Virtual Laboratory for eScience VLe Henri Bal Department

Virtual Laboratory for e-Science (VL-e) Henri Bal Department of Computer Science Vrije Universiteit Amsterdam bal@cs. vu. nl vrije Universiteit

Outline • • e-Science and virtual laboratories The VL-e project VL-e and networking Case studies: o Visualization o Interactive problem solving environments o Distributed supercomputing • Computing/networking infrastructure
e-Science • Web is about exchanging information • Grid is about sharing resources o Computers, data bases, instruments, services • e-Science supports experimental science by providing a virtual laboratory on top of Grids
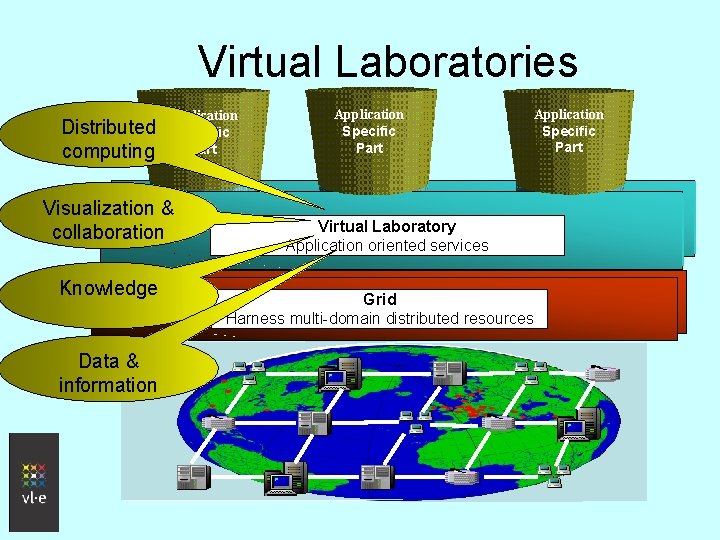
Virtual Laboratories Distributed computing Application Specific Part Potential Generic Visualization & part Management collaboration of comm. & computing Knowledge Data & information Application Specific Part Potential Generic part Virtual Laboratory Management of comm. & services Application oriented computing Application Specific Part Potential Generic part Management of comm. & computing Grid Harness multi-domain distributed resources
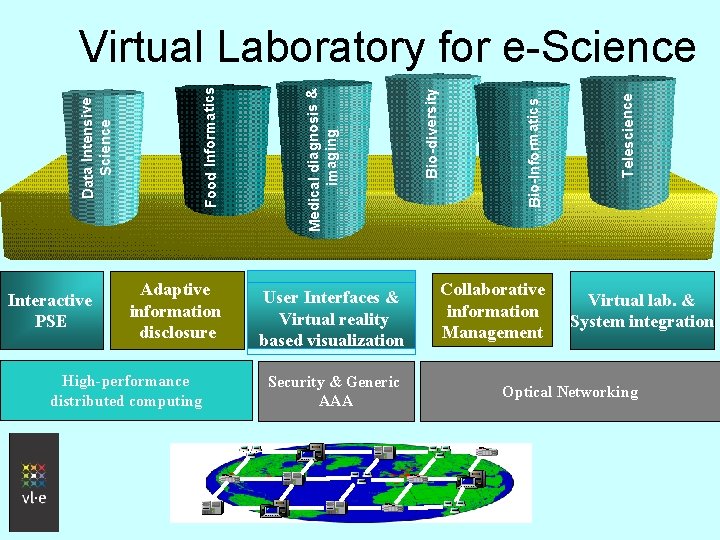
Interactive PSE Adaptive information disclosure High-performance distributed computing User Interfaces & Virtual reality based visualization Security & Generic AAA Collaborative information Management Telescience Bio-Informatics Bio-diversity Medical diagnosis & imaging Food Informatics Data Intensive Science Virtual Laboratory for e-Science Virtual lab. & System integration Optical Networking

The VL-e project • 40 M€ (20 M€ BSIK funding) • 2004 - 2008 vrije Universiteit • 20 partners • Academic - Industrial

VL-e and networking • e-Science applications generate much (distributed) data o High-resolution imaging o Bio-informatics queries o Particle physics: o Currently: 1 PByte per year o LHC (2007): 10 -30 PByte per year • Virtual laboratories need high-speed networks for o Remote visualization o Interactive problem solving environments o Distributed supercomputing
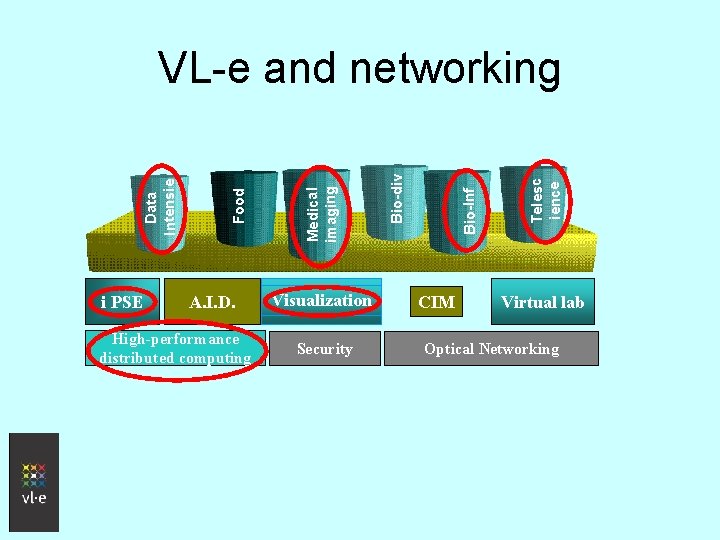
i PSE A. I. D. High-performance distributed computing Visualization Security CIM Telesc ience Bio-Inf Bio-div Medical imaging Food Data Intensie VL-e and networking Virtual lab Optical Networking

Visualization on the Grid

Visualization on the Grid
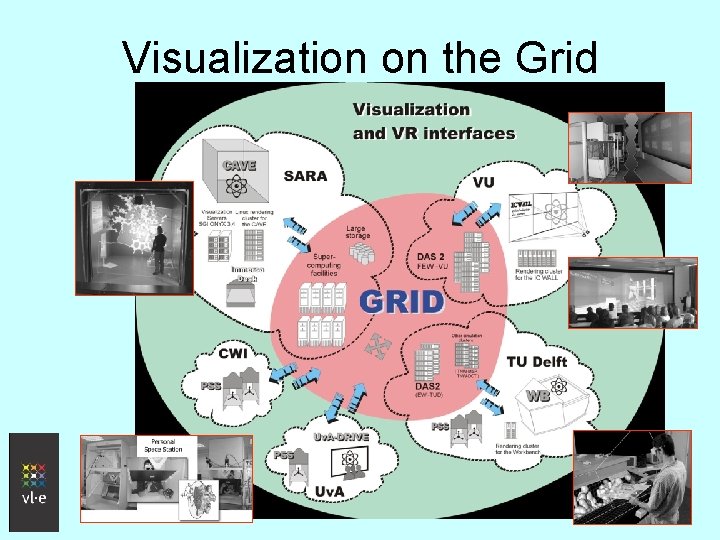
Visualization on the Grid

Visualization on the Grid
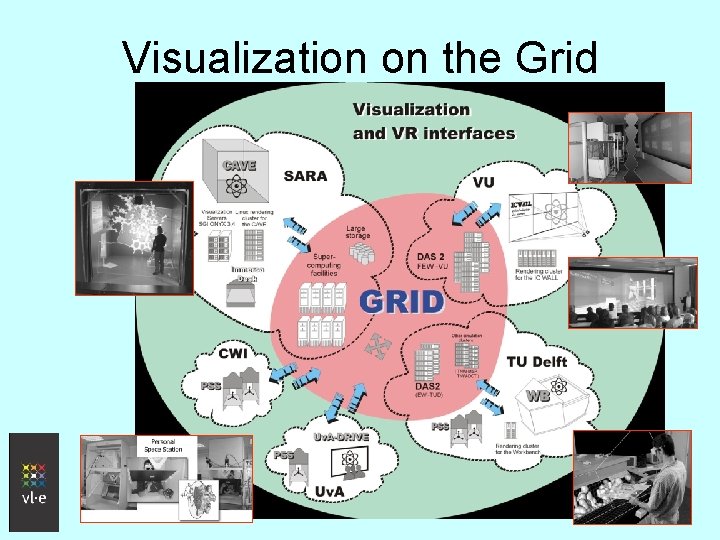
Visualization on the Grid
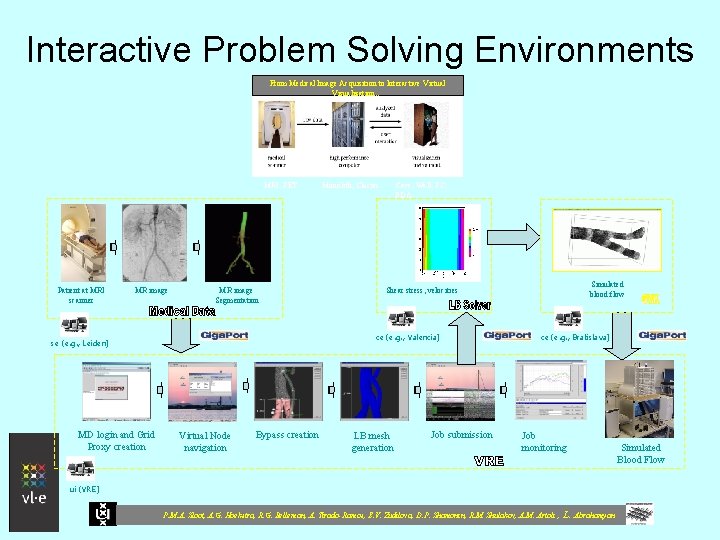
Interactive Problem Solving Environments From Medical Image Acquisition to Interactive Virtual Visualization… MRI, PET Patient at MRI scanner MR image Segmentation Cave, Wall, PC, PDA Virtual Node navigation Bypass creation Simulated blood flow Shear stress, velocities ce (e. g. , Valencia) se (e. g. , Leiden) MD login and Grid Proxy creation Monolith, Cluster LB mesh generation Job submission ce (e. g. , Bratislava) Job monitoring ui (VRE) P. M. A. Sloot, A. G. Hoekstra, R. G. Belleman, A. Tirado-Ramos, E. V. Zudilova, D. P. Shamonin, R. M. Shulakov, A. M. Artoli , L. Abrahamyan Simulated Blood Flow
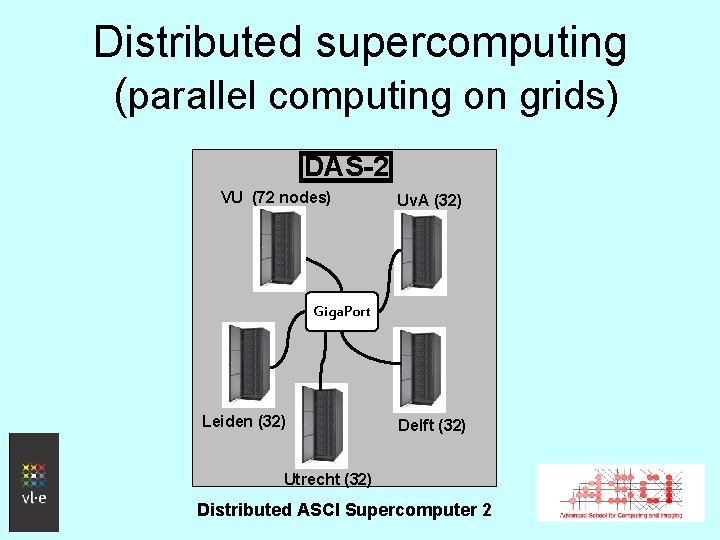
Distributed supercomputing (parallel computing on grids) DAS-2 VU (72 nodes) Uv. A (32) Giga. Port Leiden (32) Delft (32) Utrecht (32) Distributed ASCI Supercomputer 2

Distributed supercomputing (parallel computing on grids)

HPC on a grid? • Can grids be used for High-Performance Computing applications that are not trivially parallel? • Key: grids usually are hierarchical o Collections of clusters, supercomputers o Fast local links, slow wide-area links • Can optimize algorithms to exploit this hierarchy o Message combining + latency hiding on wide-area links o Optimized collective communication operations (broadcast etc. ) o Often gives latency-insensitive, throughput-bound algorithms

Ibis: a Java-centric grid programming environment • Written in pure Java, runs on heterogeneous grids o “Write once, run everywhere ” • Many applications: o Electromagnetic simulation (Jem 3 D) o Automated protein identification (VL-e application from AMOLF) o N-body simulations o SAT-solver o Raytracer Jem 3 D (see SC’ 04) Available from www. cs. vu. nl/ibis

Networking demands • Low latency is needed for o Interactive visualization o Interactive Problem Solving Environments o Synchronous, latency-sensitive parallel algorithms • High throughput is needed for o Data-intensive e-Science applications o Visualization of large data sets o Asynchronous, throughput-bound parallel algorithms • Efficient collective (group) communication for o Collaborative visualization between multiple sites o Collective operations in parallel algorithms

Outline • • e-Science and virtual laboratories The VL-e project VL-e and networking Examples: o Visualization o Interactive Problem Solving Environments o Distributed supercomputing • Computing/networking infrastructure
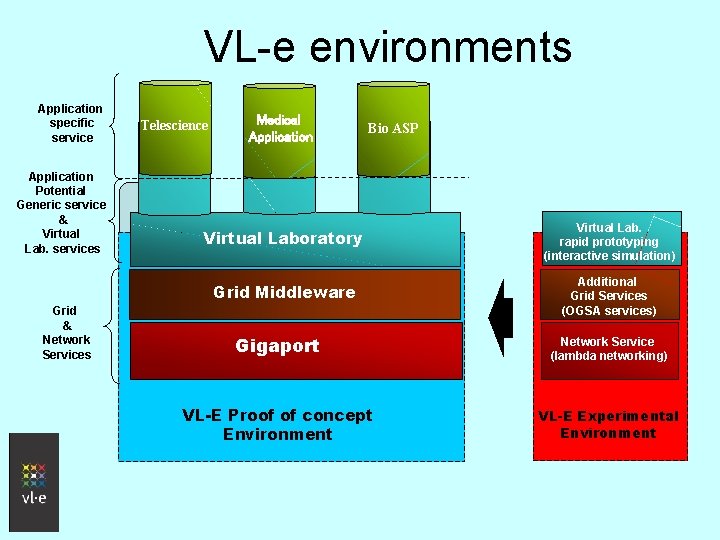
VL-e environments Application specific service Application Potential Generic service & Virtual Lab. services Grid & Network Services Telescience Medical Application Bio ASP Virtual Laboratory Virtual Lab. rapid prototyping (interactive simulation) Grid Middleware Additional Grid Services (OGSA services) Gigaport Network Service (lambda networking) VL-E Proof of concept Environment VL-E Experimental Environment

DAS-3 • Proposed next generation grid in the Netherlands • Partners: o ASCI research school (VU, Uv. A, TU Delft, Leiden) o Gigaport-NG/SURFnet: DWDM computer backplane (dedicated optical group of 8 lambdas) o VL-e and Multimedia. N BSIK projects • Topology controlled by applications through the Network Operations Center
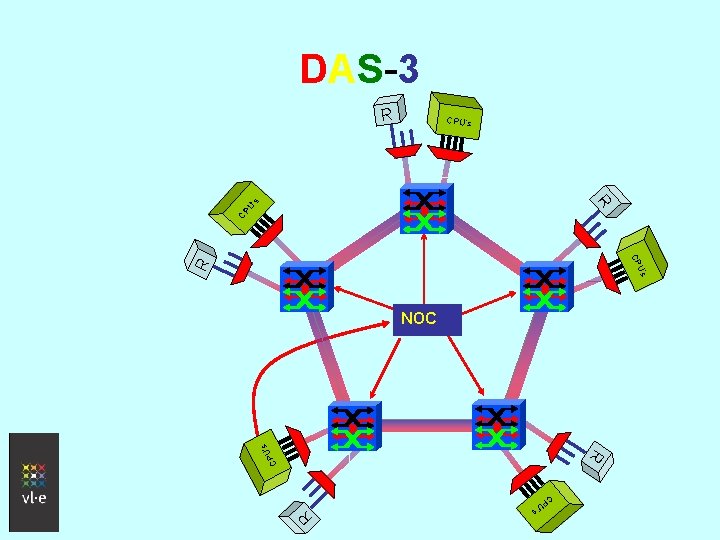
DAS-3 R CP U’ R s CPU’s R CP CP U’ s R C R ’s PU NOC

Summary • VL-e (Virtual Laboratory for e-Science) studies entire e-Science chain, including applications, middleware and grids • High networking demands from applications and generic methods • New state-of-the-art Grid infrastructure planned for 2006 using optical networking
- Slides: 24