Viruses and Bacteria Viruses Discovery of Viruses Detect

Viruses and Bacteria

Viruses

Discovery of Viruses • Detect viruses indirectly –Too small to see • Adolf Mayer –Cause of Tobacco Mosaic Disease • Stunted growth of tobacco –Contagious through sap • No microbe • Usually small bacteria • Dimitri Ivanowsky –Filter to remove bacteria

• Martinus Beijerinck –Series of infections –Effect undiluted • Pathogen reproducing –Not be cultivated in petri dish • Different properties than bacteria –Smaller and more simple • Wendel Stanley –TMV • Crystallization –Not cells • Electron microscope

Viruses • Nucleic Acids and Proteins • Genome – ds DNA, ss RNA, ds RNA – DNA or RNA virus – Unusual for any other genome – Single linear or circular

• Capsid – Protein shell – Rod shaped, polyhedral, complex – Capsomeres • Protein subunits – Many – Variety small – Bacteriophages • Most complex • Envelope (sometimes) – Accessory structure outside capsid • Help infiltrate host • Made up of – Membrane of host – Viral glycoproteins/proteins and enzymes
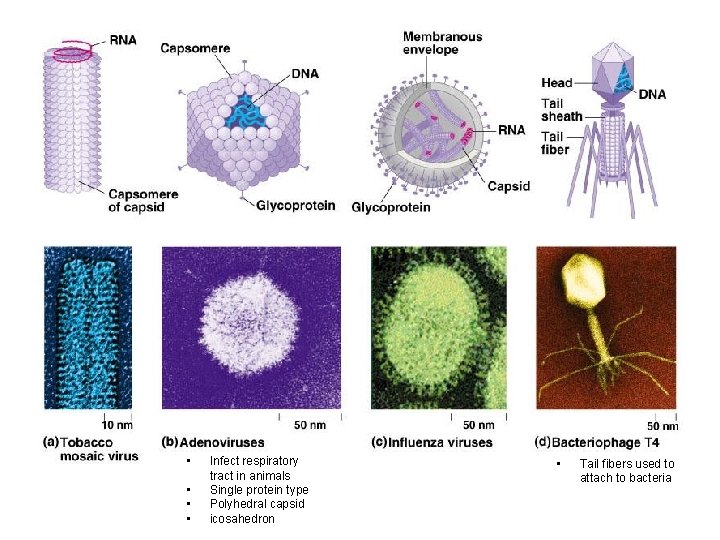
• • Infect respiratory tract in animals Single protein type Polyhedral capsid icosahedron • Tail fibers used to attach to bacteria
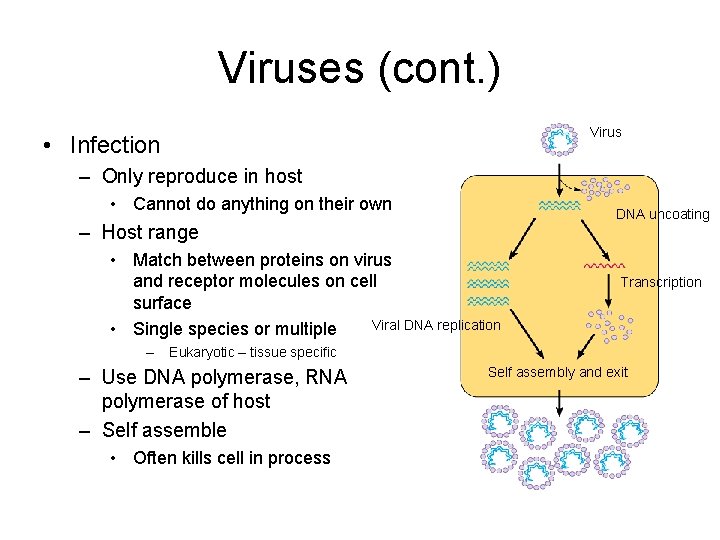
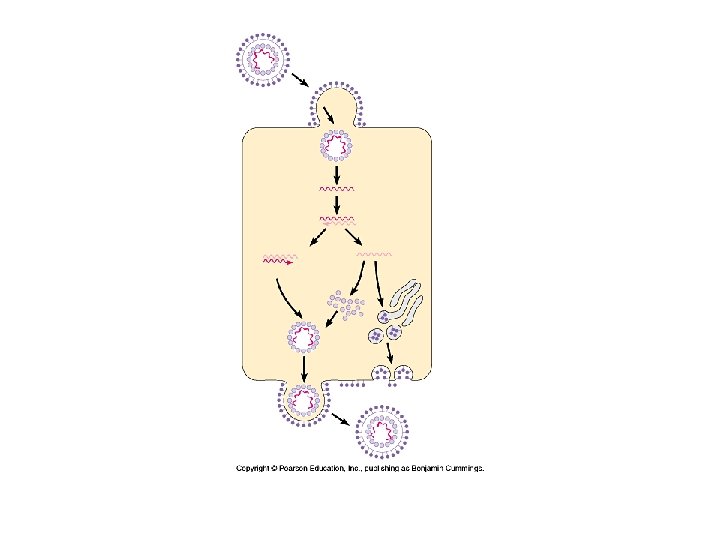
Viruses (cont. ) Virus • Infection – Only reproduce in host • Cannot do anything on their own DNA uncoating – Host range • Match between proteins on virus and receptor molecules on cell surface Viral DNA replication • Single species or multiple – Transcription Eukaryotic – tissue specific – Use DNA polymerase, RNA polymerase of host – Self assemble • Often kills cell in process Self assembly and exit

Bacterial Viruses • Bacteriophage • Lytic Cycle – Death of host – Lyses host – Virulent viruses – Defenses – Receptor sites not recognized by phages – Restriction nucleases – Cut up foreign DNA
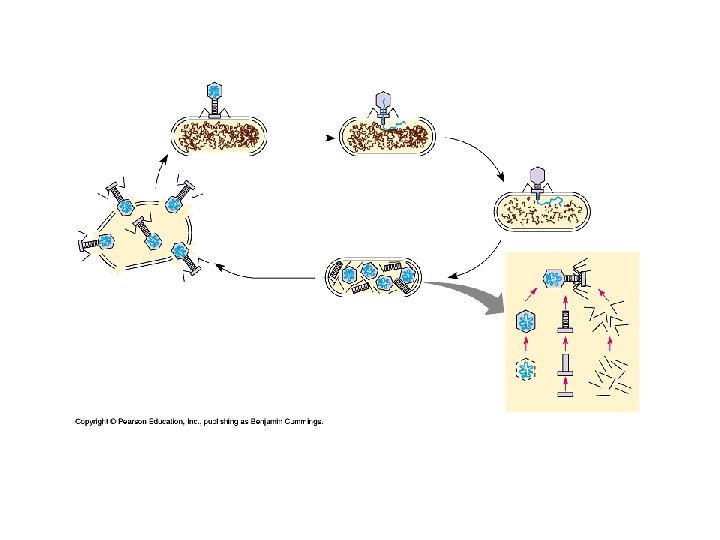

Bacterial Viruses (cont. ) • Lysogenic Cycle – Reproduces without killing host – Temperate virus • Both modes of replication – Prophage • Viral DNA incorporated into host genome – Silent in host » Repressor prophage » Alter phenotype of host cell • Active phages – Viral DNA exits host chromosome » Lytic cycle » Environmental factor
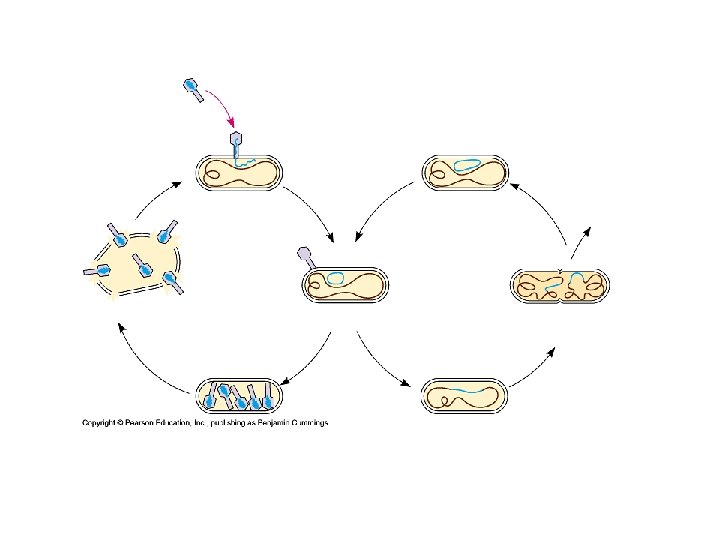

Animal Viruses • Envelopes – Lipid bilayer – like cell • Glycoproteins on surface – Spikes” – Specific receptors • Fuses with plasma membrane • In host – Creates new glycoproteins • ER • Transported to membrane • Clusters – exit points for new viruses – Like exocytosis – Does necessarily kill the cell • Cell guilty af Membrane often but not always host plasma membrane • Herpesvirus – host nuclear membrane • Reproduce within nuclear envelope • Some remain in nuclei of nerve cells

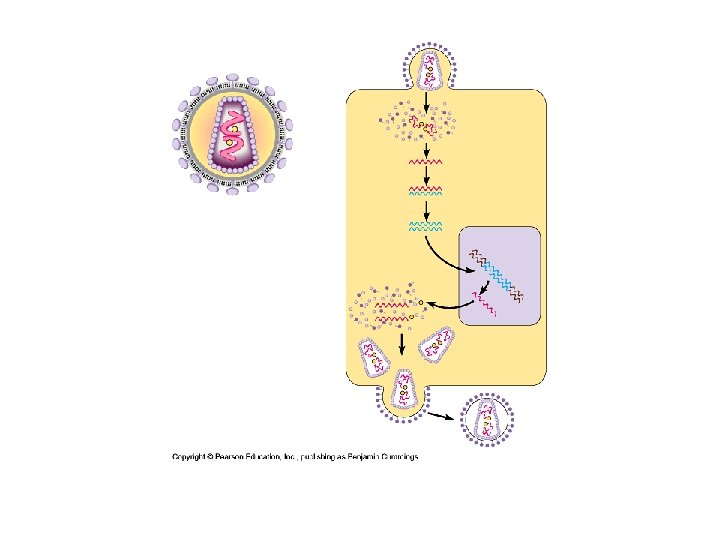
Animal Viruses (cont. ) • RNA viruses – Many different classes – Retrovirus • ss RNA • Reverse transcriptase – Viral RNA as template for DNA • Provirus – New DNA integrated into host DNA » m. RNA » protein synthesis and template for more viral DNA – Does not leave host • HIV → AIDS

Animal Viruses (cont. ) • Medical weapons – Vaccines – Harmless variants or derivatives – Antiviral drugs – Few – Interfere with nucleic acid synthesis • Tumor viruses – Transform cells – Integration of viral DNA – Oncogenes – Code for growth factors – Overexpression – Effective at causing cancer with endogenous factors • Emerging viruses

Plant Viruses • RNA viruses • Horizontal Transmission • External source – Passes the epidermis – Susceptible through weather, insects, farmers • Plasmodesmata – Viral particles through cell walls • Vertical transmission • Inherited from parents – Asexual or sexual

Viroids and Prions • Viroids – Smaller and simpler – “Naked” RNA • Circular – Replicate using host enzymes – Disrupt metabolism, development, and growth in plants • Prions – Infectious proteins • Cannot replicate itself – Misfolded brain protein • Converts normal proteins into prions – Degenerative brain illnesses

Evolutionary Origins • Evolved after first cells • Fragments of cellular nucleic acids • Sources • Plasmids • circular DNA in bacteria and yeasts • Transposons • small mobile DNA segments

Bacteria

Bacteria • Genome – Double stranded circular chromosome – Much less DNA than eukaryotic – Nucleoid – Densely packed region of DNA – No membrane – Simple – Very few associated proteins – Plasmids

• Reproduction – Binary fission – Single origin of replication – Produces clones – Mutations – Very rare – Significant impact – Rapid replication

Genetic recombination – Transformation • Process of gene transfer • Bacterial cell assimilates genetic material from surroundings • Genetic recombination • Foreign allele replaces native allele • Recombinant • Some have surface proteins recognize DNA • Only closely related species • E Coli does not • Plasmid

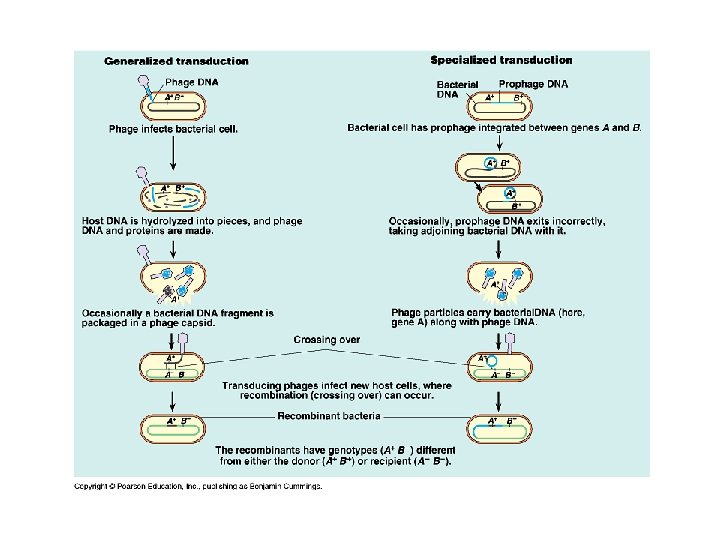
Genetic recombination • Transduction – Gene transferred by a phage – General • Host cell DNA accidentally packed in capsid – Can attach to another bacteria – Inject gene from old bacteria » Replace homologous region » Random gene transferred – Specialized • Excises host genes adjacent to prophage – Injected into new bacteria – Specific genes near the prophage

Plasmid • Self replicating DNA • Separate from bacterial DNA • F plasmid • Reversible incorporation • Episome • Can be either bacterial DNA or plasmid • Some temperate phages • No protein coat • Cannot exist outside the cell • Beneficial to host • Not usually needed • Used under stress

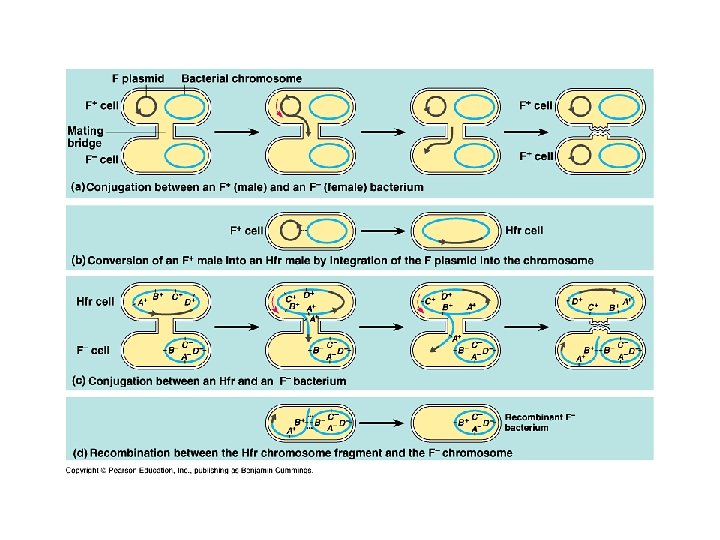
Genetic recombination – Conjugation • Transfer of genes between 2 bacteria cells • Temporarily joined

F factor • F factor – gene form • “male” • • F+ Extends sex pili • • Draws other cell in to create a cytoplasmic bridge Contains and donates F plasmid • “female” • F- • Contagious • F- becomes F+ • Hfr cell • • Integrated plasmid Donates DNA with F factor • • Usually not entire DNA Recipient becomes a recombinant

• R plasmids –Antibiotic resistance

Genetic Recombination (cont. ) • Transposons • DNA sequences that can move from one chromosomal site to another • Cannot exist independently • Recombination with target DNA site • Bacteria • • • within DNA Between plasmids • Antibiotic resistance plasmid to DNA Cut and paste Replicative • Copy inserted into gene • Move to many different locations • Not limited by similar base pairing

Bacterial Transposons • Insertion Sequences • Simplest • site specific • Composite • Help host adapt • Found in Eukaryotic cells

Metabolic control • Regulate amount of an enzyme made –Gene expression • Change activity of current enzymes –Sensitivity to chemical cues • Change catalytic activity

Control of gene expression – Operon • Operator • Single promoter region • Genes controlled • Produces polycistronic m. RNA – Transcript of several genes – Translated in separate polypeptides • Advantages • Expression can be coordinated • Entire operon can be controlled by a single operator
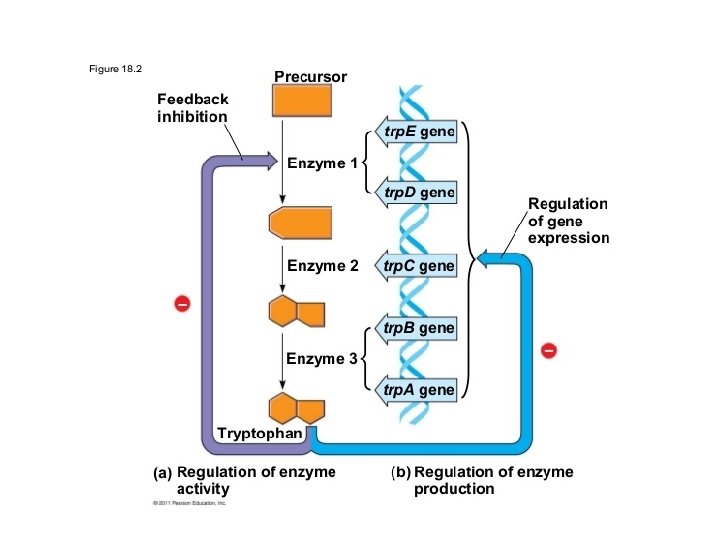

Bacteria (cont. ) • Operator • DNA segment between promoter and gene • “on/off” switch • Naturally “on”

• Repressor • Switches operator “off” • Depends on amount of repressor molecules • Protein • Operator specific • Blocks transcription • Similar to enzyme • Active site • Binds reversibly • Encoded by regulatory genes

Bacteria (cont. ) • Operon (cont. ) – Regulatory gene • Code for repressor protein • Located some distance from operon • Constantly transcribed – Repressible operon • Inhibited by metabolite • Trp operon – Trp. R » Allosteric » Two configurations » Synthesized inactively » Activated by tryptophan » Corepressor – acts with repressor to stup an operon “off”
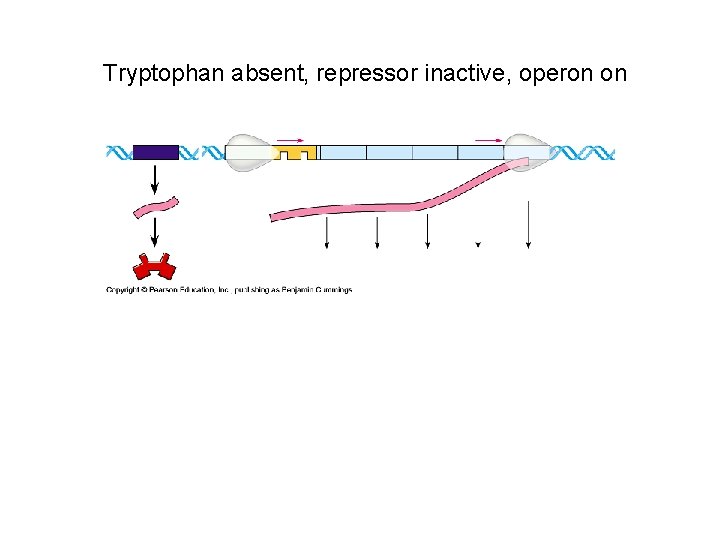
Tryptophan absent, repressor inactive, operon on
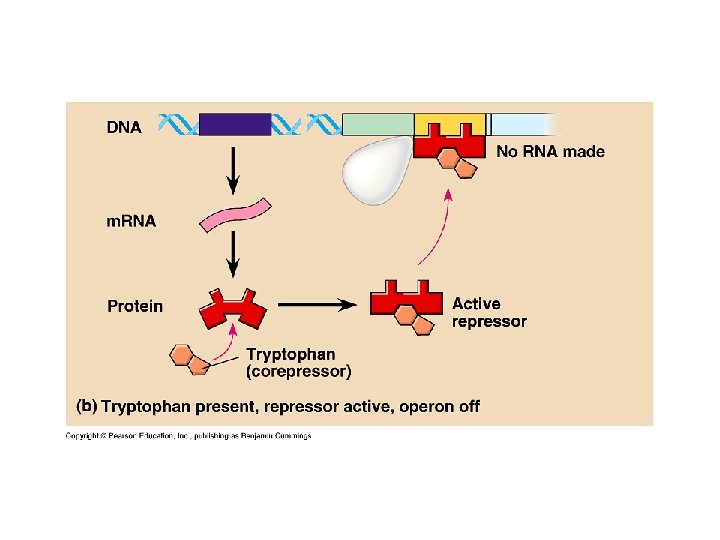

Bacteria (cont. ) • Operon (cont. ) – Inducible operon • Stimulated by metabolite • Lac operon • Inducer – allolactose • Inactivates repressor – Repressible • Anabolic pathways • Synthesize end products from precursors – Inducible • Catabolic pathways • Produce enzymes to break nutrients down – Both negative control of genes
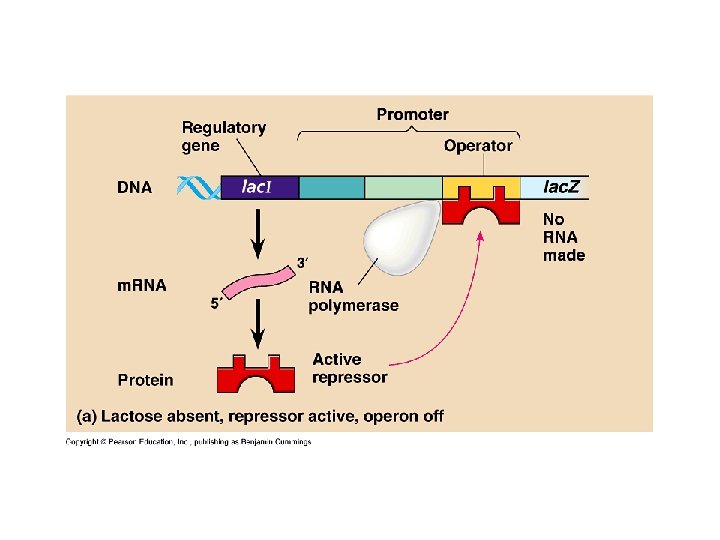
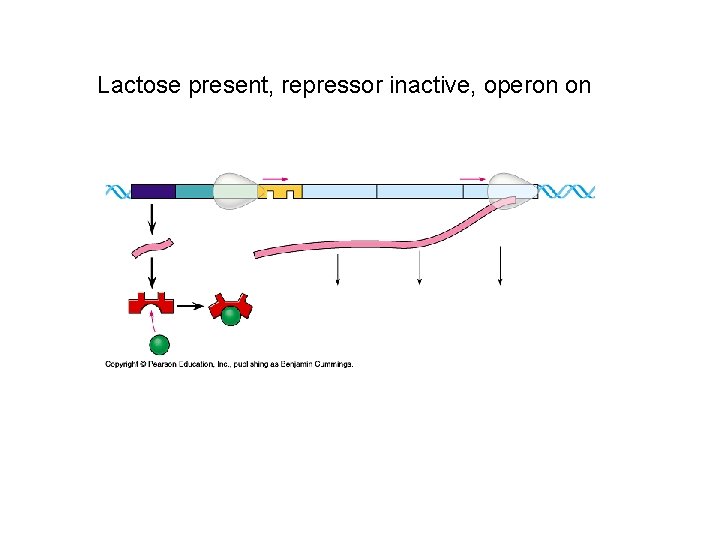
Lactose present, repressor inactive, operon on
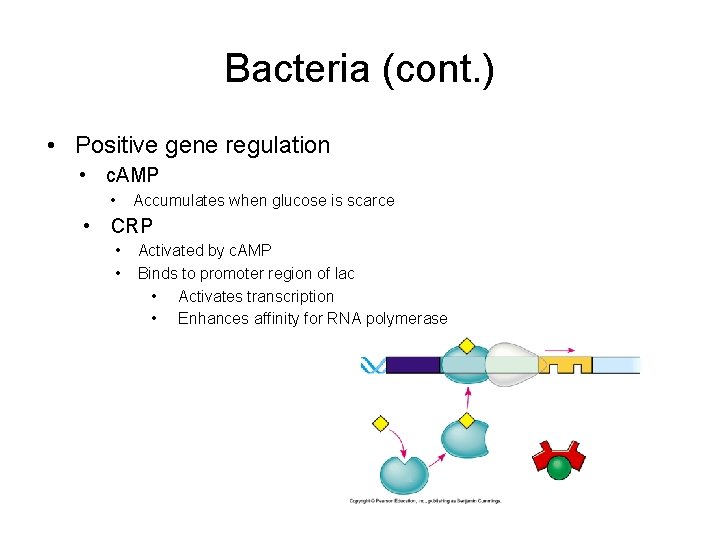
Bacteria (cont. ) • Positive gene regulation • c. AMP • Accumulates when glucose is scarce • CRP • • Activated by c. AMP Binds to promoter region of lac • Activates transcription • Enhances affinity for RNA polymerase

Organization and control of Eukaryotic cells

DNA packing • Bacterial DNA – complex coiling and looping • Eukaryotic genome –Highly folded • Chromatin –Loosely organized – 2 nm

Nucleosomes • First level of DNA packing • Histone proteins –Charged • Positive • Attach to negatively charged DNA • DNA histone complex = chromatin –DNA wound around protein core • one of four histone protein –Remains intact except during replication • Next level of packing – fifth histone protein attaches to DNA • Near bead

Higher levels • Chromosomes during mitosis –Histone H 1 • Coils/folds into 30 nm chromatin fiber –Looped domains » Attached to chromosome scaffold » Nonhistone proteins –Further coiling into metaphase chromosome –Villae

Higher levels • Interphase chromatin –Less condensed –Looped domains • Attached to nuclear lamina/ nuclear matrix –Active transcription sites • Restricted area of nucleus –Different chromosomes don’t get entangled –Euchromatin • Not as condensed –Heterochromatin • Highly condensed • Not transcribed

Eukaryotic genome • Code for proteins • Regulatory sequences • Sequences without known functions –Introns –Repetitive DNA

Repetitive DNA • Tandemly repetitive DNA –Identical repeated units adjacent –Often different density from other DNA • Satellite DNA –Classified with total length of DNA » Regular, mini, micro –Genetic disorders – Located at telomeres and centromeres • Structural role in chromosomes –Organize chromatin during interphase

Repetitive DNA • Interspersed Repetitive DNA – Repeated units scattered across genome – copies similar but not identical – 25 – 40% of mammalian genomes • Humans and primates –Alu elements » Transcribed » Function unknown

Gene evolution • Multigene family –Collection of genes similar or identical in sequence • Evolution • Pseudo genes –Non functional gene • Random mutations destroyed function –Lacks sites for expression • Introns –Similar to functional genes

Gene amplification and loss • Selective replication of certain genes –Ovum • Increased expression of r. RNA –Dna circles in nucleoli –Unable to replicate further • Able to make large amounts of ribosomes –Cancer • Resistent to chemotherapy • Amplification of chemo resistant genes • Lost genes –Insect development • Entire chromosomes can be lost

Rearrangements • Transposons – Insert into transcription regulatory sequence • Increase/decrease production – Can carry gene • activated by an upstream promoter site • Retrotransposons – move through transcripts of DNA

• Immunoglobulin genes – encode antibodies • Genes are pieced together from all over the genome

• DNA methylation – Attachment of methyl groups • Turn gene on and off – genomic imprinting • permanently turns off maternal or paternal allele – beginning of development • Histone acetylation
• Transcriptional factors – proximal control elements • promoter – TATA – Distal control elements • Enhancers – helps polymerase bind to DNA • Activator – DNA binding domain – Alternative RNA splicing • introns

Cancer • Oncogenes – potential to cause cancer • Proto-oncogenes – regulate cell growth – mutation makes it into an oncogene • Tumor suppressor
- Slides: 60