Vision and Ultrafast Chemistry Light Visual signaling Rod












- Slides: 30

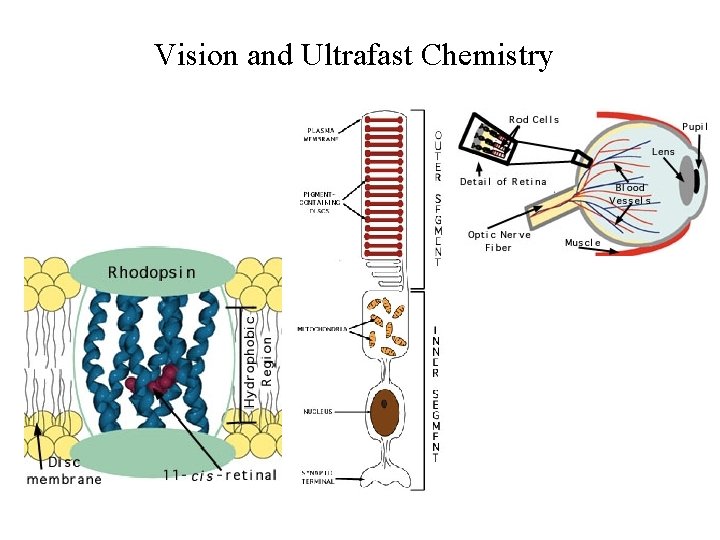
Vision and Ultrafast Chemistry

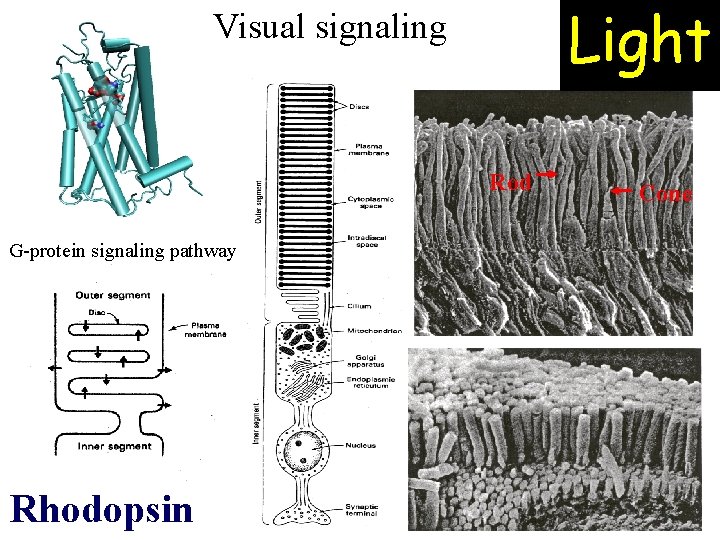
Light Visual signaling Rod G-protein signaling pathway Rhodopsin Cone

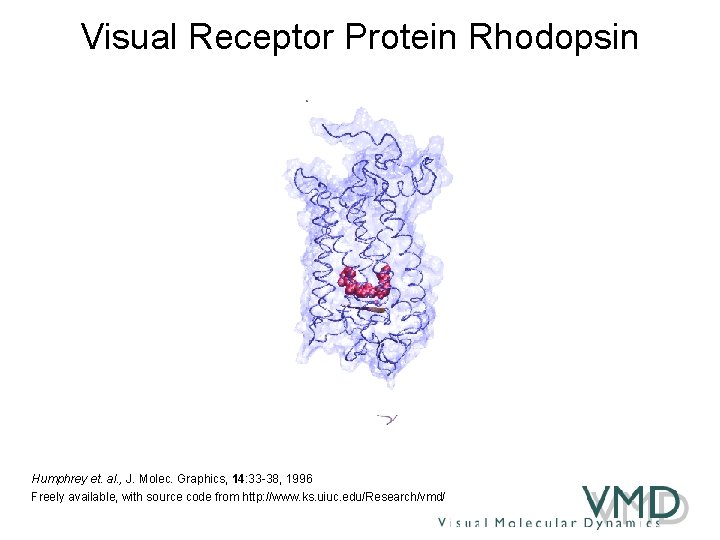
Visual Receptor Protein Rhodopsin Humphrey et. al. , J. Molec. Graphics, 14: 33 -38, 1996 Freely available, with source code from http: //www. ks. uiuc. edu/Research/vmd/

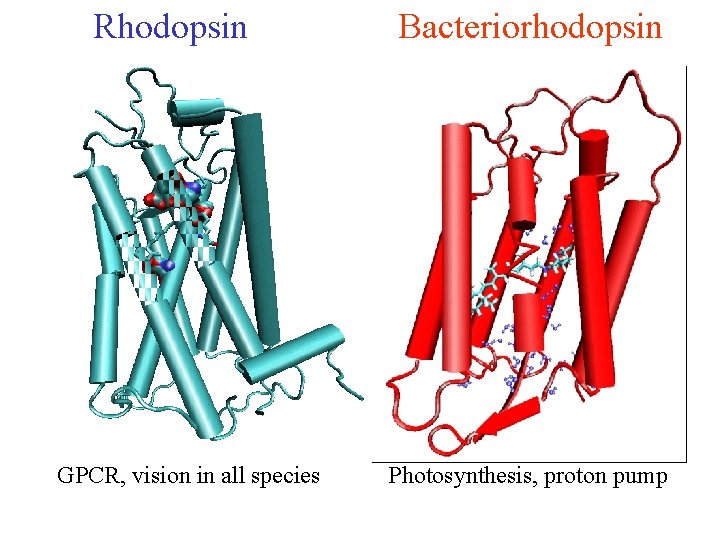
Rhodopsin GPCR, vision in all species Bacteriorhodopsin Photosynthesis, proton pump

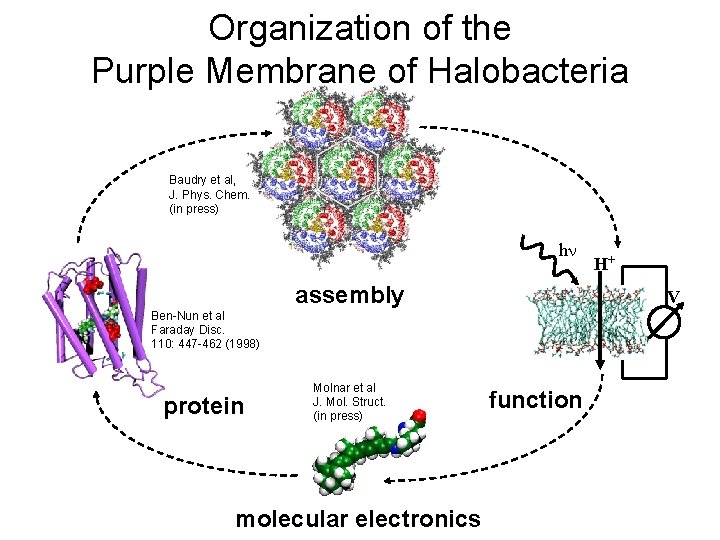
Organization of the Purple Membrane of Halobacteria Baudry et al, J. Phys. Chem. (in press) hn assembly V Ben-Nun et al Faraday Disc. 110: 447 -462 (1998) protein Molnar et al J. Mol. Struct. (in press) molecular electronics H+ function

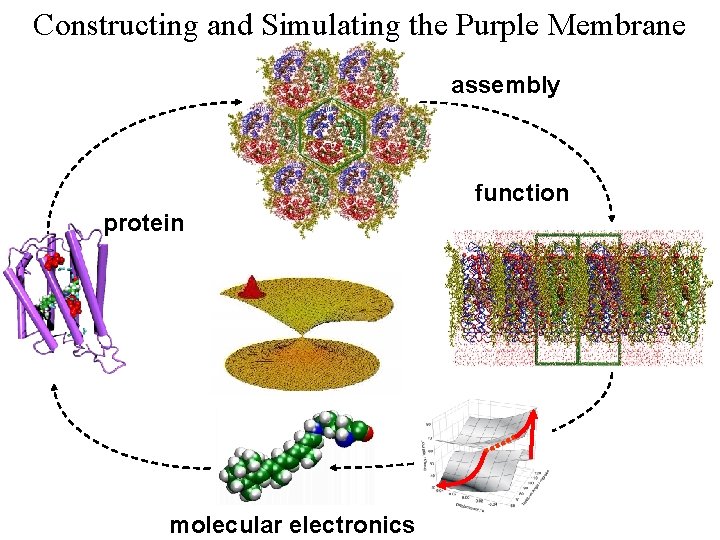
Constructing and Simulating the Purple Membrane assembly function protein b b b Vibrational Spectroscopy (Kyoto) Organic Synthesis (Rehovot) Quantum Chemistry (Heidelberg) Photophysics (Siena) Protein Simulation (Urbana) Pharmacolgy (New York) molecular electronics

Molecular Dynamics Program Used: NAMD 2 hexagonal unit cell 23700 atoms per unit cell # processors Periodic boundary conditions in 3 D (multilayers); Np. T (constant pressure) simulations; Particle Mesh Ewald (no electrostatic cutoff); ~2 weeks/ns on 4 Alpha AXP 21264 -500 Mhz procs.

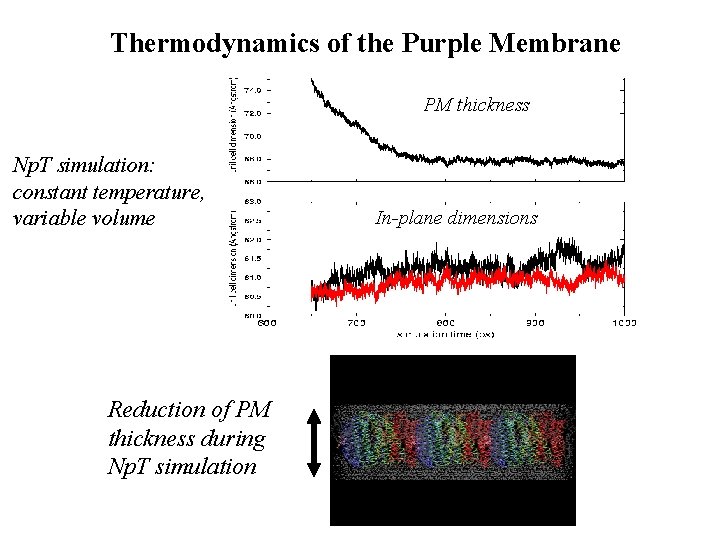
Thermodynamics of the Purple Membrane PM thickness Np. T simulation: constant temperature, variable volume Reduction of PM thickness during Np. T simulation In-plane dimensions

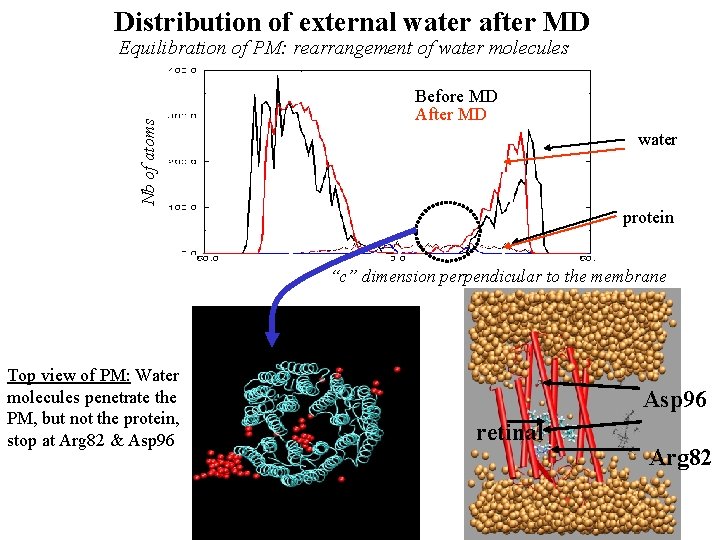
Distribution of external water after MD Nb of atoms Equilibration of PM: rearrangement of water molecules Before MD After MD water protein “c” dimension perpendicular to the membrane Top view of PM: Water molecules penetrate the PM, but not the protein, stop at Arg 82 & Asp 96 retinal Arg 82

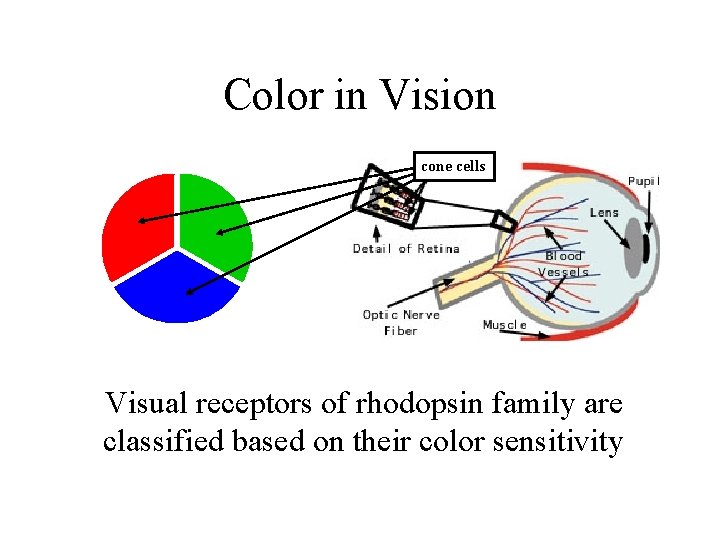
Color in Vision cone cells Visual receptors of rhodopsin family are classified based on their color sensitivity

Rhodopsin Family of Proteins • Seven transmembrane helices • Retinal chromophore bound to a lysine via the Schiff base protonated Schiff base retinal (PSBR)

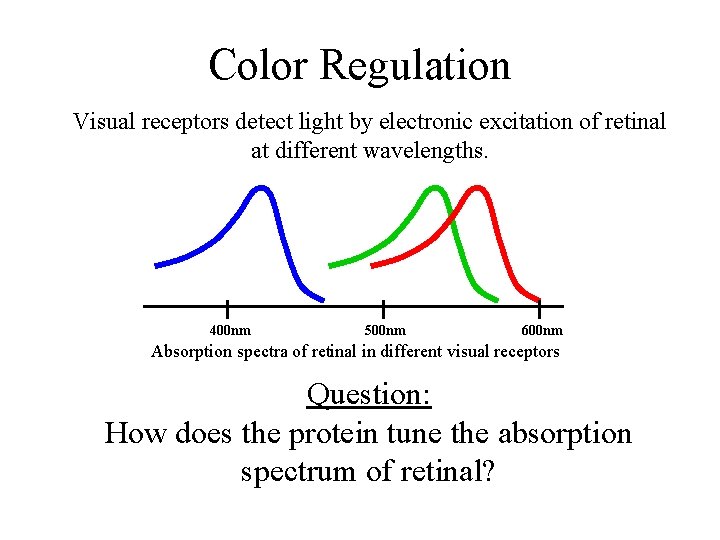
Color Regulation Visual receptors detect light by electronic excitation of retinal at different wavelengths. 400 nm 500 nm 600 nm Absorption spectra of retinal in different visual receptors Question: How does the protein tune the absorption spectrum of retinal?

Spectral Tuning in Archaeal Rhodopsins Sensory Rhodopsin II (s. RII) Repellent response to blue-green light Spectral features • Absorption maximum is strongly blue-shifted (70 nm from b. R). • Prominent sub-band. s. RII 500 nm b. R h. R s. RI 600 nm

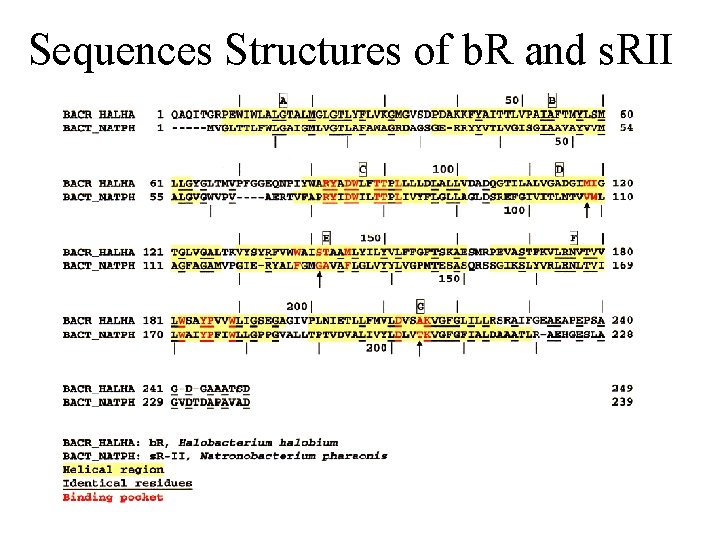
Sequences Structures of b. R and s. RII

X-ray Structures of b. R and s. RII Landau et al. Unique opportunity to study spectral shift given by the availability of X-ray structures. • Structures are homologous. (e. g. , all-trans retinals) orange: s. RII (Natronobacterium pharanois) • Spectra are significantly purple: b. R (Halobacterium salinarum) different.

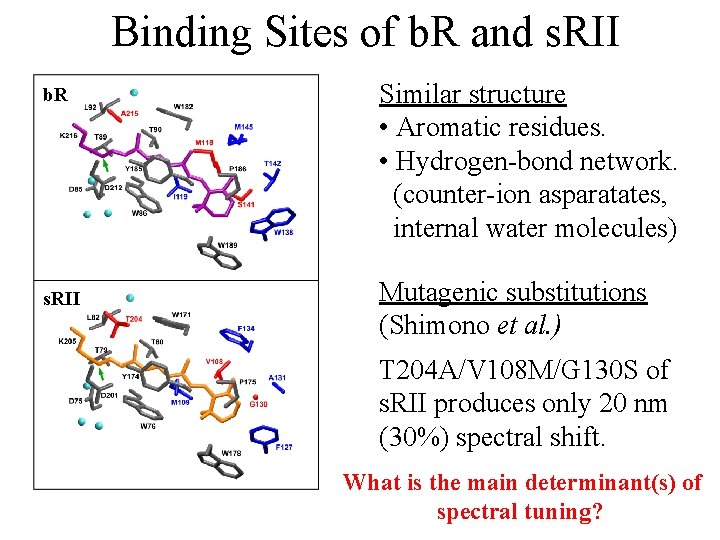
Binding Sites of b. R and s. RII b. R s. RII Similar structure • Aromatic residues. • Hydrogen-bond network. (counter-ion asparatates, internal water molecules) Mutagenic substitutions (Shimono et al. ) T 204 A/V 108 M/G 130 S of s. RII produces only 20 nm (30%) spectral shift. What is the main determinant(s) of spectral tuning?

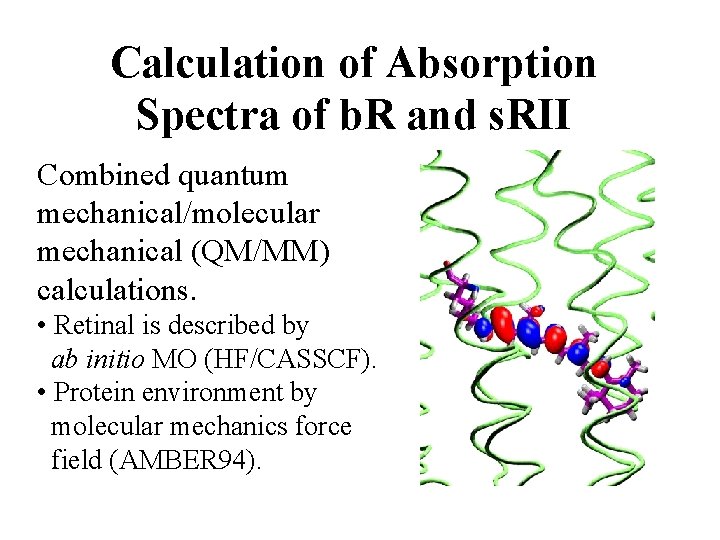
Calculation of Absorption Spectra of b. R and s. RII Combined quantum mechanical/molecular mechanical (QM/MM) calculations. • Retinal is described by ab initio MO (HF/CASSCF). • Protein environment by molecular mechanics force field (AMBER 94).

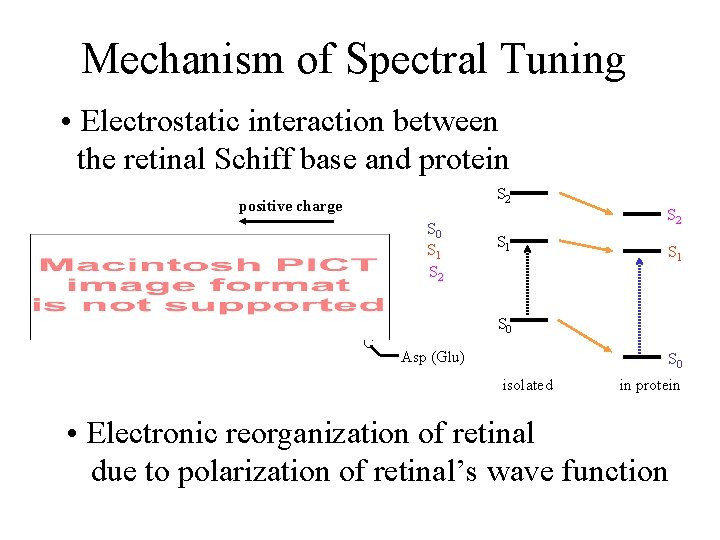
Mechanism of Spectral Tuning • Electrostatic interaction between the retinal Schiff base and protein S 2 positive charge + S 0 S 1 S 2 + O S 1 S 0 O C S 2 Asp (Glu) S 0 isolated in protein • Electronic reorganization of retinal due to polarization of retinal’s wave function

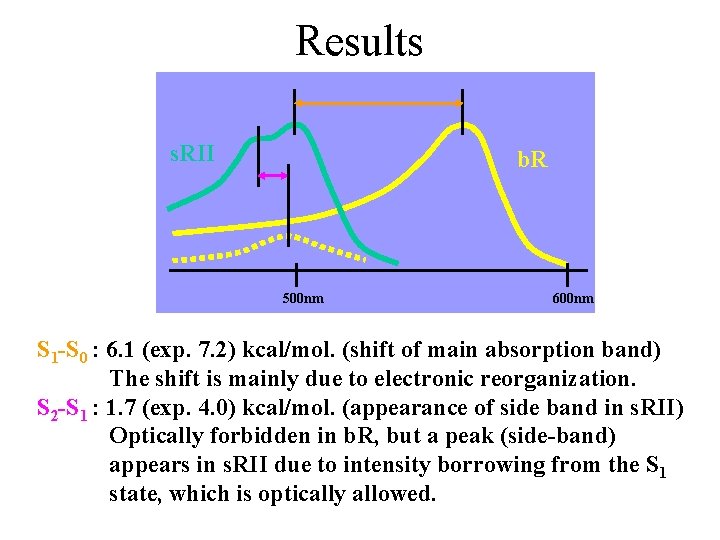
Results s. RII b. R 500 nm 600 nm S 1 -S 0 : 6. 1 (exp. 7. 2) kcal/mol. (shift of main absorption band) The shift is mainly due to electronic reorganization. S 2 -S 1 : 1. 7 (exp. 4. 0) kcal/mol. (appearance of side band in s. RII) Optically forbidden in b. R, but a peak (side-band) appears in s. RII due to intensity borrowing from the S 1 state, which is optically allowed.

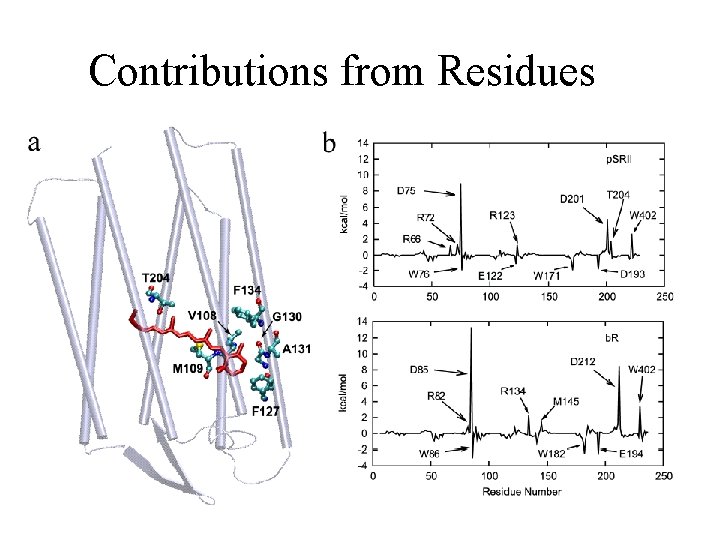
Contributions from Residues

Structural Determinants of Spectral Shift G helix is displaced in s. RII. Distance between the Schiff base and the counter-ion is shorter. G helix N 16 – Cg (Asp 201: s. RII) : 4. 5 A N 16 – Cg (Asp 212: b. R) : 5. 2 A QM/MM optimized structures orange: s. RII, purple: b. R

Rhodopsin Photodynamics Quantum (Wave Packets) Dynamic in protein, 1 -dimensional surface Ben-Nun et al. , Faraday Discussion, 110, 447 - 462 (1998)

Calculation of transition amplitude





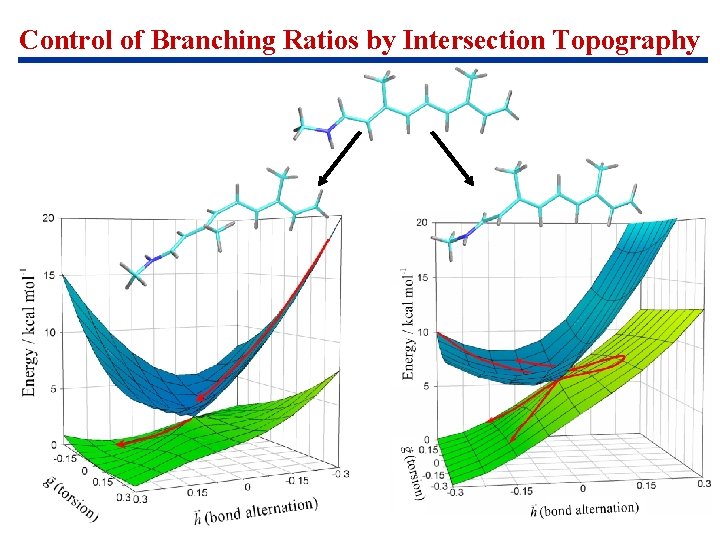
Control of Branching Ratios by Intersection Topography

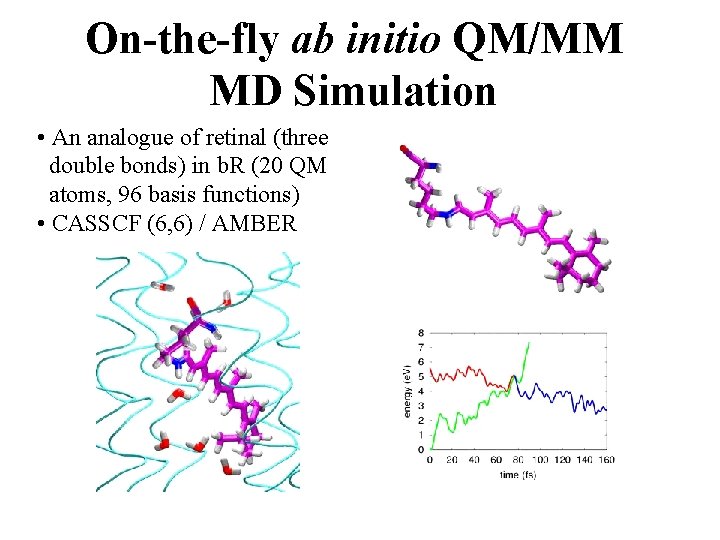
On-the-fly ab initio QM/MM MD Simulation • An analogue of retinal (three double bonds) in b. R (20 QM atoms, 96 basis functions) • CASSCF (6, 6) / AMBER

The Role of Conical Intersection Topography on the Photoisomerization of Retinal Michal Ben-nun Emad Tajkhorshid Shigehito Hayashi Jerome Baudry $$: Beckman Institute, NSF, HFSP, NIH-NCRR