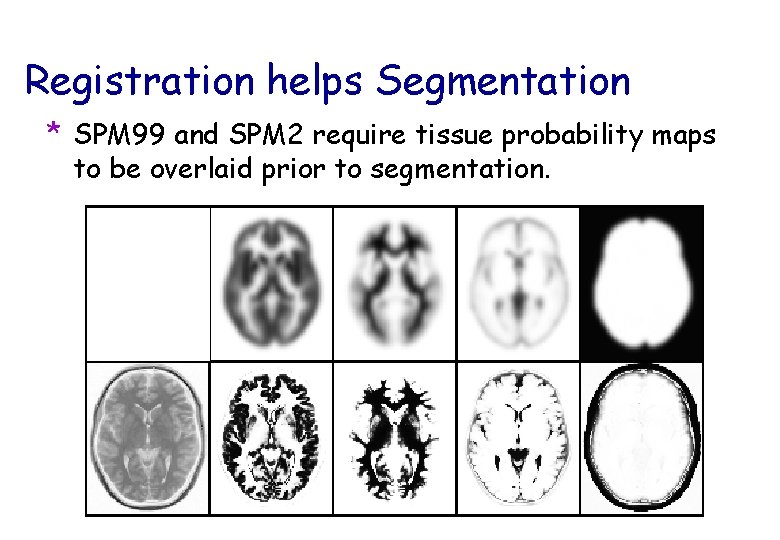
Registration helps Segmentation * SPM 99 and SPM 2 require tissue probability maps to be overlaid prior to segmentation.

Segmentation helps Bias Correction * Bias correction should not eliminate differences between tissue classes. * Can be done by * make all white matter about the same intensity * make all grey matter about the same intensity * etc * Currently fairly standard practice to combine bias correction and tissue classification
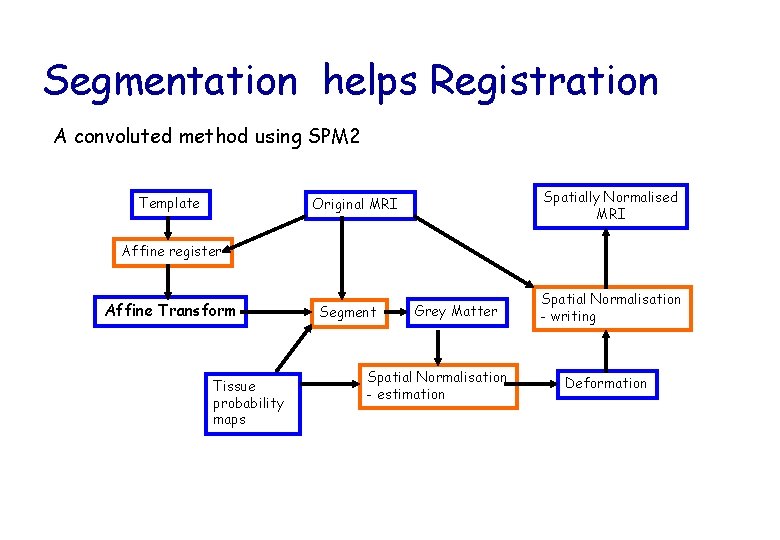
Segmentation helps Registration A convoluted method using SPM 2 Template Spatially Normalised MRI Original MRI Affine register Affine Transform Tissue probability maps Segment Grey Matter Spatial Normalisation - estimation Spatial Normalisation - writing Deformation

- Slides: 75