Visualisation of Multiple Sequence Alignments VIZBI 2011 Des

Visualisation of Multiple Sequence Alignments VIZBI 2011 Des Higgins Conway Institute University College Dublin Ireland

Multiple Alignment? • Align 3 or more sequences together – Homologous residues lined up in columns Whale myoglobin ----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT Lamprey globin GSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTP---EFFPKFKGLTT Lupin globin ---GALTESQAALVKSSWEEF--NIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE • Needed because of – Orthologues from different species But mainly: – Paralogues from Gene duplications • Multi-gene families – e. g. humans have approx. 500 protein kinases
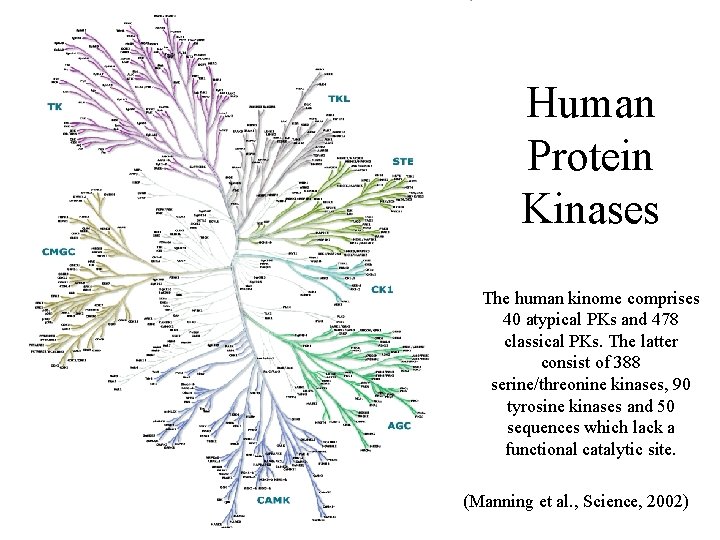
Human Protein Kinases The human kinome comprises 40 atypical PKs and 478 classical PKs. The latter consist of 388 serine/threonine kinases, 90 tyrosine kinases and 50 sequences which lack a functional catalytic site. (Manning et al. , Science, 2002)

Globin Multiple Alignment Human beta ----VHLTPEEKSAVTALWGKVN–-VDEVGGEALGRLLVVYPWTQRFFESFGDLST Horse beta ----VQLSGEEKAAVLALWDKVN–-EEEVGGEALGRLLVVYPWTQRFFDSFGDLSN Human alpha -----VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLSHorse alpha -----VLSAADKTNVKAAWSKVGGHAGEYGAEALERMFLGFPTTKTYFPHF-DLSWhale myoglobin -----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT Lamprey globin PIVDTGSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTPAAQEFFPKFKGLTT Lupin globin ----GALTESQAALVKSSWEEFNANIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE *: : : * . : . : * : . Human beta PDAVMGNPKVKAHGKKVLGAFSDGLAHLDN-----LKGTFATLSELHCDKLHVDPENFRL Horse beta PGAVMGNPKVKAHGKKVLHSFGEGVHHLDN-----LKGTFAALSELHCDKLHVDPENFRL Human alpha ----HGSAQVKGHGKKVADALTNAVAHVDD-----MPNALSALSDLHAHKLRVDPVNFKL Horse alpha ----HGSAQVKAHGKKVGDALTLAVGHLDD-----LPGALSNLSDLHAHKLRVDPVNFKL Whale myoglobin EAEMKASEDLKKHGVTVLTALGAILKKKGH-----HEAELKPLAQSHATKHKIPIKYLEF Lamprey globin ADQLKKSADVRWHAERIINAVNDAVASMDDT--EKMSMKLRDLSGKHAKSFQVDPQYFKV Lupin globin VP--QNNPELQAHAGKVFKLVYEAAIQLQVTGVVVTDATLKNLGSVHVSKGVAD-AHFPV . . : : *. : . : *. * . : . Human beta LGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH-----Horse beta LGNVLVVVLARHFGKDFTPELQASYQKVVAGVANALAHKYH-----Human alpha LSHCLLVTLAAHLPAEFTPAVHASLDKFLASVSTVLTSKYR-----Horse alpha LSHCLLSTLAVHLPNDFTPAVHASLDKFLSSVSTVLTSKYR-----Whale myoglobin ISEAIIHVLHSRHPGDFGADAQGAMNKALELFRKDIAAKYKELGYQG Lamprey globin LAAVIADTVAAG---D------AGFEKLMSMICILLRSAY------Lupin globin VKEAILKTIKEVVGAKWSEELNSAWTIAYDELAIVIKKEMNDAA--- 1. Visualise the residues/gaps?

Globin Multiple Alignment Human beta ----VHLTPEEKSAVTALWGKVN–-VDEVGGEALGRLLVVYPWTQRFFESFGDLST Horse beta ----VQLSGEEKAAVLALWDKVN–-EEEVGGEALGRLLVVYPWTQRFFDSFGDLSN Human alpha -----VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLSHorse alpha -----VLSAADKTNVKAAWSKVGGHAGEYGAEALERMFLGFPTTKTYFPHF-DLSWhale myoglobin -----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT Lamprey globin PIVDTGSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTPAAQEFFPKFKGLTT Lupin globin ----GALTESQAALVKSSWEEFNANIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE *: : : * . : . : * : . Human beta PDAVMGNPKVKAHGKKVLGAFSDGLAHLDN-----LKGTFATLSELHCDKLHVDPENFRL Horse beta PGAVMGNPKVKAHGKKVLHSFGEGVHHLDN-----LKGTFAALSELHCDKLHVDPENFRL Human alpha ----HGSAQVKGHGKKVADALTNAVAHVDD-----MPNALSALSDLHAHKLRVDPVNFKL Horse alpha ----HGSAQVKAHGKKVGDALTLAVGHLDD-----LPGALSNLSDLHAHKLRVDPVNFKL Whale myoglobin EAEMKASEDLKKHGVTVLTALGAILKKKGH-----HEAELKPLAQSHATKHKIPIKYLEF Lamprey globin ADQLKKSADVRWHAERIINAVNDAVASMDDT--EKMSMKLRDLSGKHAKSFQVDPQYFKV Lupin globin VP--QNNPELQAHAGKVFKLVYEAAIQLQVTGVVVTDATLKNLGSVHVSKGVAD-AHFPV . . : : *. : . : *. * . : . Human beta LGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH-----Horse beta LGNVLVVVLARHFGKDFTPELQASYQKVVAGVANALAHKYH-----Human alpha LSHCLLVTLAAHLPAEFTPAVHASLDKFLASVSTVLTSKYR-----Horse alpha LSHCLLSTLAVHLPNDFTPAVHASLDKFLSSVSTVLTSKYR-----Whale myoglobin ISEAIIHVLHSRHPGDFGADAQGAMNKALELFRKDIAAKYKELGYQG Lamprey globin LAAVIADTVAAG---D------AGFEKLMSMICILLRSAY------Lupin globin VKEAILKTIKEVVGAKWSEELNSAWTIAYDELAIVIKKEMNDAA---

Globin Multiple Alignment Human beta ----VHLTPEEKSAVTALWGKVN–-VDEVGGEALGRLLVVYPWTQRFFESFGDLST Horse beta ----VQLSGEEKAAVLALWDKVN–-EEEVGGEALGRLLVVYPWTQRFFDSFGDLSN Human alpha -----VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLSHorse alpha -----VLSAADKTNVKAAWSKVGGHAGEYGAEALERMFLGFPTTKTYFPHF-DLSWhale myoglobin -----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT Lamprey globin PIVDTGSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTPAAQEFFPKFKGLTT Lupin globin ----GALTESQAALVKSSWEEFNANIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE *: : : * . : . : * : . Human beta PDAVMGNPKVKAHGKKVLGAFSDGLAHLDN-----LKGTFATLSELHCDKLHVDPENFRL Horse beta PGAVMGNPKVKAHGKKVLHSFGEGVHHLDN-----LKGTFAALSELHCDKLHVDPENFRL Human alpha ----HGSAQVKGHGKKVADALTNAVAHVDD-----MPNALSALSDLHAHKLRVDPVNFKL Horse alpha ----HGSAQVKAHGKKVGDALTLAVGHLDD-----LPGALSNLSDLHAHKLRVDPVNFKL Whale myoglobin EAEMKASEDLKKHGVTVLTALGAILKKKGH-----HEAELKPLAQSHATKHKIPIKYLEF Lamprey globin ADQLKKSADVRWHAERIINAVNDAVASMDDT--EKMSMKLRDLSGKHAKSFQVDPQYFKV Lupin globin VP--QNNPELQAHAGKVFKLVYEAAIQLQVTGVVVTDATLKNLGSVHVSKGVAD-AHFPV . . : : *. : . : *. * . : . Human beta LGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH-----Horse beta LGNVLVVVLARHFGKDFTPELQASYQKVVAGVANALAHKYH-----Human alpha LSHCLLVTLAAHLPAEFTPAVHASLDKFLASVSTVLTSKYR-----Horse alpha LSHCLLSTLAVHLPNDFTPAVHASLDKFLSSVSTVLTSKYR-----Whale myoglobin ISEAIIHVLHSRHPGDFGADAQGAMNKALELFRKDIAAKYKELGYQG Lamprey globin LAAVIADTVAAG---D------AGFEKLMSMICILLRSAY------Lupin globin VKEAILKTIKEVVGAKWSEELNSAWTIAYDELAIVIKKEMNDAA--- Alpha helices

Globin Multiple Alignment Human beta ----VHLTPEEKSAVTALWGKVN–-VDEVGGEALGRLLVVYPWTQRFFESFGDLST Horse beta ----VQLSGEEKAAVLALWDKVN–-EEEVGGEALGRLLVVYPWTQRFFDSFGDLSN Human alpha -----VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLSHorse alpha -----VLSAADKTNVKAAWSKVGGHAGEYGAEALERMFLGFPTTKTYFPHF-DLSWhale myoglobin -----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT Lamprey globin PIVDTGSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTPAAQEFFPKFKGLTT Lupin globin ----GALTESQAALVKSSWEEFNANIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE *: : : * . : . : * : . Human beta PDAVMGNPKVKAHGKKVLGAFSDGLAHLDN-----LKGTFATLSELHCDKLHVDPENFRL Horse beta PGAVMGNPKVKAHGKKVLHSFGEGVHHLDN-----LKGTFAALSELHCDKLHVDPENFRL Human alpha ----HGSAQVKGHGKKVADALTNAVAHVDD-----MPNALSALSDLHAHKLRVDPVNFKL Horse alpha ----HGSAQVKAHGKKVGDALTLAVGHLDD-----LPGALSNLSDLHAHKLRVDPVNFKL Whale myoglobin EAEMKASEDLKKHGVTVLTALGAILKKKGH-----HEAELKPLAQSHATKHKIPIKYLEF Lamprey globin ADQLKKSADVRWHAERIINAVNDAVASMDDT--EKMSMKLRDLSGKHAKSFQVDPQYFKV Lupin globin VP--QNNPELQAHAGKVFKLVYEAAIQLQVTGVVVTDATLKNLGSVHVSKGVAD-AHFPV . . : : *. : . : *. * . : . Human beta LGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH-----Horse beta LGNVLVVVLARHFGKDFTPELQASYQKVVAGVANALAHKYH-----Human alpha LSHCLLVTLAAHLPAEFTPAVHASLDKFLASVSTVLTSKYR-----Horse alpha LSHCLLSTLAVHLPNDFTPAVHASLDKFLSSVSTVLTSKYR-----Whale myoglobin ISEAIIHVLHSRHPGDFGADAQGAMNKALELFRKDIAAKYKELGYQG Lamprey globin LAAVIADTVAAG---D------AGFEKLMSMICILLRSAY------Lupin globin VKEAILKTIKEVVGAKWSEELNSAWTIAYDELAIVIKKEMNDAA--- Haem binding Histidines
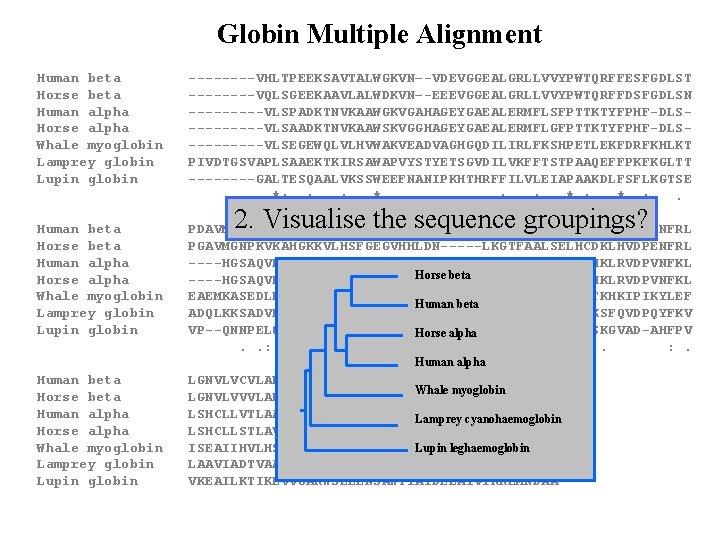
Globin Multiple Alignment Human beta ----VHLTPEEKSAVTALWGKVN–-VDEVGGEALGRLLVVYPWTQRFFESFGDLST Horse beta ----VQLSGEEKAAVLALWDKVN–-EEEVGGEALGRLLVVYPWTQRFFDSFGDLSN Human alpha -----VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLSHorse alpha -----VLSAADKTNVKAAWSKVGGHAGEYGAEALERMFLGFPTTKTYFPHF-DLSWhale myoglobin -----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT Lamprey globin PIVDTGSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTPAAQEFFPKFKGLTT Lupin globin ----GALTESQAALVKSSWEEFNANIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE *: : : * . : . : * : . Human beta PDAVMGNPKVKAHGKKVLGAFSDGLAHLDN-----LKGTFATLSELHCDKLHVDPENFRL Horse beta PGAVMGNPKVKAHGKKVLHSFGEGVHHLDN-----LKGTFAALSELHCDKLHVDPENFRL Human alpha ----HGSAQVKGHGKKVADALTNAVAHVDD-----MPNALSALSDLHAHKLRVDPVNFKL Horse beta Horse alpha ----HGSAQVKAHGKKVGDALTLAVGHLDD-----LPGALSNLSDLHAHKLRVDPVNFKL Whale myoglobin EAEMKASEDLKKHGVTVLTALGAILKKKGH-----HEAELKPLAQSHATKHKIPIKYLEF Human beta Lamprey globin ADQLKKSADVRWHAERIINAVNDAVASMDDT--EKMSMKLRDLSGKHAKSFQVDPQYFKV Lupin globin VP--QNNPELQAHAGKVFKLVYEAAIQLQVTGVVVTDATLKNLGSVHVSKGVAD-AHFPV Horse alpha . . : : *. : . : *. * . : . Human alpha Human beta LGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH-----Whale myoglobin Horse beta LGNVLVVVLARHFGKDFTPELQASYQKVVAGVANALAHKYH----- Human alpha LSHCLLVTLAAHLPAEFTPAVHASLDKFLASVSTVLTSKYR-----Lamprey cyanohaemoglobin Horse alpha LSHCLLSTLAVHLPNDFTPAVHASLDKFLSSVSTVLTSKYR----- Whale myoglobin ISEAIIHVLHSRHPGDFGADAQGAMNKALELFRKDIAAKYKELGYQG Lupin leghaemoglobin Lamprey globin LAAVIADTVAAG---D------AGFEKLMSMICILLRSAY------Lupin globin VKEAILKTIKEVVGAKWSEELNSAWTIAYDELAIVIKKEMNDAA--- 2. Visualise the sequence groupings?

So: What is the Problem? • What if N >> 100, 000? • e. g. SSU r. RNA – www. arb-silva. de – 1, 471, 257 seqs • e. g. ABC transporters – PFAM – ABC_tran PF 00005 – 127, 458 seqs • Metagenomics
• Sequence 10, 000 vertebrate genomes! =>5, 000 protein kinases, GPCRs

Sequence. Juxtaposer: Fluid Navigation For Large-Scale Sequence Comparison In Context James Slack Kristian Hildebrandy Tamara Munzner Katherine St. John. Proc. German Conference on Bioinformatics 2004, pp 3742 Poster D 03 VIZBI, 2011 Sequence Surveyor: scalable multiple sequence alignment overview visualisation. Danielle Albers, Colin Dewey, Michael Gleicher Poster D 09 VIZBI, 2011 JProfile. Grid: visualising very large multiple sequence alignments. Alberto Roca, Aaron Abajian, David Vigerust

This talk • How to make huge multiple alignments • How to cluster > 100, 000 sequences • MDS/PCA on big datasets

Multiple Sequence Alignment • NP complete • Mainly use: “Progressive Alignment” – Greedy heuristic – Use a tree/clustering of the seqs • Barton and Sternberg (1988) Feng and Doolittle (1987) Higgins and Sharp (1988) Hogeweg and Hesper (1984) Willlie Taylor (1987)

Human beta ----VHLTPEEKSAVTALWGKVN–-VDEVGGEALGRLLVVYPWTQRFFESFGDLST Horse beta ----VQLSGEEKAAVLALWDKVN–-EEEVGGEALGRLLVVYPWTQRFFDSFGDLSN Human alpha -----VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLSHorse alpha -----VLSAADKTNVKAAWSKVGGHAGEYGAEALERMFLGFPTTKTYFPHF-DLSWhale myoglobin -----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT Lamprey globin PIVDTGSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTPAAQEFFPKFKGLTT Lupin globin ----GALTESQAALVKSSWEEFNANIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE *: : : * . : . : * : . Human beta PDAVMGNPKVKAHGKKVLGAFSDGLAHLDN-----LKGTFATLSELHCDKLHVDPENFRL Horse beta PGAVMGNPKVKAHGKKVLHSFGEGVHHLDN-----LKGTFAALSELHCDKLHVDPENFRL Human alpha ----HGSAQVKGHGKKVADALTNAVAHVDD-----MPNALSALSDLHAHKLRVDPVNFKL Horse alpha ----HGSAQVKAHGKKVGDALTLAVGHLDD-----LPGALSNLSDLHAHKLRVDPVNFKL Whale myoglobin EAEMKASEDLKKHGVTVLTALGAILKKKGH-----HEAELKPLAQSHATKHKIPIKYLEF Lamprey globin ADQLKKSADVRWHAERIINAVNDAVASMDDT--EKMSMKLRDLSGKHAKSFQVDPQYFKV Lupin globin VP--QNNPELQAHAGKVFKLVYEAAIQLQVTGVVVTDATLKNLGSVHVSKGVAD-AHFPV . . : : *. : . : *. * . : . Human beta LGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH-----Horse beta LGNVLVVVLARHFGKDFTPELQASYQKVVAGVANALAHKYH-----Human alpha LSHCLLVTLAAHLPAEFTPAVHASLDKFLASVSTVLTSKYR-----Horse alpha LSHCLLSTLAVHLPNDFTPAVHASLDKFLSSVSTVLTSKYR-----Whale myoglobin ISEAIIHVLHSRHPGDFGADAQGAMNKALELFRKDIAAKYKELGYQG Lamprey globin LAAVIADTVAAG---D------AGFEKLMSMICILLRSAY------Lupin globin VKEAILKTIKEVVGAKWSEELNSAWTIAYDELAIVIKKEMNDAA-- : : . . . . : Horse beta Human beta Horse alpha Human alpha Whale myoglobin Lamprey cyanohaemoglobin Lupin leghaemoglobin “Guide Tree”
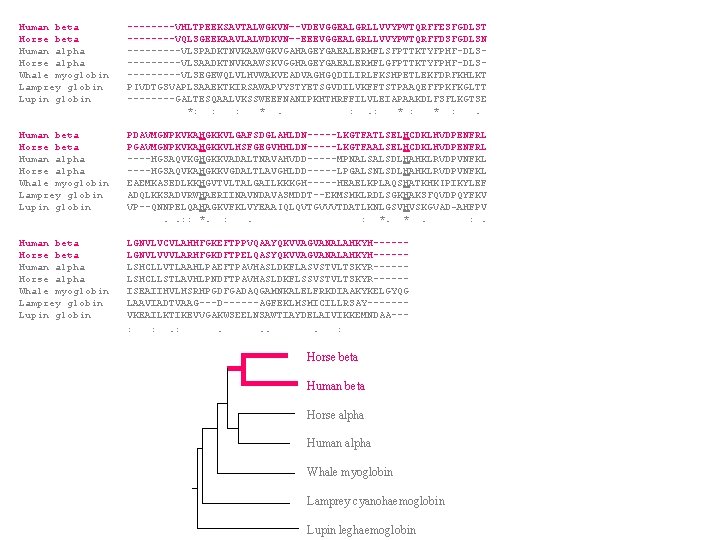
Human beta ----VHLTPEEKSAVTALWGKVN–-VDEVGGEALGRLLVVYPWTQRFFESFGDLST Horse beta ----VQLSGEEKAAVLALWDKVN–-EEEVGGEALGRLLVVYPWTQRFFDSFGDLSN Human alpha -----VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLSHorse alpha -----VLSAADKTNVKAAWSKVGGHAGEYGAEALERMFLGFPTTKTYFPHF-DLSWhale myoglobin -----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT Lamprey globin PIVDTGSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTPAAQEFFPKFKGLTT Lupin globin ----GALTESQAALVKSSWEEFNANIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE *: : : * . : . : * : . Human beta PDAVMGNPKVKAHGKKVLGAFSDGLAHLDN-----LKGTFATLSELHCDKLHVDPENFRL Horse beta PGAVMGNPKVKAHGKKVLHSFGEGVHHLDN-----LKGTFAALSELHCDKLHVDPENFRL Human alpha ----HGSAQVKGHGKKVADALTNAVAHVDD-----MPNALSALSDLHAHKLRVDPVNFKL Horse alpha ----HGSAQVKAHGKKVGDALTLAVGHLDD-----LPGALSNLSDLHAHKLRVDPVNFKL Whale myoglobin EAEMKASEDLKKHGVTVLTALGAILKKKGH-----HEAELKPLAQSHATKHKIPIKYLEF Lamprey globin ADQLKKSADVRWHAERIINAVNDAVASMDDT--EKMSMKLRDLSGKHAKSFQVDPQYFKV Lupin globin VP--QNNPELQAHAGKVFKLVYEAAIQLQVTGVVVTDATLKNLGSVHVSKGVAD-AHFPV . . : : *. : . : *. * . : . Human beta LGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH-----Horse beta LGNVLVVVLARHFGKDFTPELQASYQKVVAGVANALAHKYH-----Human alpha LSHCLLVTLAAHLPAEFTPAVHASLDKFLASVSTVLTSKYR-----Horse alpha LSHCLLSTLAVHLPNDFTPAVHASLDKFLSSVSTVLTSKYR-----Whale myoglobin ISEAIIHVLHSRHPGDFGADAQGAMNKALELFRKDIAAKYKELGYQG Lamprey globin LAAVIADTVAAG---D------AGFEKLMSMICILLRSAY------Lupin globin VKEAILKTIKEVVGAKWSEELNSAWTIAYDELAIVIKKEMNDAA-- : : . . . . : Horse beta Human beta Horse alpha Human alpha Whale myoglobin Lamprey cyanohaemoglobin Lupin leghaemoglobin
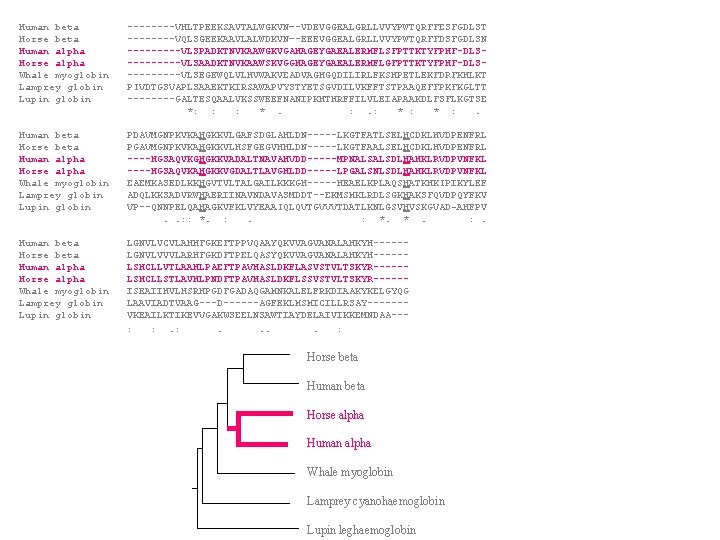
Human beta ----VHLTPEEKSAVTALWGKVN–-VDEVGGEALGRLLVVYPWTQRFFESFGDLST Horse beta ----VQLSGEEKAAVLALWDKVN–-EEEVGGEALGRLLVVYPWTQRFFDSFGDLSN Human alpha -----VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLSHorse alpha -----VLSAADKTNVKAAWSKVGGHAGEYGAEALERMFLGFPTTKTYFPHF-DLSWhale myoglobin -----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT Lamprey globin PIVDTGSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTPAAQEFFPKFKGLTT Lupin globin ----GALTESQAALVKSSWEEFNANIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE *: : : * . : . : * : . Human beta PDAVMGNPKVKAHGKKVLGAFSDGLAHLDN-----LKGTFATLSELHCDKLHVDPENFRL Horse beta PGAVMGNPKVKAHGKKVLHSFGEGVHHLDN-----LKGTFAALSELHCDKLHVDPENFRL Human alpha ----HGSAQVKGHGKKVADALTNAVAHVDD-----MPNALSALSDLHAHKLRVDPVNFKL Horse alpha ----HGSAQVKAHGKKVGDALTLAVGHLDD-----LPGALSNLSDLHAHKLRVDPVNFKL Whale myoglobin EAEMKASEDLKKHGVTVLTALGAILKKKGH-----HEAELKPLAQSHATKHKIPIKYLEF Lamprey globin ADQLKKSADVRWHAERIINAVNDAVASMDDT--EKMSMKLRDLSGKHAKSFQVDPQYFKV Lupin globin VP--QNNPELQAHAGKVFKLVYEAAIQLQVTGVVVTDATLKNLGSVHVSKGVAD-AHFPV . . : : *. : . : *. * . : . Human beta LGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH-----Horse beta LGNVLVVVLARHFGKDFTPELQASYQKVVAGVANALAHKYH-----Human alpha LSHCLLVTLAAHLPAEFTPAVHASLDKFLASVSTVLTSKYR-----Horse alpha LSHCLLSTLAVHLPNDFTPAVHASLDKFLSSVSTVLTSKYR-----Whale myoglobin ISEAIIHVLHSRHPGDFGADAQGAMNKALELFRKDIAAKYKELGYQG Lamprey globin LAAVIADTVAAG---D------AGFEKLMSMICILLRSAY------Lupin globin VKEAILKTIKEVVGAKWSEELNSAWTIAYDELAIVIKKEMNDAA-- : : . . . . : Horse beta Human beta Horse alpha Human alpha Whale myoglobin Lamprey cyanohaemoglobin Lupin leghaemoglobin
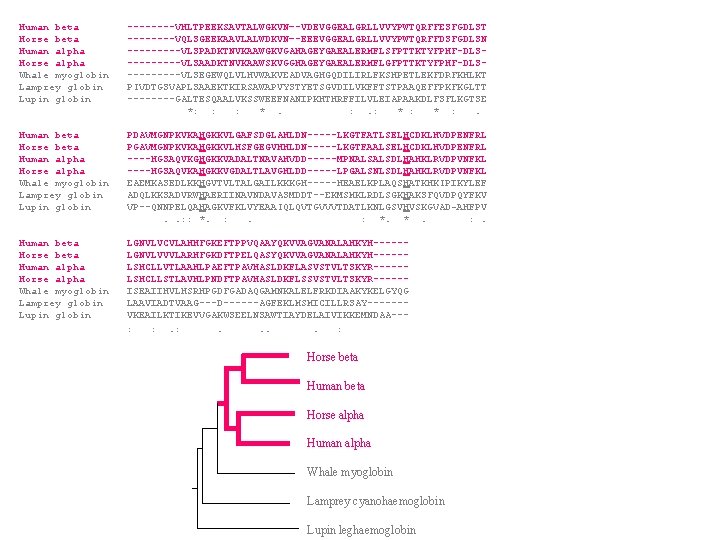
Human beta ----VHLTPEEKSAVTALWGKVN–-VDEVGGEALGRLLVVYPWTQRFFESFGDLST Horse beta ----VQLSGEEKAAVLALWDKVN–-EEEVGGEALGRLLVVYPWTQRFFDSFGDLSN Human alpha -----VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLSHorse alpha -----VLSAADKTNVKAAWSKVGGHAGEYGAEALERMFLGFPTTKTYFPHF-DLSWhale myoglobin -----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT Lamprey globin PIVDTGSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTPAAQEFFPKFKGLTT Lupin globin ----GALTESQAALVKSSWEEFNANIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE *: : : * . : . : * : . Human beta PDAVMGNPKVKAHGKKVLGAFSDGLAHLDN-----LKGTFATLSELHCDKLHVDPENFRL Horse beta PGAVMGNPKVKAHGKKVLHSFGEGVHHLDN-----LKGTFAALSELHCDKLHVDPENFRL Human alpha ----HGSAQVKGHGKKVADALTNAVAHVDD-----MPNALSALSDLHAHKLRVDPVNFKL Horse alpha ----HGSAQVKAHGKKVGDALTLAVGHLDD-----LPGALSNLSDLHAHKLRVDPVNFKL Whale myoglobin EAEMKASEDLKKHGVTVLTALGAILKKKGH-----HEAELKPLAQSHATKHKIPIKYLEF Lamprey globin ADQLKKSADVRWHAERIINAVNDAVASMDDT--EKMSMKLRDLSGKHAKSFQVDPQYFKV Lupin globin VP--QNNPELQAHAGKVFKLVYEAAIQLQVTGVVVTDATLKNLGSVHVSKGVAD-AHFPV . . : : *. : . : *. * . : . Human beta LGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH-----Horse beta LGNVLVVVLARHFGKDFTPELQASYQKVVAGVANALAHKYH-----Human alpha LSHCLLVTLAAHLPAEFTPAVHASLDKFLASVSTVLTSKYR-----Horse alpha LSHCLLSTLAVHLPNDFTPAVHASLDKFLSSVSTVLTSKYR-----Whale myoglobin ISEAIIHVLHSRHPGDFGADAQGAMNKALELFRKDIAAKYKELGYQG Lamprey globin LAAVIADTVAAG---D------AGFEKLMSMICILLRSAY------Lupin globin VKEAILKTIKEVVGAKWSEELNSAWTIAYDELAIVIKKEMNDAA-- : : . . . . : Horse beta Human beta Horse alpha Human alpha Whale myoglobin Lamprey cyanohaemoglobin Lupin leghaemoglobin
Clustal • 66, 000 citations • Clustal 1 -Clustal 4 – 1988, Paul Sharp, Dublin • Clustal V 1992 – EMBL Heidelberg, – Rainer Fuchs – Alan Bleasby • Clustal W, Clustal X 1994 -2005 – Toby Gibson, EMBL, Heidelberg – Julie Thompson, ICGEB, Strasbourg • Clustal W and Clustal X 2. 0 2007 – University College Dublin www. clustal. org

Complexity • Guide tree construction O(N 2) • Later Progressive Alignment O(N) • Guide tree construction is limiting >10, 000 seq alignment is tough

Part. Tree • • MAFFT Package Select n sequences where n << N UPGMA on n sequences Cluster the remainder (N-n) with their closest clusters Katoh, K. , Toh, H. , 2007. Part. Tree: an algorithm to build an approximate tree from a large number of unaligned sequences. Bioinformatics 23, 372– 374.

Embedding? • Replace each sequence by a Vector – Vector-Vector distances • MUCH faster than • Seq. – Seq. distances • Vectors very fast/simple to cluster • e. g. cluster 10, 000 vectors of length 150 • <<1 min on 1 processor • UPGMA • e. g. cluster 300, 000 vectors of length 300 • 6 mins • k-means, k = 300

Embedding papers • Fast. Map • Faloutsos, C. , Lin, K. (1995) Fast. Map: A Fast Algorithm for Indexing Data-Mining and Visualisation of Traditional and Multimedia Datasets, Proc. 1995 ACM SIGMOD International Con. on Management of Data, pp. 163– 174. • Sparsemap • G. Hristescu and M. Farach-Colton. Cluster-preserving embedding of proteins. Technical Report 99 -50, Computer Science Department, Rutgers University, 1999.

m. BED • Select k seqs “randomly” – k << N – k α log. N • Use distance to each of these k “references” – k long vector for each sequence • Use heuristics – avoid duplicates – find outliers • Very fast and simple – Complexity O(k. N) i. e. O(Nlog. N) • Blackshields G, Sievers F, Shi W, Wilm A, Higgins DG. (2010) Sequence embedding for fast construction of guide trees for multiple sequence alignment. Algorithms Mol Biol. 14; 5: 21.
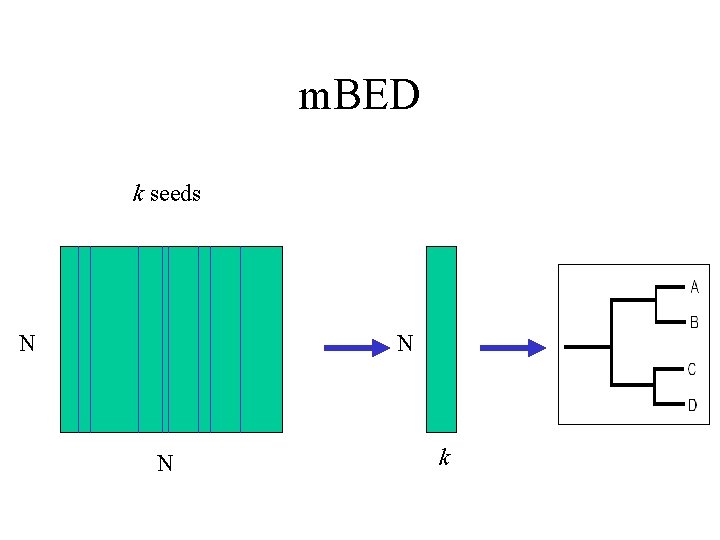
m. BED k seeds N N N k
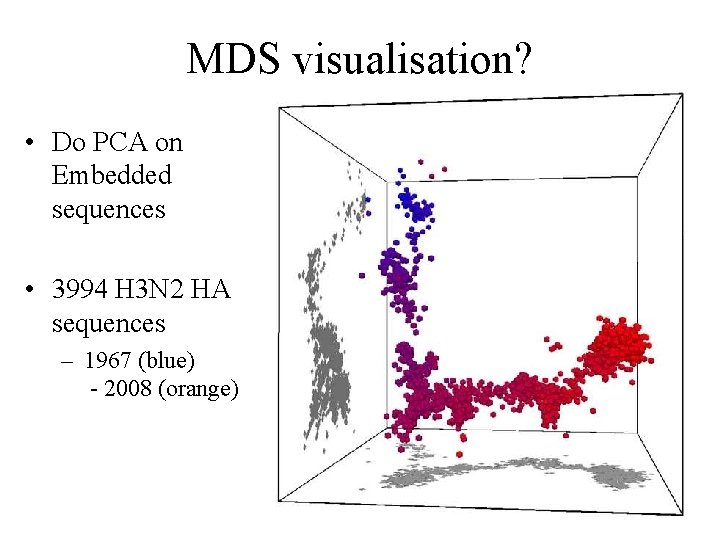
MDS visualisation? • Do PCA on Embedded sequences • 3994 H 3 N 2 HA sequences – 1967 (blue) - 2008 (orange)
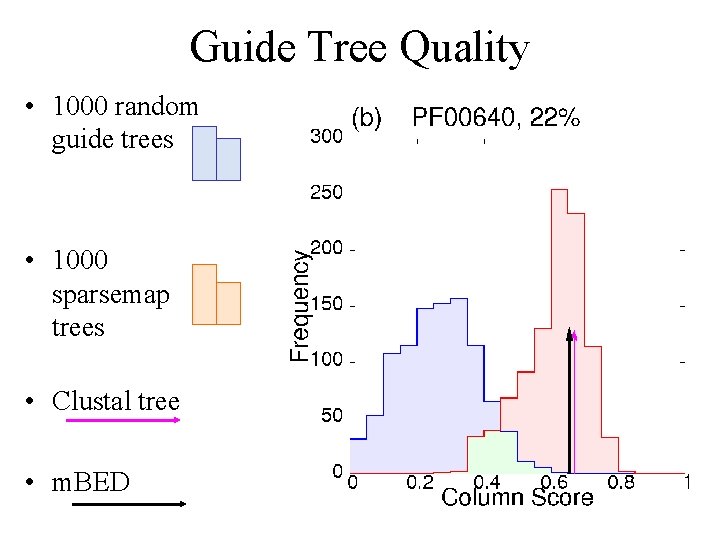
Guide Tree Quality • 1000 random guide trees • 1000 sparsemap trees • Clustal tree • m. BED

Clustal Ω • Release first version by April 2011 • Scalable – m. Bed – Gordon Blackshields • Accurate – HMM-HMM alignment – HHalign – Johannes Söding, Munich. • Re-use old alignments – Kevin Karplus – UCSC

• Align 120, 000 abc transporters – 6 hours on 1 core • More accurate than – MUSCLE or MAFFT • Coming soon. . . Fabian Sievers Andreas Wilm David Dineen

MDS/PCA etc. • Dimension reduction • Treat alignment columns as variables – PCA • Principal Components Analysis – CA • Correspondence Analysis, Jean Paul Benzécri • Use Nx. N distance matrix – MDS – PCOORD
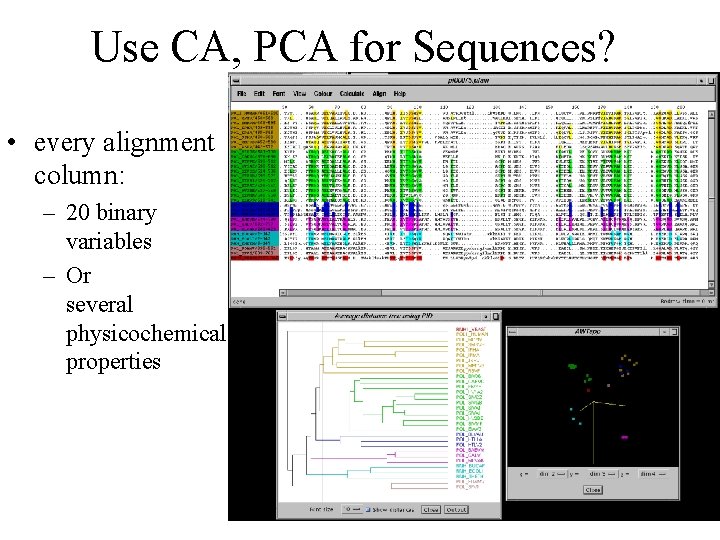
Use CA, PCA for Sequences? • every alignment column: – 20 binary variables – Or several physicochemical properties

Trypsin-like serine proteases 15 Chymotrypsins 10 Elastases • Correspondence Analysis • Supervise: • Between Groups Analysis • Dolédec and Chessel (1987) (similar to PLS discriminant analysis) 31 Trypsins
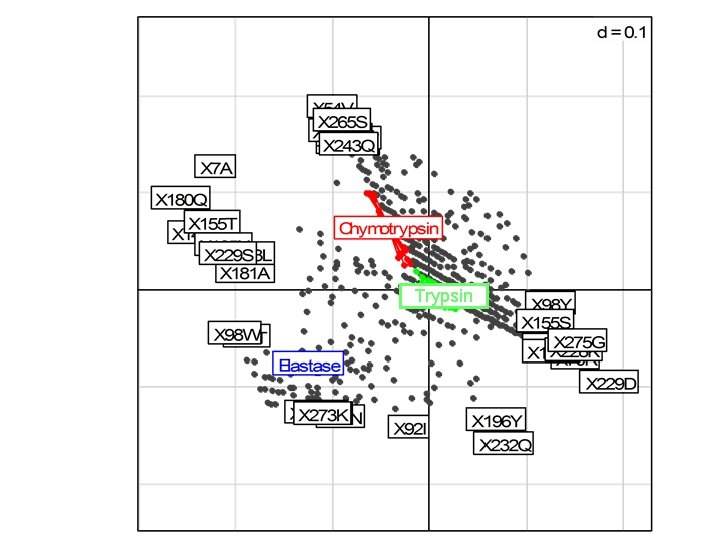
Trypsin
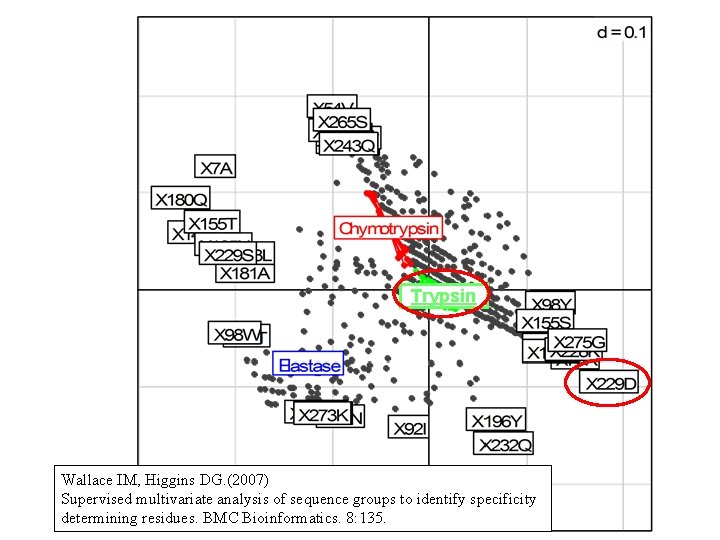
Trypsin Wallace IM, Higgins DG. (2007) Supervised multivariate analysis of sequence groups to identify specificity determining residues. BMC Bioinformatics. 8: 135.


MDS • Multidimensional Scaling • Fit distances to a Nx. N distance matrix • Use euclidean distances? – “Classical scaling” = Principal Co-Ordinates Analysis • PCOORD, John Gower – Gower, J. C. (1966). Some distance properties of latent root and vector methods used in multivariate analysis. Biometrika 53, 325 -328. – Higgins, D. G. (1992) Sequence ordinations: a multivariate analysis approach to analysing large sequence data sets. CABIOS, 8, 15 -22. – Complexity at least O(N 2)

Large scale MDS? • SC-MDS • Jengnan Tzeng, Henry Horng-Shing Lu, and Wen-Hsiung Li (2008) Multidimensional scaling for large genomic data sets BMC Bioinformatics. 2008; 9: 179. Easily do MDS on >100, 000 seqs • m. BED • Blackshields et al. , (2010) • PCOORD or MDS on a subset of the sequences • add the rest later • Landmark MDS + Nystrom approximation • V. de Silva, J. B. Tenenbaum, “Sparse multidimensional scaling using landmark points. ” (2004) Technical report, Stanford University.
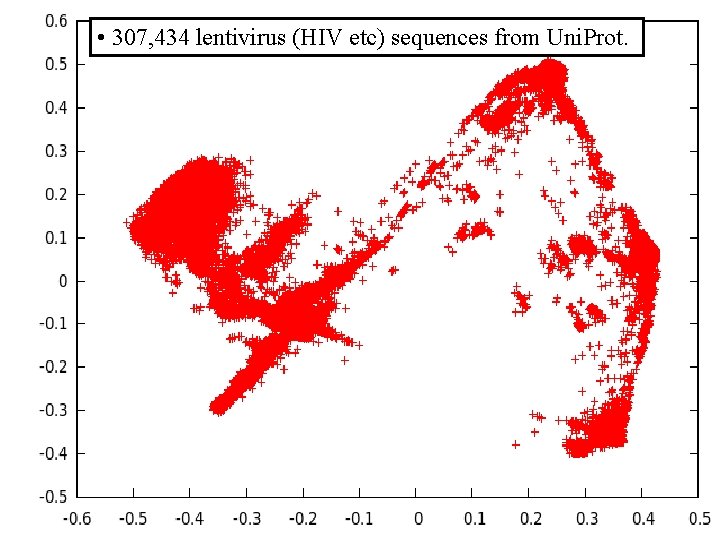
• 307, 434 lentivirus (HIV etc) sequences from Uni. Prot.

H 3 N 2 flu sequences • Weifeng Shi • 8167 HA sequences – human H 3 N 2 influenza viruses • DNAdist in Phylip – K 2 P (Kimura two parameter) model • Python: Matplotl. Ib
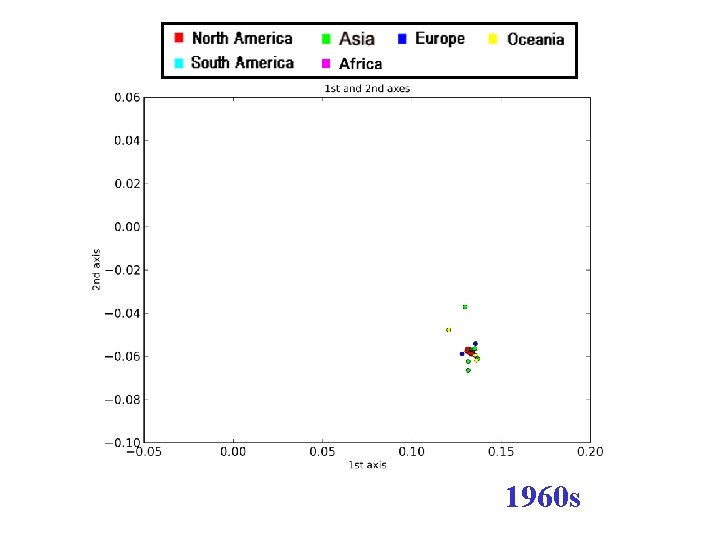
1960 s
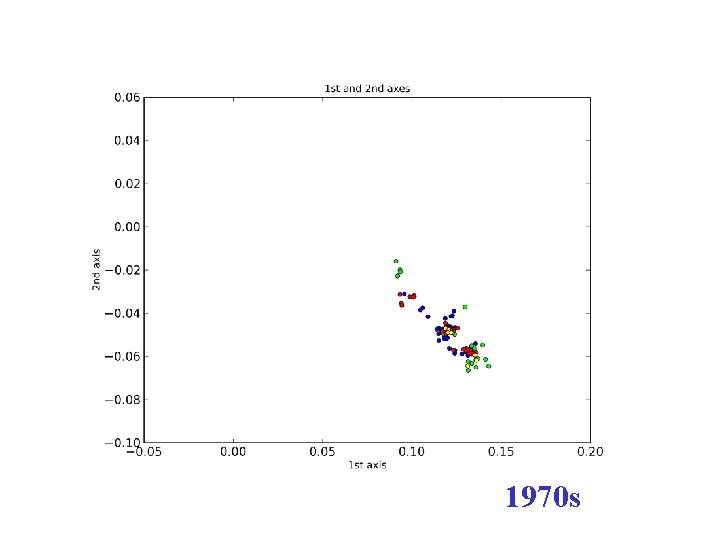
1970 s
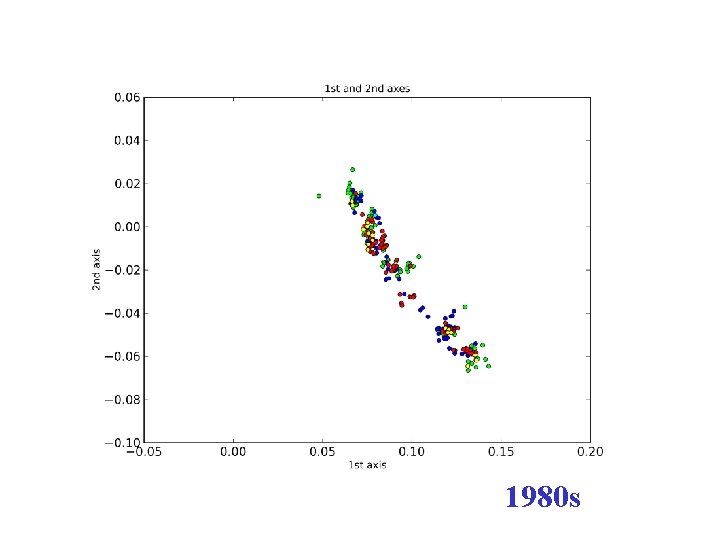
1980 s
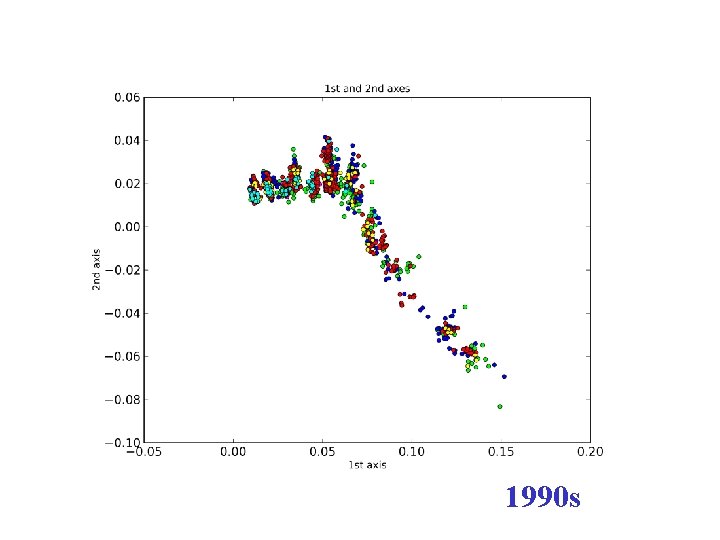
1990 s
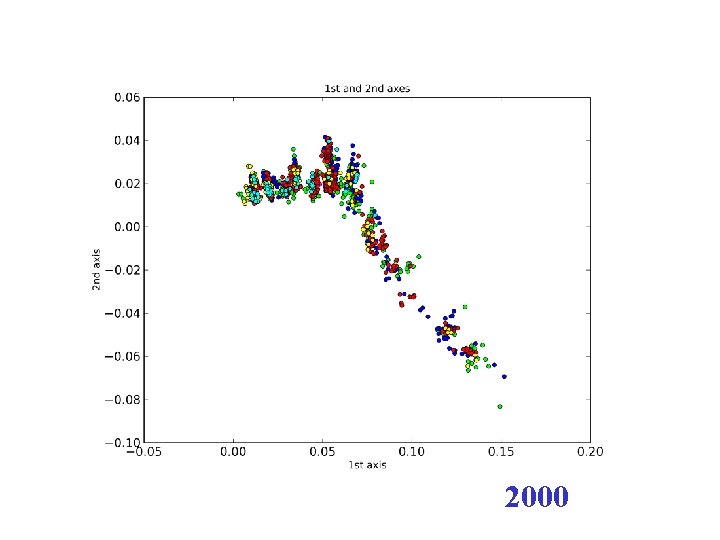
2000
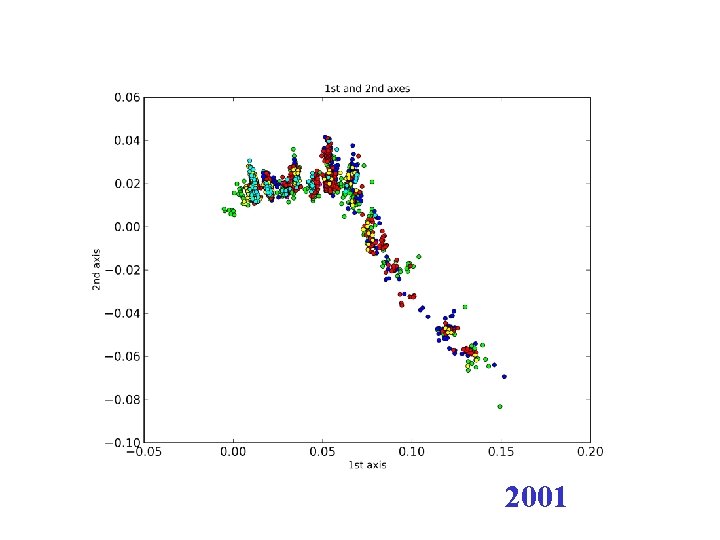
2001
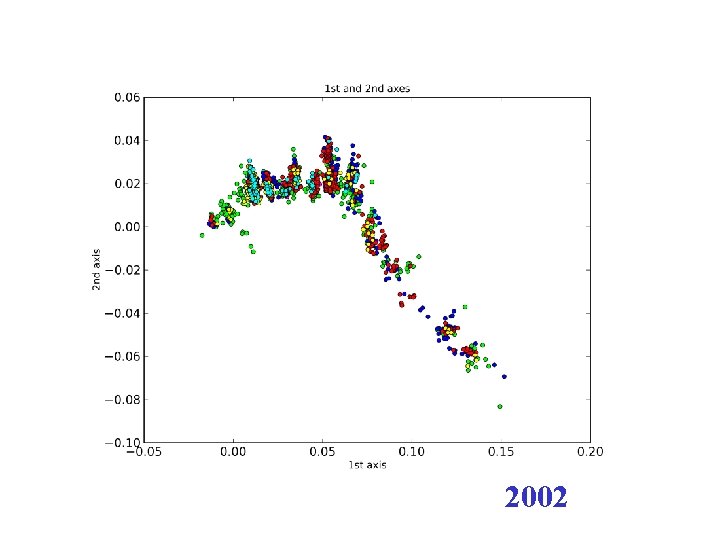
2002
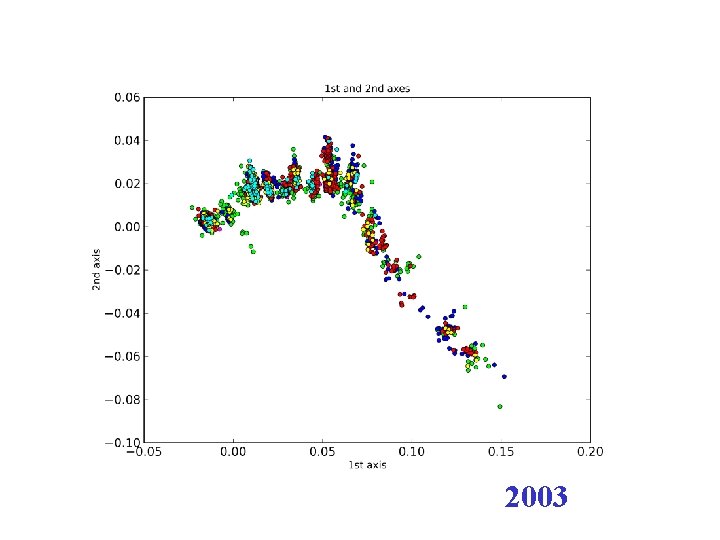
2003
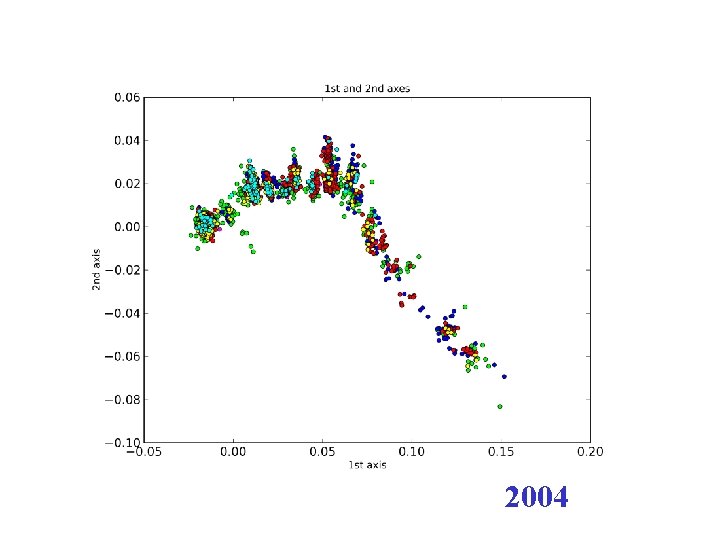
2004
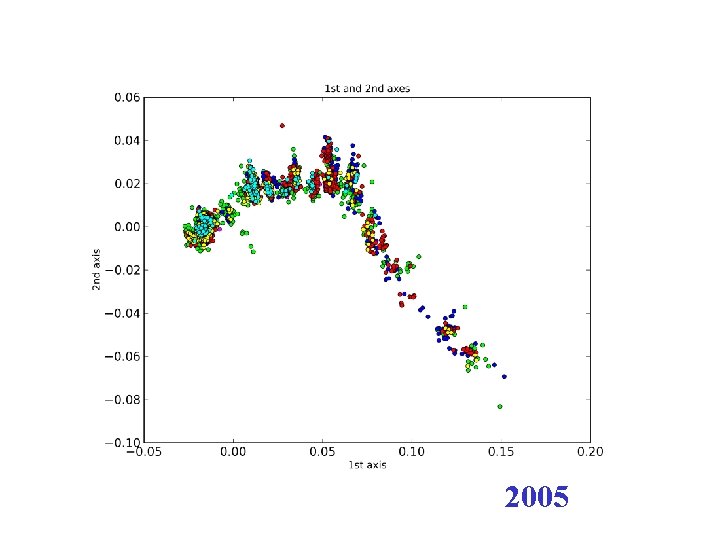
2005
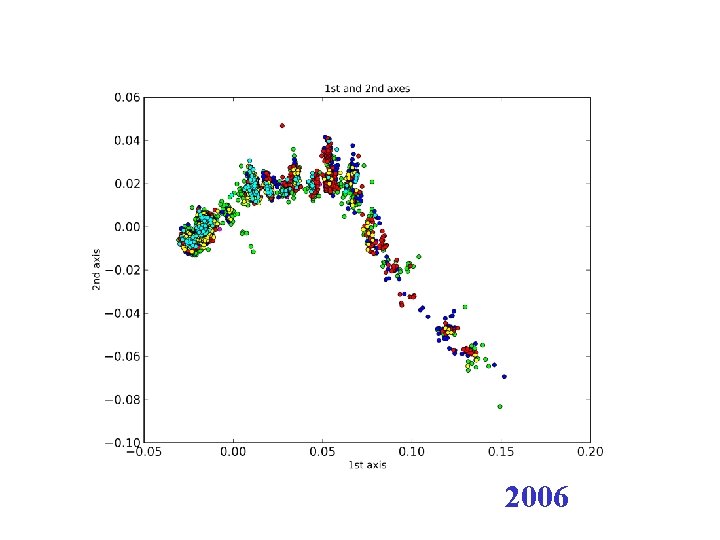
2006
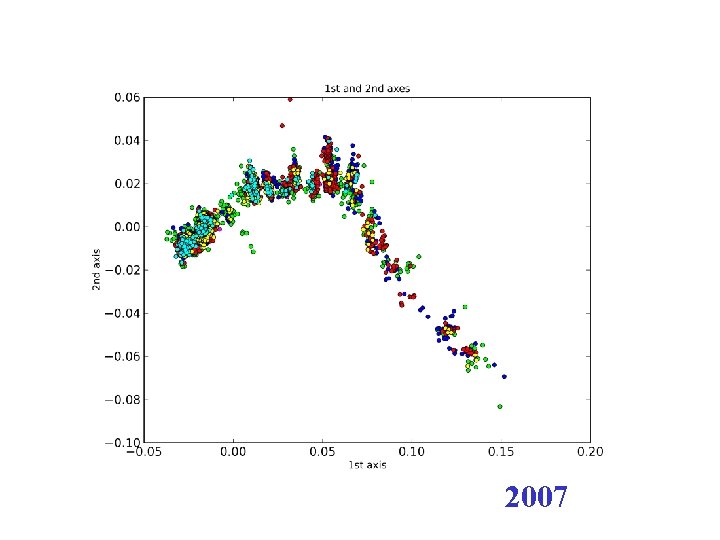
2007
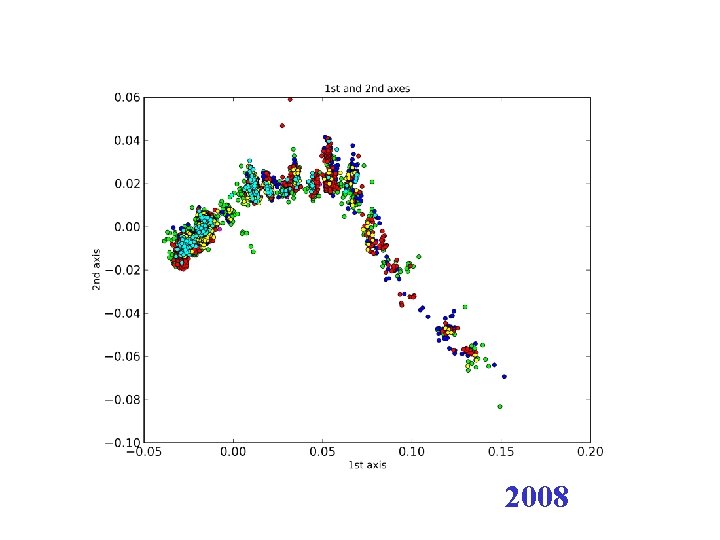
2008
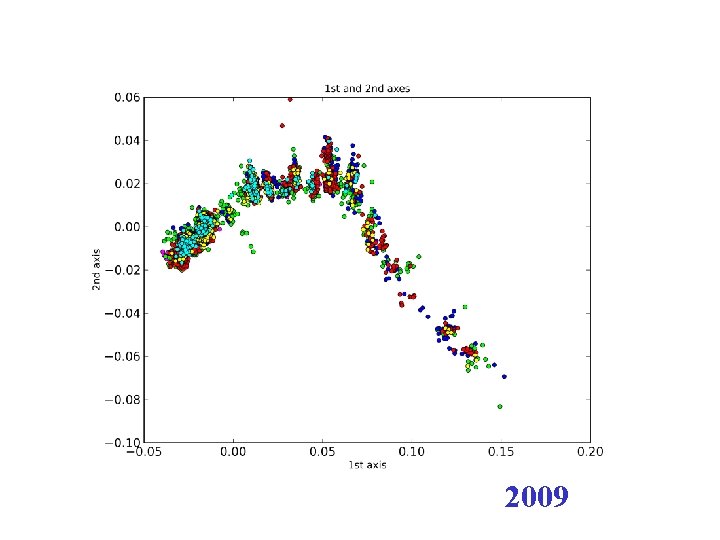
2009
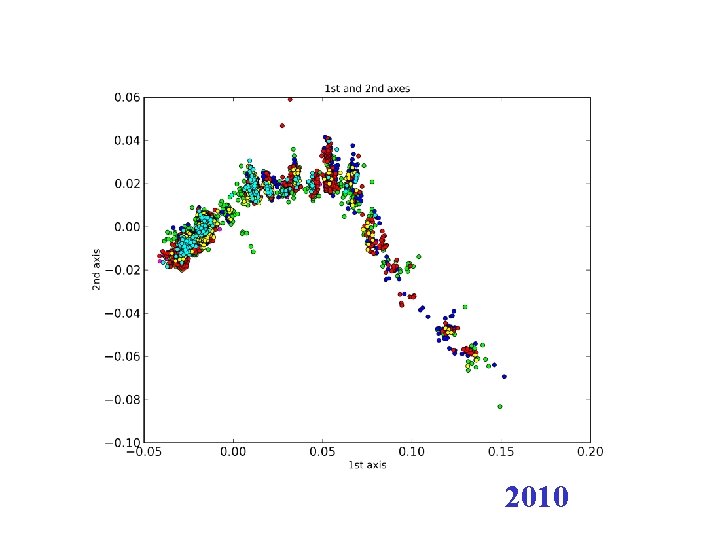
2010

BGA, CIA m. BED Aedin Culhane Ian Jeffery Stephen Madden Iain Wallace Guy Perriere, Lyons Gordon Blackshields Mark Larkin Clustal Omega Flu MDS Fabian Sievers Andreas Wilm David Dineen Johannes Soeding, Munich Rodrigo Lopez, EBI Weifeng Shi

Supervised PCA or CA? Malate Dehydrogenases Lactate Dehydrogenases

ADE-4 http: //pbil. univ-lyon 1. fr/ADE-4/ Thioulouse J. , Chessel D. , Dolédec S. , & Olivier J. M. (1997) ADE-4: a multivariate analysis and graphical display software. Statistics and Computing, 7, 1, 75 -83.

Between Group Analysis BGA Dolédec, S. & Chessel, D. (1987) Acta Oecologica, Oecologica Generalis, 8, 3, 403 -426. Supervised Correspondence Analysis or PCA CO-Inertia Analysis CIA Dolédec, S. & Chessel, D. (1994) Freshwater Biology, 31, 277 -294. Thioulouse, J. & Lobry, J. R. (1995) CABIOS, 11, 321 -329 2 datasets; Simultaneous CA or PCA • MADE 4 – Culhane, A. , Thiolouse, J. , Perriere, G. , Higgins, D. G. (2005) MADE 4: an R package for multivariate analysis of gene expression data. Bioinformatics. 21(11): 2789 -2790.
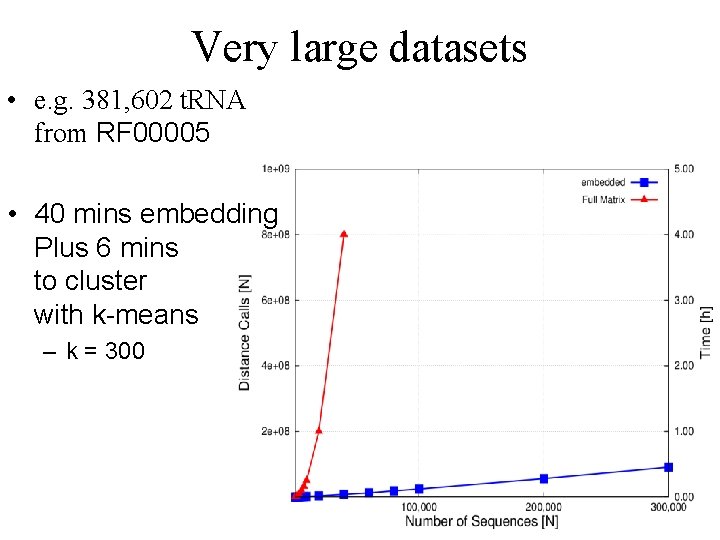
Very large datasets • e. g. 381, 602 t. RNA from RF 00005 • 40 mins embedding Plus 6 mins to cluster with k-means – k = 300
- Slides: 59